120-08-RV
| 4969. | +/- | 6. |
| 5013. | +/- | 93. |
| 2.90 | +/- | 0. |
| 2.54 | +/- | 0. |
| 1.51 | +/- | 0. |
| 1.51 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.37 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.94 |
| -9999.00 |

120-08-RV
| 4769. | +/- | 5. |
| 4833. | +/- | 87. |
| 2.72 | +/- | 0. |
| 2.40 | +/- | 0. |
| 1.45 | +/- | 0. |
| 1.45 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.17 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.26 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.68 |
| -9999.00 |

PERSIST_LOW
120-08-RV
| 4301. | +/- | 2. |
| 4386. | +/- | 73. |
| 2.27 | +/- | 0. |
| 2.02 | +/- | 0. |
| 1.42 | +/- | 0. |
| 1.42 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.81 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.24 |
| -9999.00 |

PERSIST_MED,PERSIST_LOW
120-08-RV
| 4131. | +/- | 1. |
| 4234. | +/- | 73. |
| 1.77 | +/- | 0. |
| 1.60 | +/- | 0. |
| 1.65 | +/- | 0. |
| 1.65 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -0.35 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.07 | +/- | 0. |
| 3.63 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.83 |
| -9999.00 |

PERSIST_MED,PERSIST_LOW
120-08-RV
| 4443. | +/- | 3. |
| 4546. | +/- | 81. |
| 2.19 | +/- | 0. |
| 2.00 | +/- | 0. |
| 1.69 | +/- | 0. |
| 1.69 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -0.35 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.06 | +/- | 0. |
| 3.62 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.43 |
| -9999.00 |

120-08-RV
| 4196. | +/- | 2. |
| 4295. | +/- | 74. |
| 1.61 | +/- | 0. |
| 1.44 | +/- | 0. |
| 1.55 | +/- | 0. |
| 1.55 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -0.25 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.40 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.00 | +/- | 0. |
| 3.42 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.58 |
| -9999.00 |

LOW_SNR,PERSIST_LOW
STAR_WARN,SN_WARN
120-08-RV
| 4371. | +/- | 6. |
| 4467. | +/- | 107. |
| 2.16 | +/- | 0. |
| 1.94 | +/- | 0. |
| 1.36 | +/- | 0. |
| 1.36 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -0.18 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.29 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| -0.55 |
| -9999.00 |

BRIGHT_NEIGHBOR
120-08-RV
| 4642. | +/- | 6. |
| 4729. | +/- | 97. |
| 2.58 | +/- | 0. |
| 2.40 | +/- | 0. |
| 1.27 | +/- | 0. |
| 1.27 | +/- | -NaN |
| -0.34 | +/- | 0. |
| -0.34 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.07 | +/- | 0. |
| 3.61 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -1.20 |
| -9999.00 |
PERSIST_MED,PERSIST_LOW
120-08-RV
| 4218. | +/- | 1. |
| 4312. | +/- | 73. |
| 2.00 | +/- | 0. |
| 1.79 | +/- | 0. |
| 1.50 | +/- | 0. |
| 1.50 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.13 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.19 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.39 |
| -9999.00 |

PERSIST_MED,PERSIST_LOW
120-08-RV
| 4807. | +/- | 5. |
| 4870. | +/- | 89. |
| 2.75 | +/- | 0. |
| 2.42 | +/- | 0. |
| 1.52 | +/- | 0. |
| 1.52 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.24 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.41 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.51 |
| -9999.00 |

120-08-RV
| 4467. | +/- | 2. |
| 4558. | +/- | 78. |
| 2.45 | +/- | 0. |
| 2.26 | +/- | 0. |
| 1.37 | +/- | 0. |
| 1.37 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.11 | +/- | 0. |
| 3.06 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.32 |
| -9999.00 |

120-08-RV
| 5039. | +/- | 7. |
| 5073. | +/- | 95. |
| 2.99 | +/- | 0. |
| 2.61 | +/- | 0. |
| 1.47 | +/- | 0. |
| 1.47 | +/- | -NaN |
| -0.21 | +/- | 0. |
| -0.21 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.34 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.14 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -1.00 |
| -9999.00 |

120-08-RV
| 4524. | +/- | 3. |
| 4613. | +/- | 80. |
| 2.59 | +/- | 0. |
| 2.38 | +/- | 0. |
| 1.30 | +/- | 0. |
| 1.30 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.13 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES
STAR_WARN,COLORTE_WARN
120-08-RV
| 8216. | +/- | 28. |
| 8115. | +/- | 312. |
| 4.05 | +/- | 0. |
| 4.60 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 75.56 | +/- | 0. |
| 1.88 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.38 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -2.41 |
| -9999.00 |
PERSIST_MED,PERSIST_LOW
120-08-RV
| 4463. | +/- | 2. |
| 4568. | +/- | 81. |
| 2.51 | +/- | 0. |
| 2.33 | +/- | 0. |
| 1.22 | +/- | 0. |
| 1.22 | +/- | -NaN |
| -0.39 | +/- | 0. |
| -0.39 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.17 | +/- | 0. |
| 3.71 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.69 |
| -9999.00 |
PERSIST_MED,PERSIST_LOW
120-08-RV
| 4065. | +/- | 1. |
| 4166. | +/- | 71. |
| 1.61 | +/- | 0. |
| 1.44 | +/- | 0. |
| 1.67 | +/- | 0. |
| 1.67 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -0.28 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.49 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -1.15 |
| -9999.00 |

SUSPECT_RV_COMBINATION
STAR_BAD
120-08-RV
| 8382. | +/- | 31. |
| -10000. | +/- | -NaN |
| 4.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -1.45 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 90.43 | +/- | 0. |
| 1.96 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.58 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -1.62 |
| -9999.00 |
PERSIST_LOW
120-08-RV
| 4714. | +/- | 4. |
| 4771. | +/- | 82. |
| 3.34 | +/- | 0. |
| 3.15 | +/- | 0. |
| 0.91 | +/- | 0. |
| 0.91 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.69 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.06 |
| -9999.00 |
PERSIST_MED,PERSIST_LOW
120-08-RV
| 4718. | +/- | 4. |
| 4784. | +/- | 84. |
| 2.73 | +/- | 0. |
| 2.41 | +/- | 0. |
| 1.53 | +/- | 0. |
| 1.53 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 3.05 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.21 |
| -9999.00 |

120-08-RV
| 4404. | +/- | 2. |
| 4477. | +/- | 73. |
| 2.66 | +/- | 0. |
| 2.36 | +/- | 0. |
| 0.98 | +/- | 0. |
| 0.98 | +/- | -NaN |
| 0.36 | +/- | 0. |
| 0.36 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.40 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| 0.36 |
| -9999.00 |

PERSIST_LOW
120-08-RV
| 3840. | +/- | 1. |
| 3927. | +/- | 64. |
| 1.44 | +/- | 0. |
| 1.25 | +/- | 0. |
| 1.61 | +/- | 0. |
| 1.61 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.05 | +/- | 0. |
| 2.89 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.11 |
| -9999.00 |
120-08-RV
| 4789. | +/- | 5. |
| 4848. | +/- | 87. |
| 2.76 | +/- | 0. |
| 2.43 | +/- | 0. |
| 1.57 | +/- | 0. |
| 1.57 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.10 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.02 |
| -9999.00 |
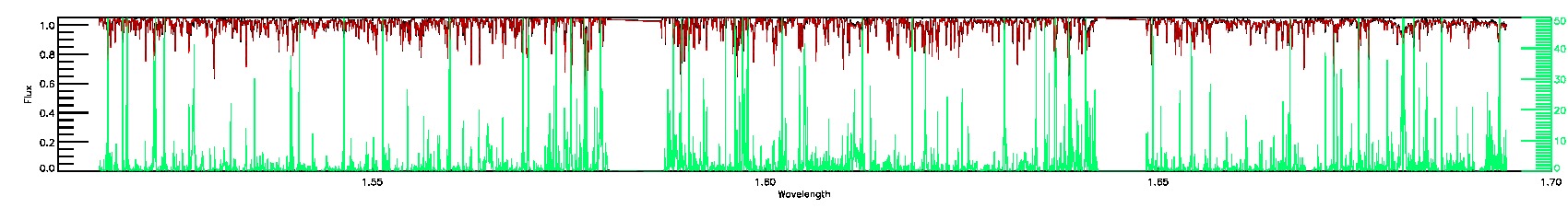
PERSIST_LOW,SUSPECT_RV_COMBINATION
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN
120-08-RV
| 7872. | +/- | 17. |
| -10000. | +/- | -NaN |
| 4.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.63 | +/- | 0. |
| 1.63 | +/- | -NaN |
| -1.84 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 92.60 | +/- | 0. |
| 1.97 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.89 |
| -9999.00 |
120-08-RV
| 4153. | +/- | 1. |
| 4243. | +/- | 71. |
| 2.02 | +/- | 0. |
| 1.79 | +/- | 0. |
| 1.50 | +/- | 0. |
| 1.50 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.03 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.41 |
| -9999.00 |

PERSIST_MED,PERSIST_LOW
120-08-RV
| 4417. | +/- | 2. |
| 4514. | +/- | 79. |
| 2.52 | +/- | 0. |
| 2.31 | +/- | 0. |
| 1.30 | +/- | 0. |
| 1.30 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -0.20 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.11 | +/- | 0. |
| 3.32 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.65 |
| -9999.00 |

PERSIST_MED,PERSIST_LOW
120-08-RV
| 4872. | +/- | 6. |
| 4934. | +/- | 92. |
| 2.75 | +/- | 0. |
| 2.42 | +/- | 0. |
| 1.58 | +/- | 0. |
| 1.58 | +/- | -NaN |
| -0.39 | +/- | 0. |
| -0.39 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.06 | +/- | 0. |
| 3.71 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.88 |
| -9999.00 |

PERSIST_LOW
120-08-RV
| 4402. | +/- | 2. |
| 4496. | +/- | 78. |
| 2.43 | +/- | 0. |
| 2.22 | +/- | 0. |
| 1.29 | +/- | 0. |
| 1.29 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.18 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.62 |
| -9999.00 |
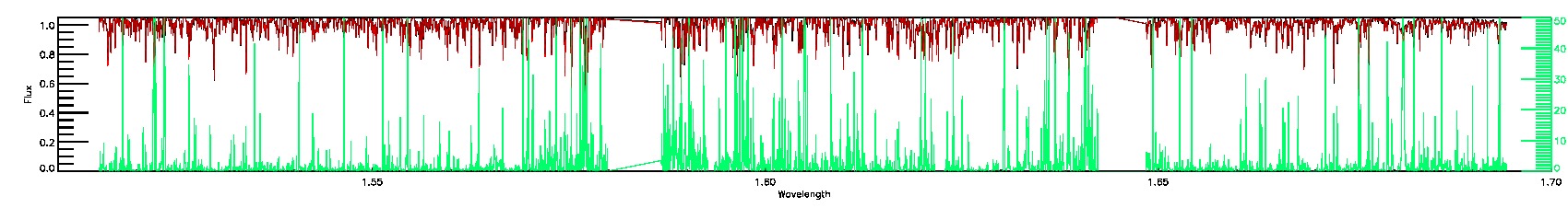
PERSIST_MED,PERSIST_LOW
120-08-RV
| 4848. | +/- | 5. |
| 4894. | +/- | 87. |
| 2.92 | +/- | 0. |
| 2.55 | +/- | 0. |
| 1.38 | +/- | 0. |
| 1.38 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.85 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.49 |
| -9999.00 |

120-08-RV
| 4068. | +/- | 1. |
| 4152. | +/- | 68. |
| 1.96 | +/- | 0. |
| 1.72 | +/- | 0. |
| 1.34 | +/- | 0. |
| 1.34 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.07 | +/- | 0. |
| 2.80 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.23 |
| -9999.00 |

PERSIST_LOW
120-08-RV
| 4423. | +/- | 2. |
| 4513. | +/- | 77. |
| 2.45 | +/- | 0. |
| 2.23 | +/- | 0. |
| 1.35 | +/- | 0. |
| 1.35 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.02 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.30 |
| -9999.00 |
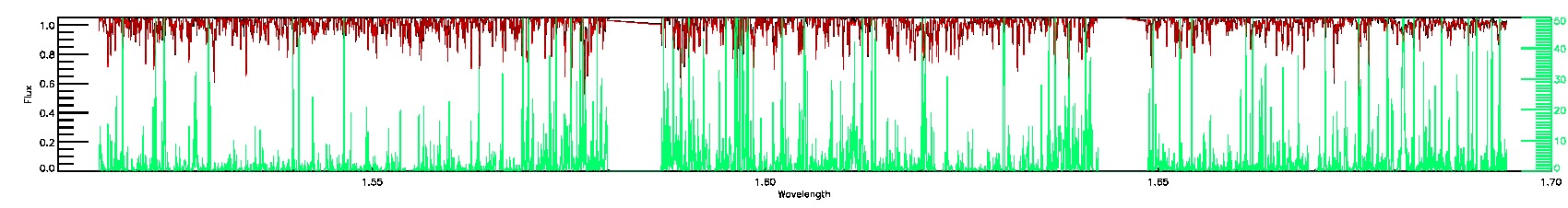
PERSIST_MED,PERSIST_LOW
120-08-RV
| 4840. | +/- | 5. |
| 4892. | +/- | 88. |
| 2.89 | +/- | 0. |
| 2.53 | +/- | 0. |
| 1.47 | +/- | 0. |
| 1.47 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.00 | +/- | 0. |
| 3.08 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.21 |
| -9999.00 |
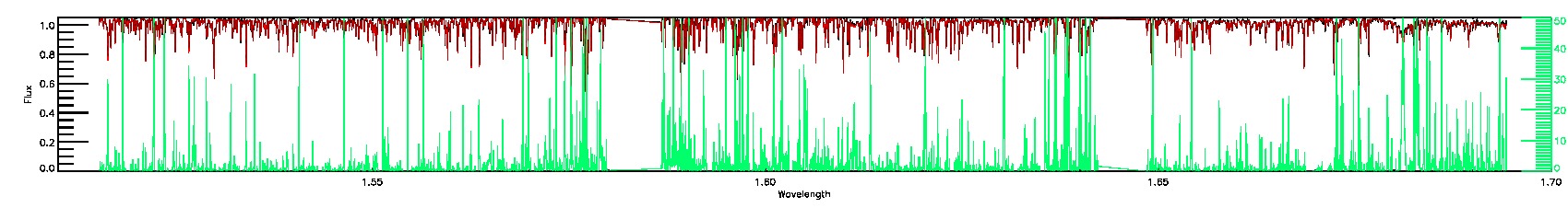
PERSIST_MED,PERSIST_LOW
120-08-RV
| 4572. | +/- | 3. |
| 4644. | +/- | 78. |
| 2.76 | +/- | 0. |
| 2.43 | +/- | 0. |
| 1.31 | +/- | 0. |
| 1.31 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.62 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.21 |
| -9999.00 |

120-08-RV
| 5078. | +/- | 7. |
| 5104. | +/- | 95. |
| 3.05 | +/- | 0. |
| 2.68 | +/- | 0. |
| 1.50 | +/- | 0. |
| 1.50 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| 0.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| 3.17 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.14 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.42 |
| -9999.00 |
120-08-RV
| 4722. | +/- | 4. |
| 4781. | +/- | 83. |
| 2.83 | +/- | 0. |
| 2.48 | +/- | 0. |
| 1.36 | +/- | 0. |
| 1.36 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.80 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.06 |
| -9999.00 |

120-08-RV
| 4875. | +/- | 5. |
| 4923. | +/- | 89. |
| 2.89 | +/- | 0. |
| 2.52 | +/- | 0. |
| 1.46 | +/- | 0. |
| 1.46 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 3.10 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.56 |
| -9999.00 |

PERSIST_MED,PERSIST_LOW
120-08-RV
| 4540. | +/- | 3. |
| 4637. | +/- | 82. |
| 2.44 | +/- | 0. |
| 2.24 | +/- | 0. |
| 1.57 | +/- | 0. |
| 1.57 | +/- | -NaN |
| -0.30 | +/- | 0. |
| -0.30 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.53 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.76 |
| -9999.00 |
120-08-RV
| 4701. | +/- | 4. |
| 4770. | +/- | 85. |
| 2.60 | +/- | 0. |
| 2.33 | +/- | 0. |
| 1.55 | +/- | 0. |
| 1.55 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 3.15 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.19 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
120-08-RV
| 4619. | +/- | 3. |
| 4679. | +/- | 78. |
| 2.87 | +/- | 0. |
| 2.51 | +/- | 0. |
| 1.23 | +/- | 0. |
| 1.23 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.43 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.55 |
| -9999.00 |

120-08-RV
| 4879. | +/- | 6. |
| 4941. | +/- | 92. |
| 2.69 | +/- | 0. |
| 2.38 | +/- | 0. |
| 1.72 | +/- | 0. |
| 1.72 | +/- | -NaN |
| -0.41 | +/- | 0. |
| -0.41 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.06 | +/- | 0. |
| 3.76 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.29 |
| -9999.00 |
PERSIST_MED,PERSIST_LOW
120-08-RV
| 4699. | +/- | 4. |
| 4763. | +/- | 83. |
| 2.73 | +/- | 0. |
| 2.41 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.43 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.90 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.03 |
| -9999.00 |

PERSIST_LOW,SUSPECT_RV_COMBINATION
120-08-RV
| 8674. | +/- | 39. |
| 8572. | +/- | 357. |
| 4.27 | +/- | 0. |
| 4.81 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.35 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.02 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.36 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| -2.02 |
| -9999.00 |
BRIGHT_NEIGHBOR,PERSIST_MED,PERSIST_LOW
120-08-RV
| 4088. | +/- | 1. |
| 4170. | +/- | 68. |
| 2.04 | +/- | 0. |
| 1.79 | +/- | 0. |
| 1.38 | +/- | 0. |
| 1.38 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.69 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.25 |
| -9999.00 |

PERSIST_MED,PERSIST_LOW
120-08-RV
| 3864. | +/- | 1. |
| 3972. | +/- | 69. |
| 1.12 | +/- | 0. |
| 1.03 | +/- | 0. |
| 1.98 | +/- | 0. |
| 1.98 | +/- | -NaN |
| -0.46 | +/- | 0. |
| -0.46 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.09 | +/- | 0. |
| 3.87 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.60 |
| -9999.00 |
PERSIST_MED,PERSIST_LOW
120-08-RV
| 4935. | +/- | 6. |
| 4990. | +/- | 94. |
| 2.80 | +/- | 0. |
| 2.45 | +/- | 0. |
| 1.62 | +/- | 0. |
| 1.62 | +/- | -NaN |
| -0.39 | +/- | 0. |
| -0.39 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.72 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.73 |
| -9999.00 |
120-08-RV
| 4250. | +/- | 2. |
| 4353. | +/- | 76. |
| 2.12 | +/- | 0. |
| 1.93 | +/- | 0. |
| 1.42 | +/- | 0. |
| 1.42 | +/- | -NaN |
| -0.34 | +/- | 0. |
| -0.34 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.09 | +/- | 0. |
| 3.61 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.50 |
| -9999.00 |

BRIGHT_NEIGHBOR,PERSIST_MED,PERSIST_LOW
120-08-RV
| 4629. | +/- | 4. |
| 4710. | +/- | 83. |
| 2.68 | +/- | 0. |
| 2.37 | +/- | 0. |
| 1.33 | +/- | 0. |
| 1.33 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.17 | +/- | 0. |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.14 | +/- | 0. |
| 3.27 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.18 |
| -9999.00 |
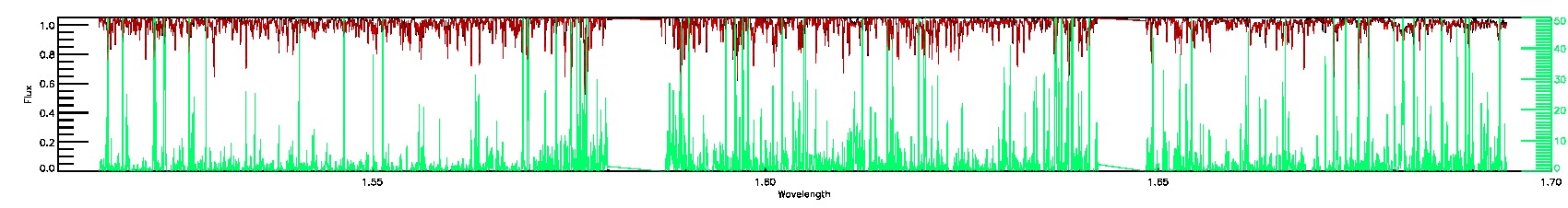
PERSIST_LOW
120-08-RV
| 4222. | +/- | 1. |
| 4311. | +/- | 73. |
| 2.05 | +/- | 0. |
| 1.82 | +/- | 0. |
| 1.53 | +/- | 0. |
| 1.53 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.01 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.02 |
| -9999.00 |
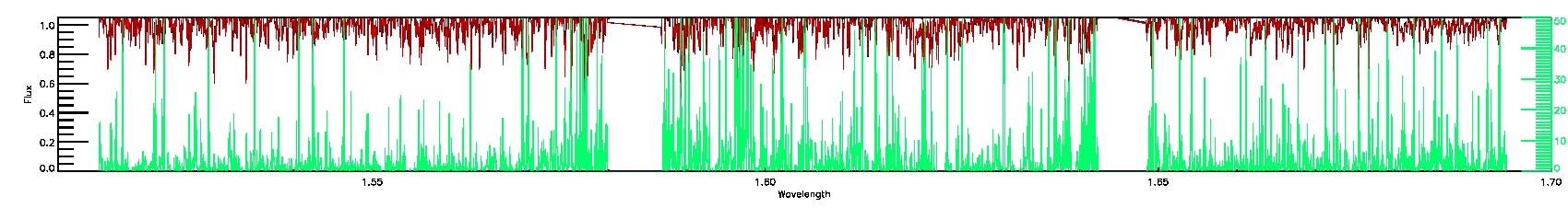
BRIGHT_NEIGHBOR,PERSIST_LOW
120-08-RV
| 4450. | +/- | 2. |
| 4524. | +/- | 75. |
| 2.69 | +/- | 0. |
| 2.38 | +/- | 0. |
| 1.28 | +/- | 0. |
| 1.28 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.44 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| 0.29 |
| -9999.00 |

120-08-RV
| 4606. | +/- | 4. |
| 4692. | +/- | 83. |
| 2.62 | +/- | 0. |
| 2.43 | +/- | 0. |
| 1.41 | +/- | 0. |
| 1.41 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.24 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.41 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.29 |
| -9999.00 |
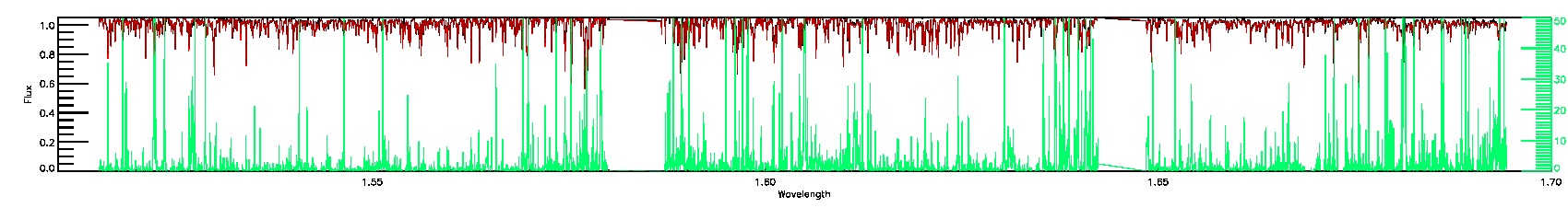
PERSIST_MED,PERSIST_LOW
120-08-RV
| 4531. | +/- | 2. |
| 4600. | +/- | 76. |
| 2.82 | +/- | 0. |
| 2.47 | +/- | 0. |
| 1.25 | +/- | 0. |
| 1.25 | +/- | -NaN |
| 0.37 | +/- | 0. |
| 0.37 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| 0.50 |
| -9999.00 |
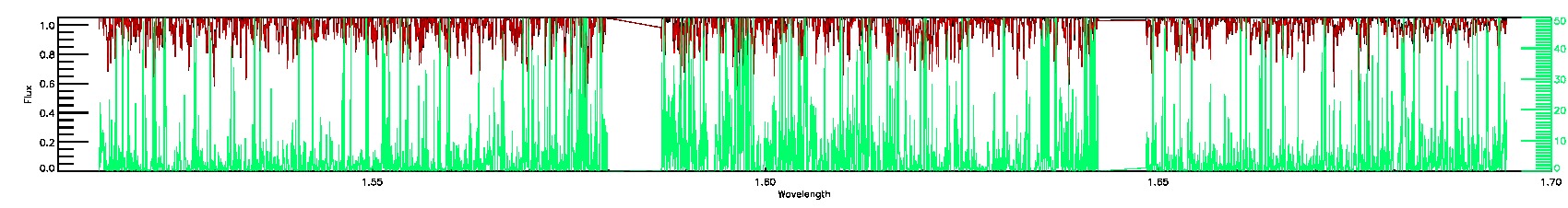
PERSIST_MED,PERSIST_LOW
120-08-RV
| 4915. | +/- | 6. |
| 4957. | +/- | 90. |
| 3.26 | +/- | 0. |
| 3.10 | +/- | 0. |
| 1.22 | +/- | 0. |
| 1.22 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 3.05 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.32 |
| -9999.00 |
PERSIST_LOW
120-08-RV
| 4087. | +/- | 1. |
| 4176. | +/- | 70. |
| 1.67 | +/- | 0. |
| 1.46 | +/- | 0. |
| 1.48 | +/- | 0. |
| 1.48 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.97 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.32 |
| -9999.00 |

120-08-RV
| 4836. | +/- | 5. |
| 4884. | +/- | 87. |
| 2.92 | +/- | 0. |
| 2.55 | +/- | 0. |
| 1.42 | +/- | 0. |
| 1.42 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.89 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.05 |
| -9999.00 |
BRIGHT_NEIGHBOR,PERSIST_LOW
120-08-RV
| 4313. | +/- | 2. |
| 4399. | +/- | 74. |
| 2.53 | +/- | 0. |
| 2.30 | +/- | 0. |
| 0.91 | +/- | 0. |
| 0.91 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.09 | +/- | 0. |
| 2.86 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.04 |
| -9999.00 |

120-08-RV
| 4805. | +/- | 4. |
| 4846. | +/- | 83. |
| 3.39 | +/- | 0. |
| 3.18 | +/- | 0. |
| 0.88 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.29 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.50 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.27 |
| -9999.00 |
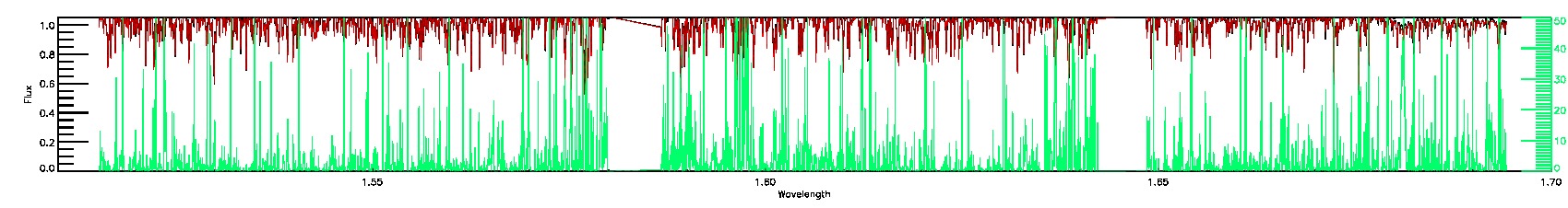
PERSIST_LOW
120-08-RV
| 4545. | +/- | 3. |
| 4616. | +/- | 77. |
| 2.74 | +/- | 0. |
| 2.41 | +/- | 0. |
| 1.28 | +/- | 0. |
| 1.28 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.29 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.49 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| 0.23 |
| -9999.00 |
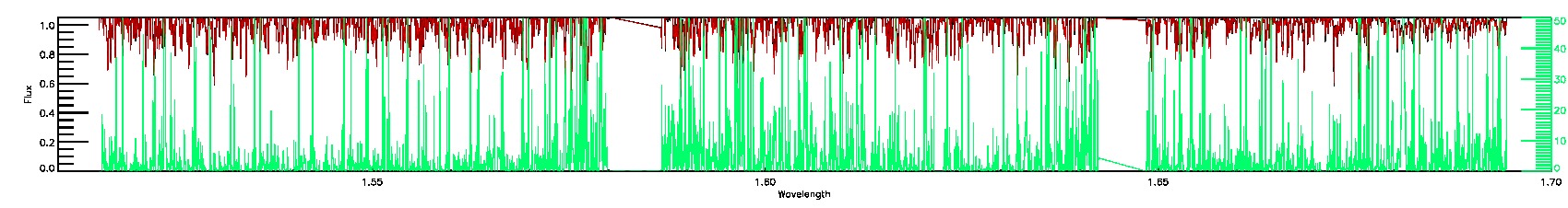
120-08-RV
| 5163. | +/- | 9. |
| 5181. | +/- | 97. |
| 3.22 | +/- | 0. |
| 2.86 | +/- | 0. |
| 0.94 | +/- | 0. |
| 0.94 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.03 | +/- | 0. |
| 3.23 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.16 |
| -9999.00 |
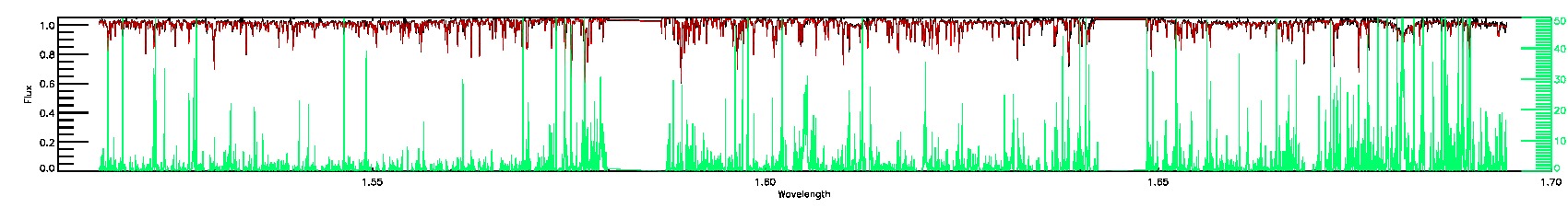
PERSIST_LOW
120-08-RV
| 4594. | +/- | 3. |
| 4667. | +/- | 80. |
| 2.65 | +/- | 0. |
| 2.36 | +/- | 0. |
| 1.40 | +/- | 0. |
| 1.40 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.80 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.25 |
| -9999.00 |

PERSIST_MED,PERSIST_LOW
120-08-RV
| 4804. | +/- | 5. |
| 4866. | +/- | 90. |
| 2.74 | +/- | 0. |
| 2.41 | +/- | 0. |
| 1.54 | +/- | 0. |
| 1.54 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -0.20 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.32 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -0.90 |
| -9999.00 |
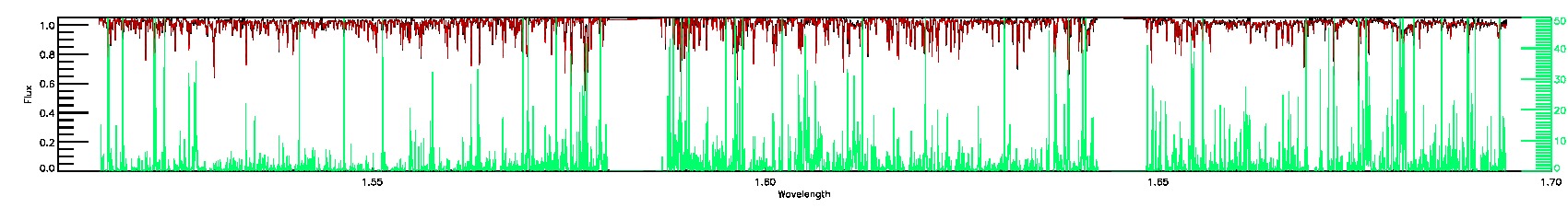
120-08-RV
| 4748. | +/- | 5. |
| 4807. | +/- | 85. |
| 3.09 | +/- | 0. |
| 2.90 | +/- | 0. |
| 1.18 | +/- | 0. |
| 1.18 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.95 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.16 |
| -9999.00 |

120-08-RV
| 4463. | +/- | 2. |
| 4554. | +/- | 78. |
| 2.48 | +/- | 0. |
| 2.25 | +/- | 0. |
| 1.39 | +/- | 0. |
| 1.39 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.06 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.16 |
| -9999.00 |
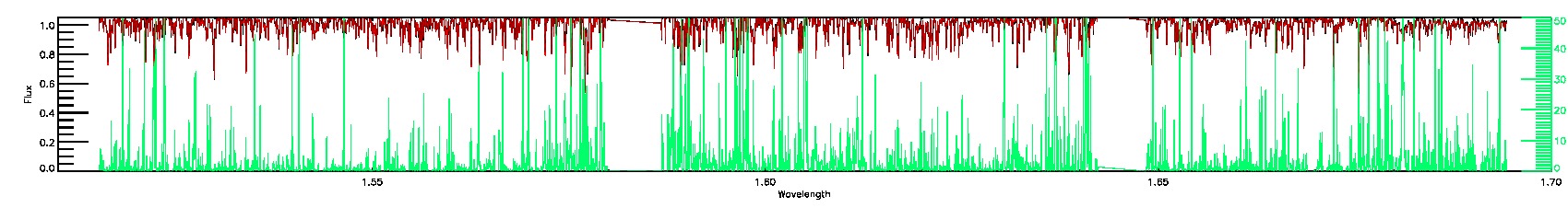
120-08-RV
| 4230. | +/- | 1. |
| 4334. | +/- | 76. |
| 1.79 | +/- | 0. |
| 1.61 | +/- | 0. |
| 1.69 | +/- | 0. |
| 1.69 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -0.35 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.63 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.66 |
| -9999.00 |

120-08-RV
| 5111. | +/- | 7. |
| 5133. | +/- | 95. |
| 3.16 | +/- | 0. |
| 2.79 | +/- | 0. |
| 1.53 | +/- | 0. |
| 1.53 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.01 | +/- | 0. |
| 3.15 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.36 |
| -9999.00 |

120-08-RV
| 4737. | +/- | 4. |
| 4795. | +/- | 84. |
| 2.77 | +/- | 0. |
| 2.44 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.43 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.85 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.82 |
| -9999.00 |

PERSIST_MED
120-08-RV
| 3973. | +/- | 1. |
| 4066. | +/- | 68. |
| 1.49 | +/- | 0. |
| 1.32 | +/- | 0. |
| 1.80 | +/- | 0. |
| 1.80 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.18 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.29 |
| -9999.00 |
120-08-RV
| 4476. | +/- | 3. |
| 4577. | +/- | 81. |
| 2.31 | +/- | 0. |
| 2.11 | +/- | 0. |
| 1.53 | +/- | 0. |
| 1.53 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -0.29 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.50 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -1.02 |
| -9999.00 |

120-08-RV
| 4370. | +/- | 2. |
| 4464. | +/- | 77. |
| 2.18 | +/- | 0. |
| 1.96 | +/- | 0. |
| 1.55 | +/- | 0. |
| 1.55 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.13 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.19 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.32 |
| -9999.00 |

LOW_SNR
120-08-RV
| 4804. | +/- | 14. |
| 4860. | +/- | 114. |
| 2.75 | +/- | 0. |
| 2.42 | +/- | 0. |
| 1.53 | +/- | 0. |
| 1.53 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 3.07 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
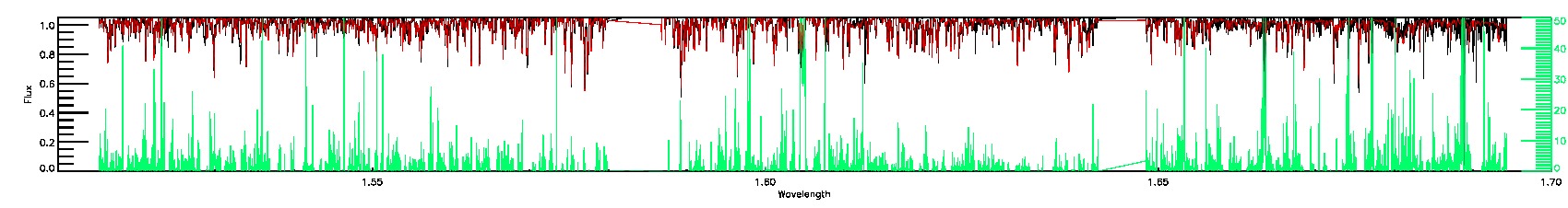
PERSIST_MED,PERSIST_LOW,SUSPECT_RV_COMBINATION
120-08-RV
| 8158. | +/- | 26. |
| 8047. | +/- | 298. |
| 4.42 | +/- | 0. |
| 4.87 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 35.49 | +/- | 0. |
| 1.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.31 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.29 |
| -9999.00 |
120-08-RV
| 4346. | +/- | 2. |
| 4432. | +/- | 75. |
| 2.22 | +/- | 0. |
| 1.97 | +/- | 0. |
| 1.51 | +/- | 0. |
| 1.51 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.12 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.29 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.86 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.08 |
| -9999.00 |
PERSIST_MED,PERSIST_LOW
120-08-RV
| 4291. | +/- | 2. |
| 4376. | +/- | 73. |
| 2.34 | +/- | 0. |
| 2.10 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.43 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.84 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.10 |
| -9999.00 |
120-08-RV
| 4808. | +/- | 5. |
| 4871. | +/- | 89. |
| 2.77 | +/- | 0. |
| 2.43 | +/- | 0. |
| 1.56 | +/- | 0. |
| 1.56 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -0.26 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.44 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.73 |
| -9999.00 |

BRIGHT_NEIGHBOR,PERSIST_LOW
120-08-RV
| 4712. | +/- | 4. |
| 4779. | +/- | 85. |
| 2.69 | +/- | 0. |
| 2.38 | +/- | 0. |
| 1.54 | +/- | 0. |
| 1.54 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.11 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.34 |
| -9999.00 |

120-08-RV
| 3935. | +/- | 1. |
| 4024. | +/- | 67. |
| 1.52 | +/- | 0. |
| 1.33 | +/- | 0. |
| 1.55 | +/- | 0. |
| 1.55 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.05 | +/- | 0. |
| 2.99 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.15 |
| -9999.00 |
PERSIST_LOW
120-08-RV
| 4016. | +/- | 1. |
| 4105. | +/- | 68. |
| 1.72 | +/- | 0. |
| 1.50 | +/- | 0. |
| 1.67 | +/- | 0. |
| 1.67 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.97 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.25 |
| -9999.00 |

120-08-RV
| 4485. | +/- | 2. |
| 4561. | +/- | 76. |
| 2.65 | +/- | 0. |
| 2.36 | +/- | 0. |
| 1.30 | +/- | 0. |
| 1.30 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.29 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.49 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| 0.30 |
| -9999.00 |
PERSIST_MED
120-08-RV
| 4486. | +/- | 3. |
| 4582. | +/- | 80. |
| 2.38 | +/- | 0. |
| 2.17 | +/- | 0. |
| 1.50 | +/- | 0. |
| 1.50 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -0.18 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.28 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.55 |
| -9999.00 |
PERSIST_MED
120-08-RV
| 4770. | +/- | 4. |
| 4823. | +/- | 84. |
| 2.88 | +/- | 0. |
| 2.52 | +/- | 0. |
| 1.39 | +/- | 0. |
| 1.39 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.29 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.82 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.10 |
| -9999.00 |

PERSIST_MED
120-08-RV
| 4796. | +/- | 5. |
| 4854. | +/- | 87. |
| 2.99 | +/- | 0. |
| 2.81 | +/- | 0. |
| 1.16 | +/- | 0. |
| 1.16 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 3.13 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.22 |
| -9999.00 |

120-08-RV
| 4933. | +/- | 6. |
| 4967. | +/- | 88. |
| 3.36 | +/- | 0. |
| 3.18 | +/- | 0. |
| 1.19 | +/- | 0. |
| 1.19 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.79 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.19 |
| -9999.00 |

120-08-RV
| 3884. | +/- | 1. |
| 3996. | +/- | 70. |
| 1.04 | +/- | 0. |
| 0.96 | +/- | 0. |
| 1.84 | +/- | 0. |
| 1.84 | +/- | -NaN |
| -0.54 | +/- | 0. |
| -0.54 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.09 | +/- | 0. |
| 4.06 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.86 |
| -9999.00 |
SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD
120-08-RV
| 8234. | +/- | 25. |
| -10000. | +/- | -NaN |
| 4.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -0.90 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 92.11 | +/- | 0. |
| 1.96 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.20 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.16 |
| -9999.00 |
SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES
120-08-RV
| 8312. | +/- | 28. |
| 8169. | +/- | 286. |
| 4.16 | +/- | 0. |
| 4.31 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -1.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 84.76 | +/- | 0. |
| 1.93 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.68 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -1.38 |
| -9999.00 |
120-08-RV
| 4658. | +/- | 4. |
| 4727. | +/- | 82. |
| 2.72 | +/- | 0. |
| 2.40 | +/- | 0. |
| 1.41 | +/- | 0. |
| 1.41 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.03 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.90 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.08 |
| -9999.00 |

120-08-RV
| 4624. | +/- | 4. |
| 4699. | +/- | 82. |
| 2.85 | +/- | 0. |
| 2.65 | +/- | 0. |
| 1.28 | +/- | 0. |
| 1.28 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.01 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |

PERSIST_LOW
120-08-RV
| 4608. | +/- | 3. |
| 4682. | +/- | 81. |
| 2.94 | +/- | 0. |
| 2.73 | +/- | 0. |
| 1.15 | +/- | 0. |
| 1.15 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.85 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.21 |
| -9999.00 |
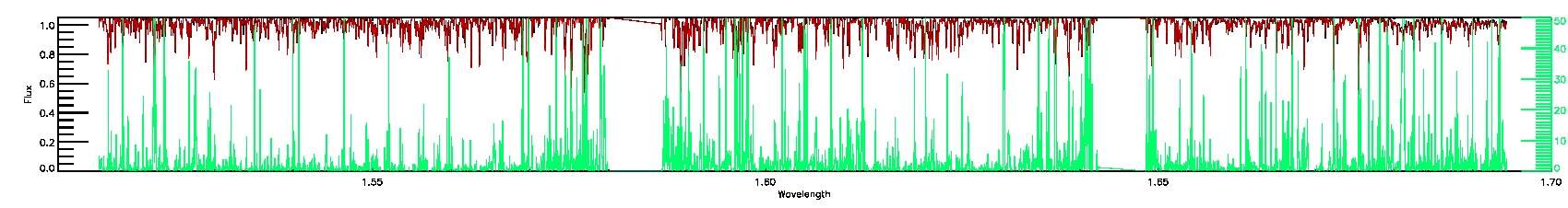
120-08-RV
| 4220. | +/- | 1. |
| 4306. | +/- | 72. |
| 2.04 | +/- | 0. |
| 1.80 | +/- | 0. |
| 1.52 | +/- | 0. |
| 1.52 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.88 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.01 |
| -9999.00 |
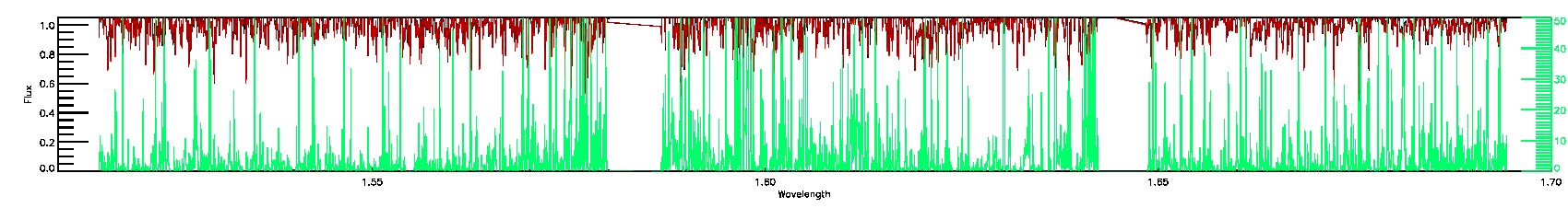
120-08-RV
| 4349. | +/- | 3. |
| 4454. | +/- | 83. |
| 2.06 | +/- | 0. |
| 1.88 | +/- | 0. |
| 1.51 | +/- | 0. |
| 1.51 | +/- | -NaN |
| -0.38 | +/- | 0. |
| -0.38 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.08 | +/- | 0. |
| 3.70 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.52 |
| -9999.00 |
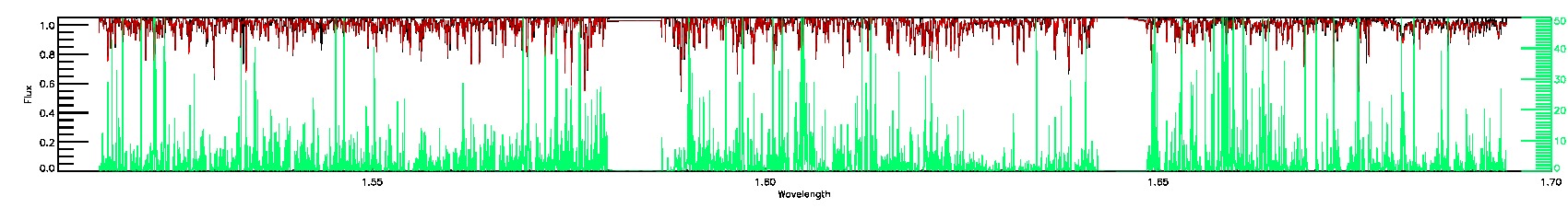
PERSIST_MED
120-08-RV
| 4935. | +/- | 6. |
| 4974. | +/- | 90. |
| 3.05 | +/- | 0. |
| 2.67 | +/- | 0. |
| 1.37 | +/- | 0. |
| 1.37 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.99 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.14 |
| -9999.00 |

PERSIST_LOW
120-08-RV
| 4291. | +/- | 2. |
| 4383. | +/- | 75. |
| 2.20 | +/- | 0. |
| 1.98 | +/- | 0. |
| 1.38 | +/- | 0. |
| 1.38 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.14 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.05 |
| -9999.00 |
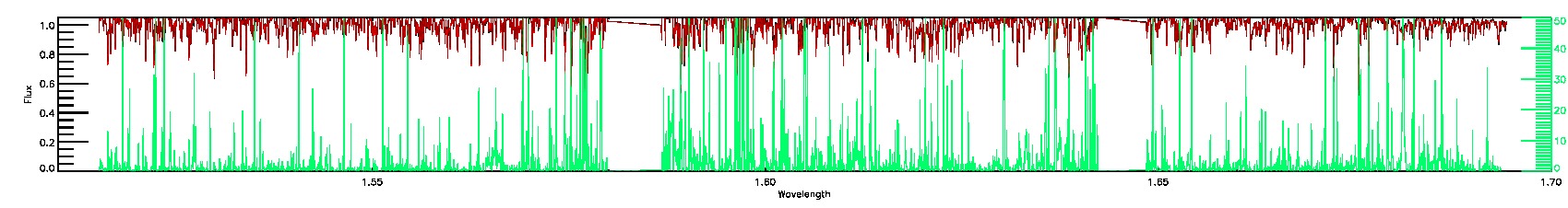
PERSIST_LOW
120-08-RV
| 4517. | +/- | 3. |
| 4610. | +/- | 80. |
| 2.40 | +/- | 0. |
| 2.18 | +/- | 0. |
| 1.53 | +/- | 0. |
| 1.53 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -0.14 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.00 | +/- | 0. |
| 3.20 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.46 |
| -9999.00 |

PERSIST_LOW
120-08-RV
| 4482. | +/- | 2. |
| 4559. | +/- | 76. |
| 2.59 | +/- | 0. |
| 2.33 | +/- | 0. |
| 1.39 | +/- | 0. |
| 1.39 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.54 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| 0.28 |
| -9999.00 |
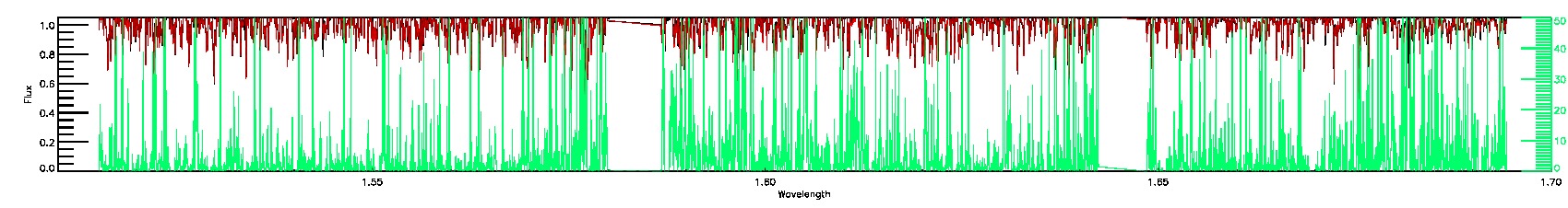
120-08-RV
| 4055. | +/- | 1. |
| 4164. | +/- | 73. |
| 1.52 | +/- | 0. |
| 1.39 | +/- | 0. |
| 1.80 | +/- | 0. |
| 1.80 | +/- | -NaN |
| -0.48 | +/- | 0. |
| -0.48 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.08 | +/- | 0. |
| 3.91 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.55 |
| -9999.00 |

SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_WARN,COLORTE_WARN
120-08-RV
| 8158. | +/- | 22. |
| 8054. | +/- | 304. |
| 4.44 | +/- | 0. |
| 4.96 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 23.37 | +/- | 0. |
| 1.37 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.46 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -2.14 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -2.14 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -2.13 |
| -9999.00 |
PERSIST_LOW
120-08-RV
| 4751. | +/- | 4. |
| 4804. | +/- | 83. |
| 2.89 | +/- | 0. |
| 2.53 | +/- | 0. |
| 1.37 | +/- | 0. |
| 1.37 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.71 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.56 |
| -9999.00 |
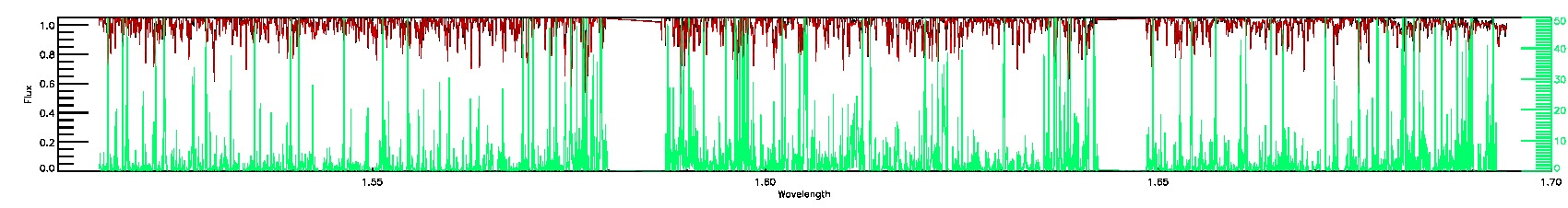
120-08-RV
| 4387. | +/- | 2. |
| 4478. | +/- | 77. |
| 2.21 | +/- | 0. |
| 1.98 | +/- | 0. |
| 1.56 | +/- | 0. |
| 1.56 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.07 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.50 |
| -9999.00 |
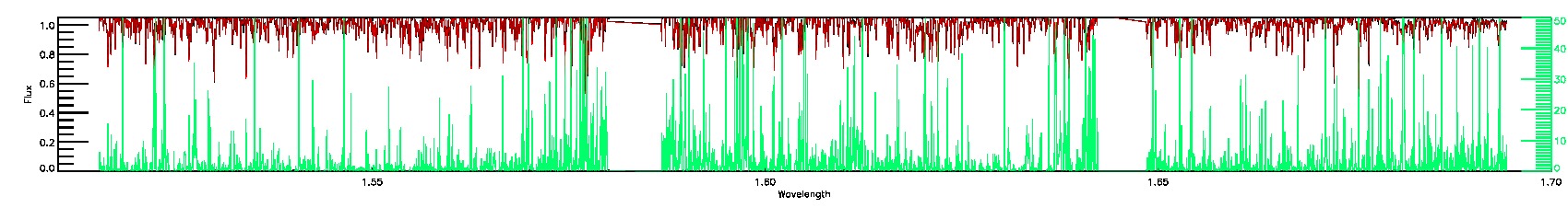
PERSIST_LOW
120-08-RV
| 4845. | +/- | 6. |
| 4895. | +/- | 88. |
| 2.99 | +/- | 0. |
| 2.61 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.43 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.14 | +/- | 0. |
| -0.14 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 3.03 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.14 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.31 |
| -9999.00 |
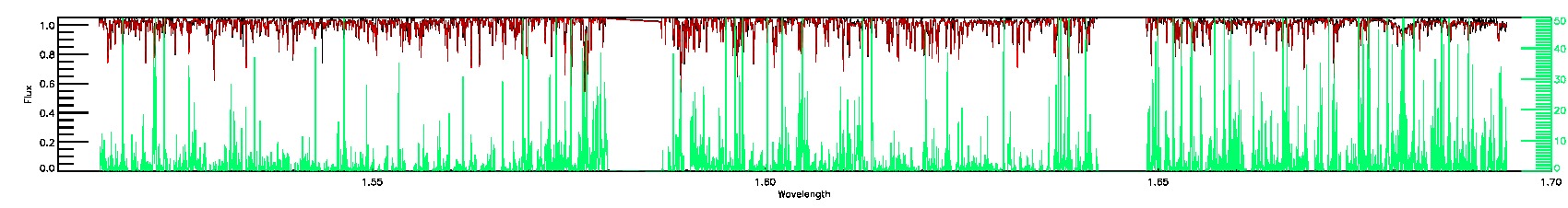
PERSIST_LOW
120-08-RV
| 4388. | +/- | 2. |
| 4480. | +/- | 77. |
| 2.38 | +/- | 0. |
| 2.16 | +/- | 0. |
| 1.40 | +/- | 0. |
| 1.40 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.10 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.12 |
| -9999.00 |
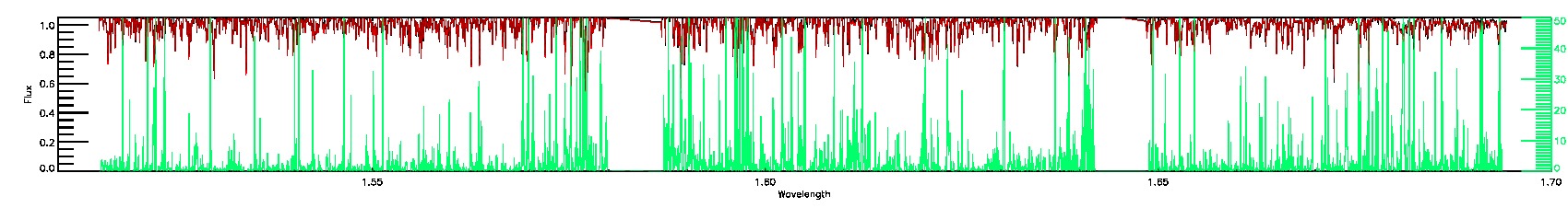
120-08-RV
| 4457. | +/- | 3. |
| 4556. | +/- | 80. |
| 2.25 | +/- | 0. |
| 2.05 | +/- | 0. |
| 1.61 | +/- | 0. |
| 1.61 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -0.26 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.44 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.88 |
| -9999.00 |
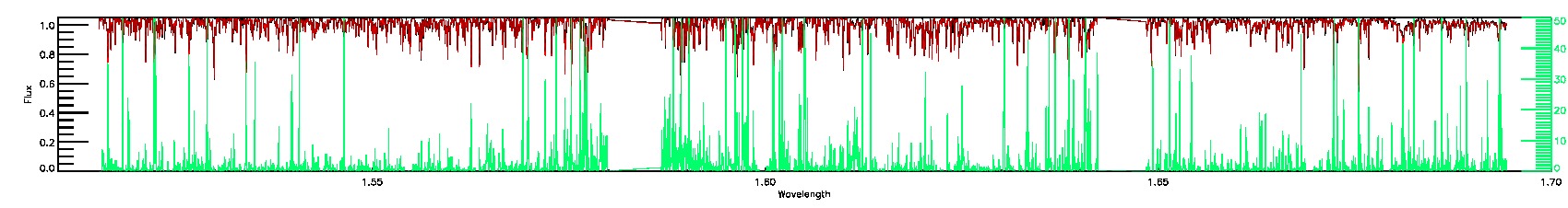
PERSIST_LOW
120-08-RV
| 4167. | +/- | 1. |
| 4262. | +/- | 73. |
| 1.83 | +/- | 0. |
| 1.62 | +/- | 0. |
| 1.64 | +/- | 0. |
| 1.64 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.23 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.56 |
| -9999.00 |

120-08-RV
| 4147. | +/- | 1. |
| 4247. | +/- | 73. |
| 1.87 | +/- | 0. |
| 1.68 | +/- | 0. |
| 1.59 | +/- | 0. |
| 1.59 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -0.29 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.41 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.15 | +/- | 0. |
| 3.50 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |

120-08-RV
| 4733. | +/- | 4. |
| 4798. | +/- | 85. |
| 2.75 | +/- | 0. |
| 2.42 | +/- | 0. |
| 1.50 | +/- | 0. |
| 1.50 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.09 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.14 |
| -9999.00 |
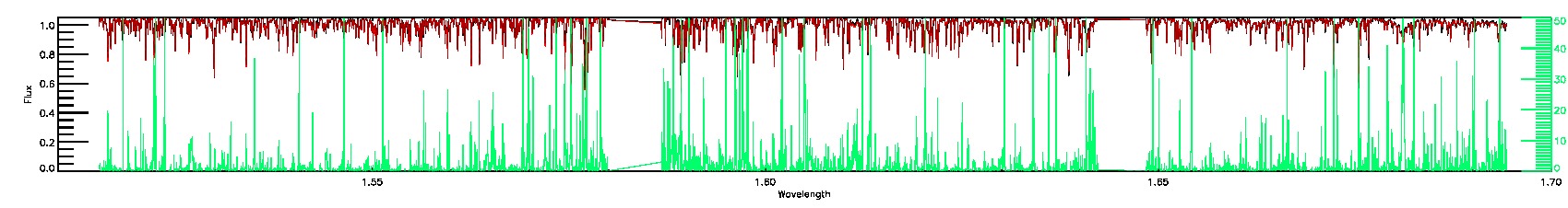
120-08-RV
| 4558. | +/- | 3. |
| 4646. | +/- | 81. |
| 2.66 | +/- | 0. |
| 2.45 | +/- | 0. |
| 1.38 | +/- | 0. |
| 1.38 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.21 | +/- | 0. |
| -0.21 | +/- | -NaN |
| 0.45 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.16 | +/- | 0. |
| 3.22 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.22 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.54 |
| -9999.00 |

PERSIST_LOW
120-08-RV
| 4892. | +/- | 6. |
| 4946. | +/- | 91. |
| 2.81 | +/- | 0. |
| 2.46 | +/- | 0. |
| 1.50 | +/- | 0. |
| 1.50 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -0.25 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.42 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.37 |
| -9999.00 |

120-08-RV
| 4737. | +/- | 5. |
| 4807. | +/- | 87. |
| 2.80 | +/- | 0. |
| 2.62 | +/- | 0. |
| 1.33 | +/- | 0. |
| 1.33 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.36 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.45 |
| -9999.00 |
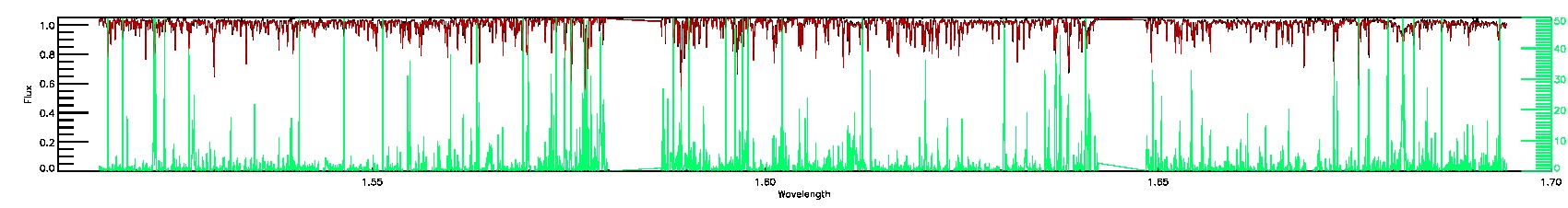
PERSIST_LOW,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
120-08-RV
| 8250. | +/- | 23. |
| 8144. | +/- | 311. |
| 4.15 | +/- | 0. |
| 4.65 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.69 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.43 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| -1.93 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -1.93 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -1.93 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -2.29 |
| -9999.00 |
120-08-RV
| 5104. | +/- | 8. |
| 5133. | +/- | 97. |
| 3.49 | +/- | 0. |
| 3.22 | +/- | 0. |
| 1.30 | +/- | 0. |
| 1.30 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -0.25 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.42 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.08 |
| -9999.00 |

PERSIST_LOW
120-08-RV
| 3998. | +/- | 1. |
| 4093. | +/- | 69. |
| 1.45 | +/- | 0. |
| 1.29 | +/- | 0. |
| 1.71 | +/- | 0. |
| 1.71 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.17 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.27 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.87 |
| -9999.00 |

PERSIST_LOW
120-08-RV
| 3746. | +/- | 1. |
| 3856. | +/- | 67. |
| 0.75 | +/- | 0. |
| 0.66 | +/- | 0. |
| 2.21 | +/- | 0. |
| 2.21 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -0.49 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.94 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.34 |
| -9999.00 |
PERSIST_LOW,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD
STAR_WARN,COLORTE_WARN
120-08-RV
| 7999. | +/- | 12. |
| -10000. | +/- | -NaN |
| 4.71 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.12 | +/- | 2. |
| 1.12 | +/- | -NaN |
| -2.30 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.64 | +/- | 1. |
| 0.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.45 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -1.98 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| -2.14 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -2.14 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.14 |
| -9999.00 |
120-08-RV
| 4236. | +/- | 1. |
| 4321. | +/- | 72. |
| 2.34 | +/- | 0. |
| 2.09 | +/- | 0. |
| 1.32 | +/- | 0. |
| 1.32 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.08 | +/- | 0. |
| 2.84 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.06 |
| -9999.00 |

PERSIST_LOW,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
120-08-RV
| 8396. | +/- | 27. |
| 8282. | +/- | 317. |
| 4.30 | +/- | 0. |
| 4.72 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.00 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.23 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -2.03 |
| -9999.00 |
PERSIST_LOW
120-08-RV
| 4716. | +/- | 4. |
| 4786. | +/- | 86. |
| 2.64 | +/- | 0. |
| 2.35 | +/- | 0. |
| 1.55 | +/- | 0. |
| 1.55 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.17 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.26 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.55 |
| -9999.00 |

120-08-RV
| 4879. | +/- | 6. |
| 4930. | +/- | 90. |
| 2.80 | +/- | 0. |
| 2.46 | +/- | 0. |
| 1.50 | +/- | 0. |
| 1.50 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -0.14 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.21 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.16 |
| -9999.00 |
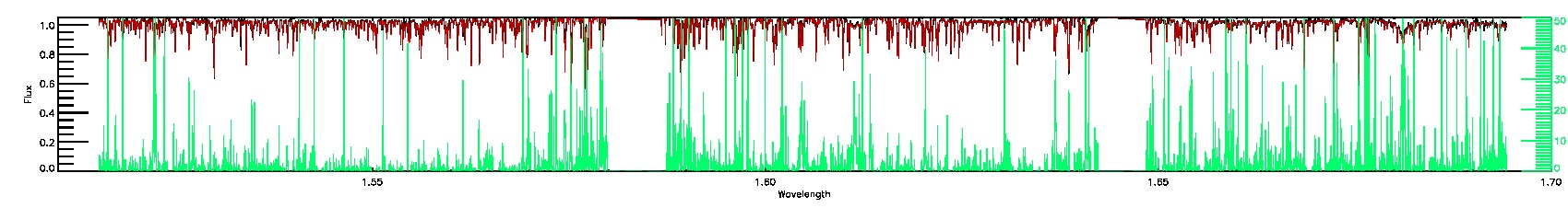
120-08-RV
| 3927. | +/- | 1. |
| 4041. | +/- | 71. |
| 1.14 | +/- | 0. |
| 1.06 | +/- | 0. |
| 2.08 | +/- | 0. |
| 2.08 | +/- | -NaN |
| -0.60 | +/- | 0. |
| -0.60 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.12 | +/- | -NaN |
| 0.43 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.07 | +/- | 0. |
| 4.20 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -1.10 |
| -9999.00 |
120-08-RV
| 4685. | +/- | 4. |
| 4748. | +/- | 82. |
| 2.76 | +/- | 0. |
| 2.43 | +/- | 0. |
| 1.37 | +/- | 0. |
| 1.37 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.80 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.13 |
| -9999.00 |

120-08-RV
| 4198. | +/- | 1. |
| 4311. | +/- | 77. |
| 1.80 | +/- | 0. |
| 1.65 | +/- | 0. |
| 1.57 | +/- | 0. |
| 1.57 | +/- | -NaN |
| -0.58 | +/- | 0. |
| -0.58 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.18 | +/- | 0. |
| 4.16 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.83 |
| -9999.00 |
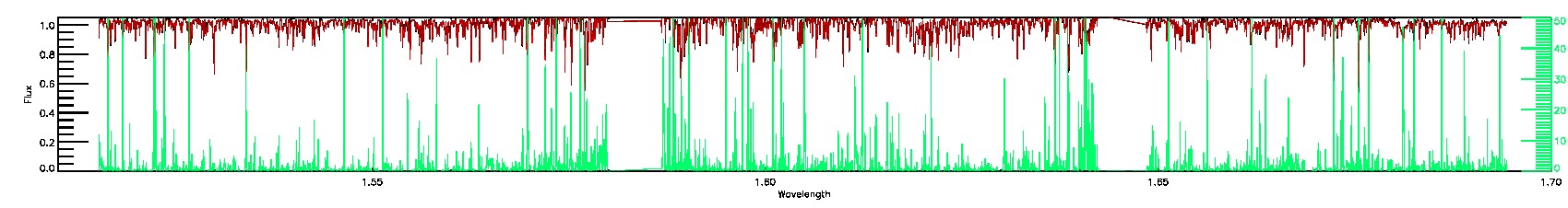
PERSIST_LOW
120-08-RV
| 4797. | +/- | 5. |
| 4855. | +/- | 87. |
| 3.15 | +/- | 0. |
| 2.98 | +/- | 0. |
| 1.32 | +/- | 0. |
| 1.32 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.11 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.50 |
| -9999.00 |
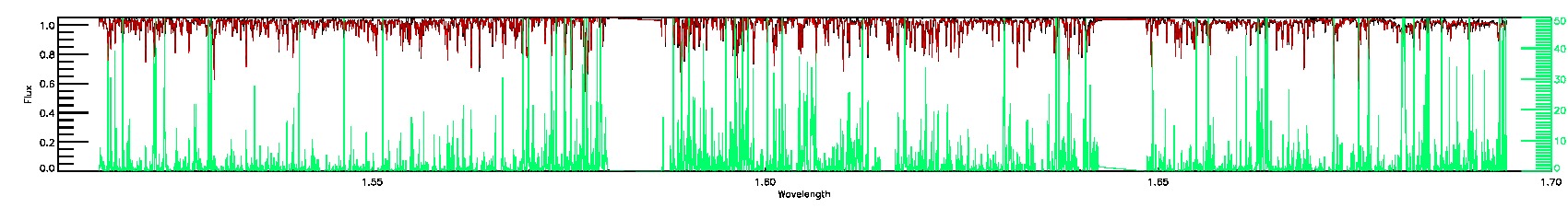
SUSPECT_RV_COMBINATION
120-08-RV
| 8301. | +/- | 32. |
| 8203. | +/- | 322. |
| 4.24 | +/- | 0. |
| 4.81 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 46.17 | +/- | 0. |
| 1.66 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.44 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -2.11 |
| -9999.00 |
PERSIST_LOW
120-08-RV
| 4156. | +/- | 1. |
| 4232. | +/- | 69. |
| 2.24 | +/- | 0. |
| 1.96 | +/- | 0. |
| 1.35 | +/- | 0. |
| 1.35 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.29 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.50 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.55 |
| -9999.00 |
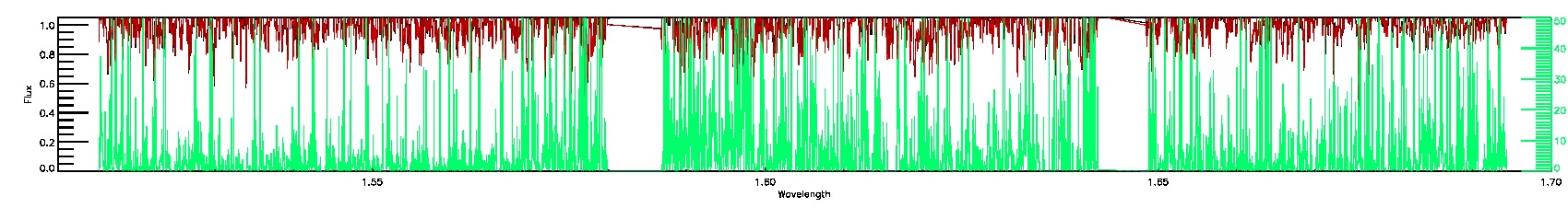
SUSPECT_RV_COMBINATION
STAR_BAD
STAR_WARN,COLORTE_WARN
120-08-RV
| 8223. | +/- | 26. |
| -10000. | +/- | -NaN |
| 4.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -1.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.04 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.38 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -1.25 |
| -9999.00 |
PERSIST_LOW
120-08-RV
| 4710. | +/- | 5. |
| 4789. | +/- | 87. |
| 2.68 | +/- | 0. |
| 2.51 | +/- | 0. |
| 1.45 | +/- | 0. |
| 1.45 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -0.35 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.64 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.32 |
| -9999.00 |
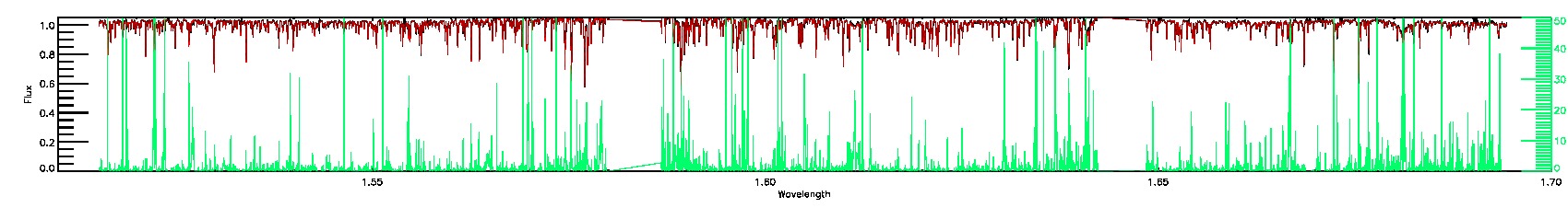
PERSIST_LOW
120-08-RV
| 3820. | +/- | 1. |
| 3929. | +/- | 68. |
| 1.01 | +/- | 0. |
| 0.93 | +/- | 0. |
| 1.92 | +/- | 0. |
| 1.92 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -0.49 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.09 | +/- | 0. |
| 3.93 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.78 |
| -9999.00 |
PERSIST_LOW
120-08-RV
| 4541. | +/- | 3. |
| 4630. | +/- | 81. |
| 2.44 | +/- | 0. |
| 2.22 | +/- | 0. |
| 1.51 | +/- | 0. |
| 1.51 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.00 | +/- | 0. |
| 3.17 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.59 |
| -9999.00 |

120-08-RV
| 4531. | +/- | 3. |
| 4639. | +/- | 85. |
| 2.39 | +/- | 0. |
| 2.23 | +/- | 0. |
| 1.42 | +/- | 0. |
| 1.42 | +/- | -NaN |
| -0.54 | +/- | 0. |
| -0.54 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.10 | +/- | 0. |
| 4.07 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.88 |
| -9999.00 |

BRIGHT_NEIGHBOR
120-08-RV
| 4716. | +/- | 4. |
| 4793. | +/- | 87. |
| 2.61 | +/- | 0. |
| 2.33 | +/- | 0. |
| 1.48 | +/- | 0. |
| 1.48 | +/- | -NaN |
| -0.32 | +/- | 0. |
| -0.31 | +/- | 0. |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.23 | +/- | 0. |
| 3.56 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.24 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.45 |
| -9999.00 |

120-08-RV
| 4494. | +/- | 3. |
| 4592. | +/- | 81. |
| 2.49 | +/- | 0. |
| 2.30 | +/- | 0. |
| 1.39 | +/- | 0. |
| 1.39 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -0.23 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.38 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.53 |
| -9999.00 |
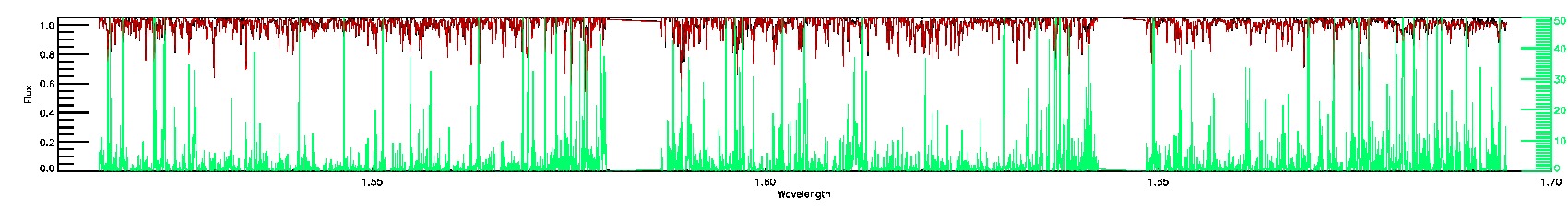
PERSIST_LOW
120-08-RV
| 4173. | +/- | 1. |
| 4284. | +/- | 76. |
| 1.70 | +/- | 0. |
| 1.55 | +/- | 0. |
| 1.82 | +/- | 0. |
| 1.82 | +/- | -NaN |
| -0.53 | +/- | 0. |
| -0.53 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.10 | +/- | 0. |
| 4.03 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.69 |
| -9999.00 |

120-08-RV
| 4685. | +/- | 4. |
| 4759. | +/- | 85. |
| 2.75 | +/- | 0. |
| 2.55 | +/- | 0. |
| 1.36 | +/- | 0. |
| 1.36 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -0.16 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 3.25 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.82 |
| -9999.00 |

120-08-RV
| 4740. | +/- | 5. |
| 4816. | +/- | 88. |
| 2.73 | +/- | 0. |
| 2.56 | +/- | 0. |
| 1.45 | +/- | 0. |
| 1.45 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -0.36 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.06 | +/- | 0. |
| 3.65 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.53 |
| -9999.00 |
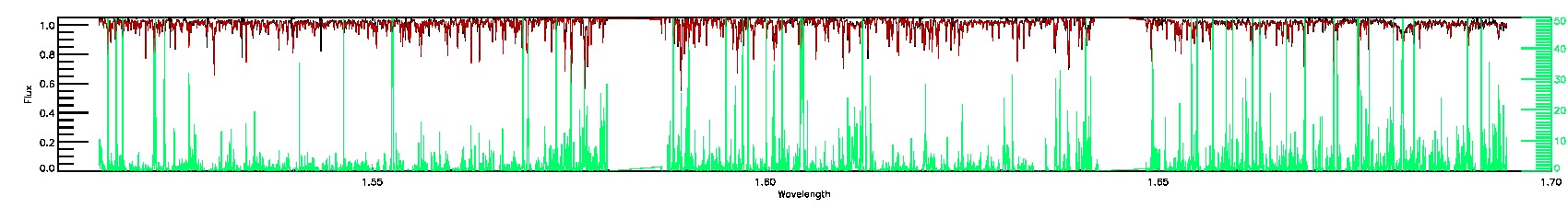
PERSIST_LOW
120-08-RV
| 5131. | +/- | 8. |
| 5147. | +/- | 95. |
| 3.33 | +/- | 0. |
| 3.00 | +/- | 0. |
| 1.13 | +/- | 0. |
| 1.13 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.41 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.03 | +/- | 0. |
| 3.00 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.10 |
| -9999.00 |

STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN
120-08-RV
| 4793. | +/- | 5. |
| -10000. | +/- | -NaN |
| 2.64 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.65 | +/- | 0. |
| 1.65 | +/- | -NaN |
| -0.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.70 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.95 |
| -9999.00 |

PERSIST_LOW,SUSPECT_RV_COMBINATION
STAR_BAD
120-08-RV
| 7999. | +/- | 15. |
| -10000. | +/- | -NaN |
| 4.31 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.50 | +/- | 0. |
| 0.50 | +/- | -NaN |
| -1.41 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 91.18 | +/- | 0. |
| 1.96 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| 0.60 |
| -9999.00 |
PERSIST_LOW
120-08-RV
| 5080. | +/- | 7. |
| 5106. | +/- | 95. |
| 3.05 | +/- | 0. |
| 2.68 | +/- | 0. |
| 1.62 | +/- | 0. |
| 1.62 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.16 | +/- | 0. |
| -0.16 | +/- | -NaN |
| 0.37 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| 3.16 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.17 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.07 |
| -9999.00 |

PERSIST_LOW,SUSPECT_RV_COMBINATION
STAR_BAD
120-08-RV
| 7999. | +/- | 10. |
| -10000. | +/- | -NaN |
| 4.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.28 | +/- | 1. |
| 1.28 | +/- | -NaN |
| -2.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.94 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.49 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -1.93 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -2.24 |
| -9999.00 |
BRIGHT_NEIGHBOR
120-08-RV
| 4518. | +/- | 3. |
| 4603. | +/- | 79. |
| 2.71 | +/- | 0. |
| 2.49 | +/- | 0. |
| 1.32 | +/- | 0. |
| 1.32 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.90 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.22 |
| -9999.00 |

PERSIST_LOW
120-08-RV
| 4557. | +/- | 3. |
| 4648. | +/- | 82. |
| 2.52 | +/- | 0. |
| 2.32 | +/- | 0. |
| 1.45 | +/- | 0. |
| 1.45 | +/- | -NaN |
| -0.21 | +/- | 0. |
| -0.21 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.34 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.16 |
| -9999.00 |
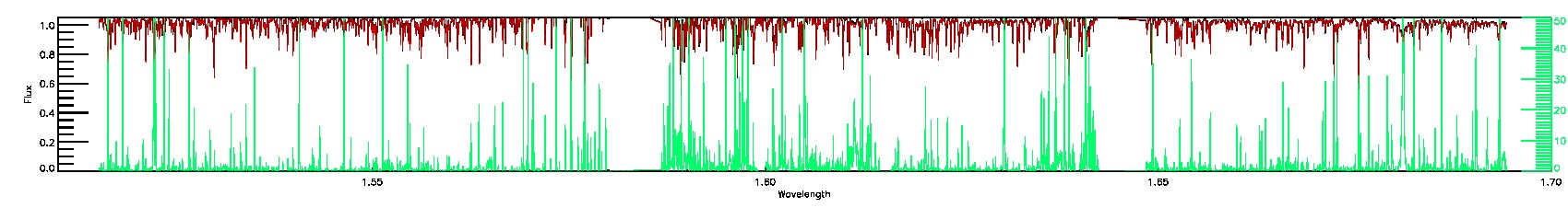
PERSIST_LOW
120-08-RV
| 4860. | +/- | 5. |
| 4908. | +/- | 88. |
| 2.85 | +/- | 0. |
| 2.50 | +/- | 0. |
| 1.49 | +/- | 0. |
| 1.49 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 3.01 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.23 |
| -9999.00 |

120-08-RV
| 4692. | +/- | 4. |
| 4755. | +/- | 83. |
| 2.81 | +/- | 0. |
| 2.46 | +/- | 0. |
| 1.31 | +/- | 0. |
| 1.31 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.84 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.46 |
| -9999.00 |
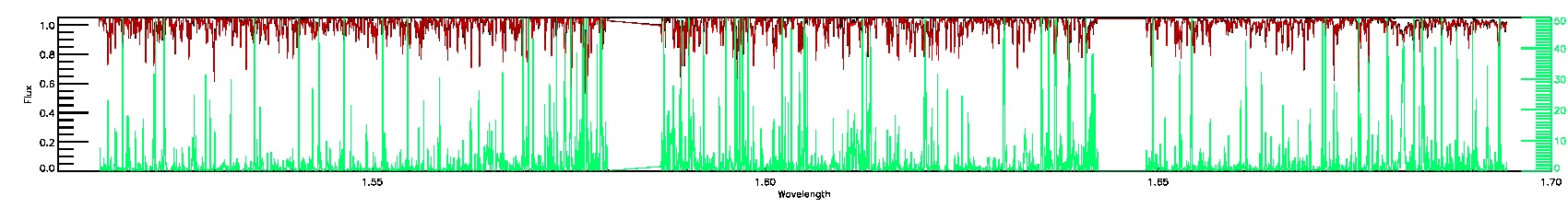
120-08-RV
| 4722. | +/- | 4. |
| 4787. | +/- | 85. |
| 2.74 | +/- | 0. |
| 2.41 | +/- | 0. |
| 1.39 | +/- | 0. |
| 1.39 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.05 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.14 |
| -9999.00 |

120-08-RV
| 4424. | +/- | 2. |
| 4513. | +/- | 77. |
| 2.49 | +/- | 0. |
| 2.26 | +/- | 0. |
| 1.37 | +/- | 0. |
| 1.37 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.99 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.25 |
| -9999.00 |

PERSIST_LOW,SUSPECT_RV_COMBINATION
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN
120-08-RV
| 7227. | +/- | 14. |
| -10000. | +/- | -NaN |
| 3.80 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.71 | +/- | 0. |
| 0.71 | +/- | -NaN |
| -1.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.76 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.17 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.55 |
| -9999.00 |
120-08-RV
| 4785. | +/- | 5. |
| 4843. | +/- | 86. |
| 3.20 | +/- | 0. |
| 3.03 | +/- | 0. |
| 1.14 | +/- | 0. |
| 1.14 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.08 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.32 |
| -9999.00 |

120-08-RV
| 4522. | +/- | 3. |
| 4602. | +/- | 78. |
| 2.78 | +/- | 0. |
| 2.55 | +/- | 0. |
| 1.33 | +/- | 0. |
| 1.33 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.76 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.17 |
| -9999.00 |

PERSIST_LOW,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES
120-08-RV
| 8234. | +/- | 23. |
| 8120. | +/- | 302. |
| 4.24 | +/- | 0. |
| 4.66 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.67 | +/- | 0. |
| 0.75 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.22 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -1.75 |
| -9999.00 |
PERSIST_LOW
120-08-RV
| 3990. | +/- | 1. |
| 4102. | +/- | 72. |
| 1.32 | +/- | 0. |
| 1.24 | +/- | 0. |
| 1.84 | +/- | 0. |
| 1.84 | +/- | -NaN |
| -0.56 | +/- | 0. |
| -0.56 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.09 | +/- | 0. |
| 4.11 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.80 |
| -9999.00 |
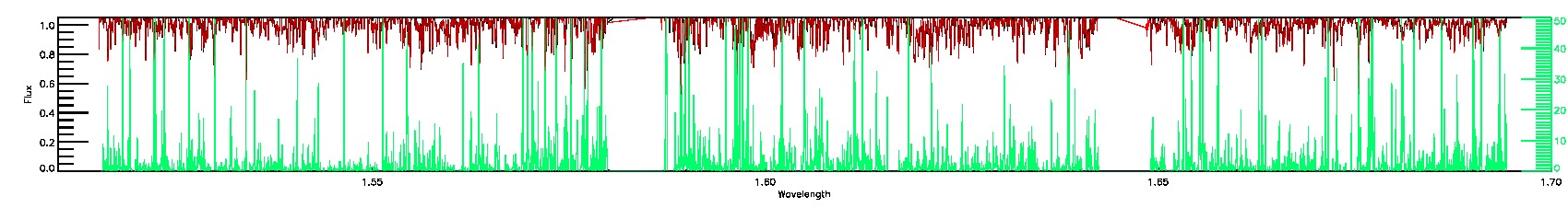
PERSIST_LOW
120-08-RV
| 5145. | +/- | 7. |
| 5170. | +/- | 98. |
| 2.87 | +/- | 0. |
| 2.51 | +/- | 0. |
| 1.85 | +/- | 0. |
| 1.85 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -0.25 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.06 | +/- | 0. |
| 3.43 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.41 |
| -9999.00 |
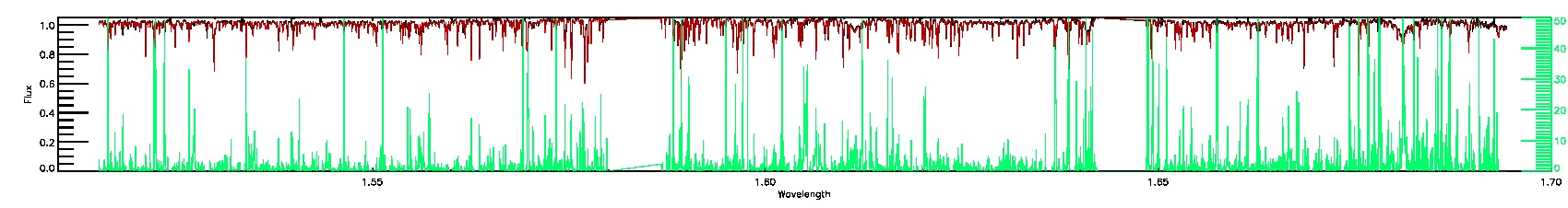
120-08-RV
| 4835. | +/- | 5. |
| 4893. | +/- | 89. |
| 3.27 | +/- | 0. |
| 3.13 | +/- | 0. |
| 1.11 | +/- | 0. |
| 1.11 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -0.20 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.06 | +/- | 0. |
| 3.32 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.05 |
| -9999.00 |
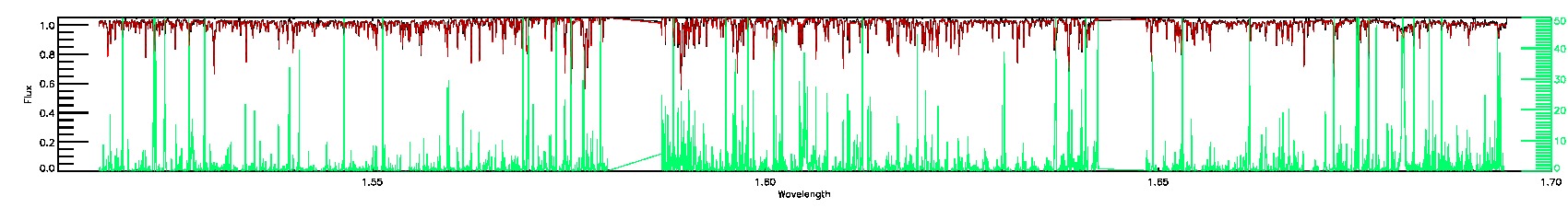
PERSIST_LOW
120-08-RV
| 4392. | +/- | 2. |
| 4476. | +/- | 75. |
| 2.33 | +/- | 0. |
| 2.08 | +/- | 0. |
| 1.39 | +/- | 0. |
| 1.39 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.44 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.77 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.17 |
| -9999.00 |

PERSIST_LOW
120-08-RV
| 4874. | +/- | 6. |
| 4931. | +/- | 91. |
| 2.81 | +/- | 0. |
| 2.46 | +/- | 0. |
| 1.60 | +/- | 0. |
| 1.60 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -0.28 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.48 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.34 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD
120-08-RV
| 5500. | +/- | 0. |
| -10000. | +/- | -NaN |
| 3.35 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.20 | +/- | 0. |
| 2.20 | +/- | -NaN |
| 0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 15.21 | +/- | 0. |
| 1.18 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.46 |
| -9999.00 |
PERSIST_LOW
120-08-RV
| 4315. | +/- | 2. |
| 4401. | +/- | 74. |
| 2.30 | +/- | 0. |
| 2.06 | +/- | 0. |
| 1.38 | +/- | 0. |
| 1.38 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.84 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| -0.01 |
| -9999.00 |
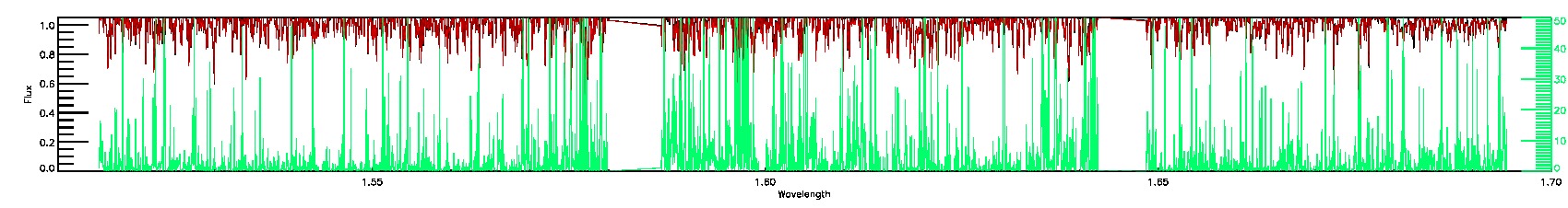
120-08-RV
| 3688. | +/- | 1. |
| 3773. | +/- | 61. |
| 1.18 | +/- | 0. |
| 1.02 | +/- | 0. |
| 1.55 | +/- | 0. |
| 1.55 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.05 | +/- | 0. |
| 2.82 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
BRIGHT_NEIGHBOR,PERSIST_LOW
120-08-RV
| 4200. | +/- | 1. |
| 4302. | +/- | 75. |
| 2.07 | +/- | 0. |
| 1.87 | +/- | 0. |
| 1.38 | +/- | 0. |
| 1.38 | +/- | -NaN |
| -0.30 | +/- | 0. |
| -0.30 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.22 | +/- | 0. |
| 3.53 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.38 |
| -9999.00 |
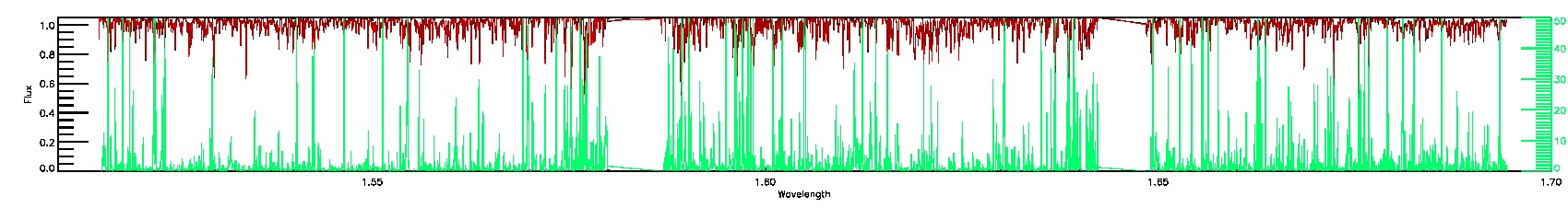
SUSPECT_RV_COMBINATION
120-08-RV
| 7955. | +/- | 12. |
| 7781. | +/- | 236. |
| 4.57 | +/- | 0. |
| 4.42 | +/- | 0. |
| 0.71 | +/- | 0. |
| 0.71 | +/- | -NaN |
| -0.65 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| 0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 36.08 | +/- | 0. |
| 1.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.49 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -1.36 |
| -9999.00 |
BRIGHT_NEIGHBOR,PERSIST_LOW
120-08-RV
| 4870. | +/- | 5. |
| 4904. | +/- | 85. |
| 3.06 | +/- | 0. |
| 2.69 | +/- | 0. |
| 1.26 | +/- | 0. |
| 1.26 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.49 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.51 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| 0.10 |
| -9999.00 |

PERSIST_LOW
120-08-RV
| 5188. | +/- | 8. |
| 5199. | +/- | 97. |
| 3.13 | +/- | 0. |
| 2.76 | +/- | 0. |
| 1.47 | +/- | 0. |
| 1.47 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.18 | +/- | 0. |
| -0.18 | +/- | -NaN |
| 0.42 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 3.06 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.20 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |

120-08-RV
| 4916. | +/- | 5. |
| 4954. | +/- | 89. |
| 3.09 | +/- | 0. |
| 2.72 | +/- | 0. |
| 1.27 | +/- | 0. |
| 1.27 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.88 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.24 |
| -9999.00 |
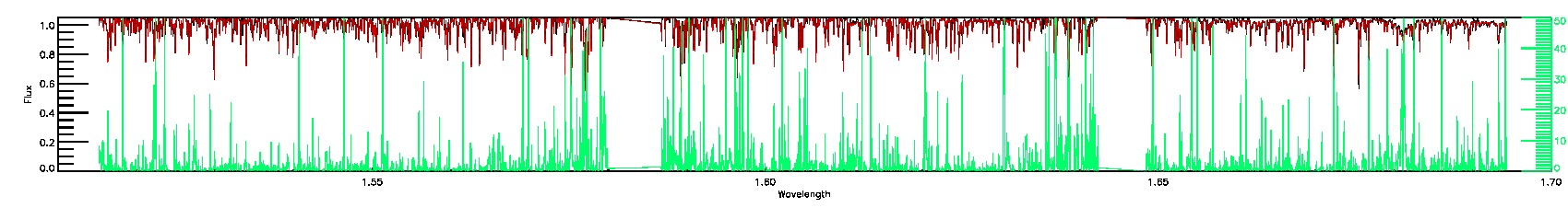
PERSIST_LOW,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN
120-08-RV
| 6369. | +/- | 27. |
| -10000. | +/- | -NaN |
| 4.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.46 | +/- | 0. |
| 2.46 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 14.87 | +/- | 0. |
| 1.17 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -2.41 |
| -9999.00 |
120-08-RV
| 4082. | +/- | 1. |
| 4167. | +/- | 69. |
| 1.94 | +/- | 0. |
| 1.70 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.43 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.83 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.02 |
| -9999.00 |

PERSIST_LOW
120-08-RV
| 4731. | +/- | 4. |
| 4798. | +/- | 85. |
| 2.67 | +/- | 0. |
| 2.37 | +/- | 0. |
| 1.48 | +/- | 0. |
| 1.48 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.17 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.22 |
| -9999.00 |
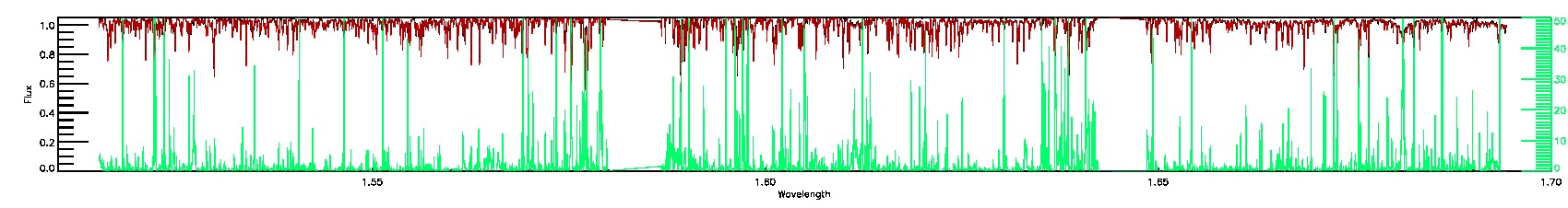
120-08-RV
| 4611. | +/- | 4. |
| 4701. | +/- | 85. |
| 2.54 | +/- | 0. |
| 2.36 | +/- | 0. |
| 1.40 | +/- | 0. |
| 1.40 | +/- | -NaN |
| -0.33 | +/- | 0. |
| -0.33 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.60 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -1.03 |
| -9999.00 |

PERSIST_LOW,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
120-08-RV
| 8371. | +/- | 28. |
| 8226. | +/- | 289. |
| 4.37 | +/- | 0. |
| 4.49 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -1.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 80.17 | +/- | 0. |
| 1.90 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.80 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -1.07 |
| -9999.00 |
PERSIST_LOW,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_WARN,COLORTE_WARN
120-08-RV
| 11970. | +/- | 63. |
| -10000. | +/- | -NaN |
| 4.26 | +/- | 0. |
| 4.14 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -0.74 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.16 | +/- | 0. |
| 1.01 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.74 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.74 |
| -9999.00 |
PERSIST_LOW,SUSPECT_RV_COMBINATION
STAR_BAD
120-08-RV
| 8131. | +/- | 26. |
| -10000. | +/- | -NaN |
| 4.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -1.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 94.75 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.36 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -1.63 |
| -9999.00 |
SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
120-08-RV
| 8356. | +/- | 26. |
| 8251. | +/- | 322. |
| 4.19 | +/- | 0. |
| 4.70 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 21.99 | +/- | 0. |
| 1.34 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.40 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -2.43 |
| -9999.00 |
PERSIST_LOW
120-08-RV
| 4220. | +/- | 1. |
| 4321. | +/- | 75. |
| 2.01 | +/- | 0. |
| 1.82 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.43 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -0.29 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.50 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.67 |
| -9999.00 |

120-08-RV
| 3640. | +/- | 1. |
| 3733. | +/- | 62. |
| 0.83 | +/- | 0. |
| 0.69 | +/- | 0. |
| 1.72 | +/- | 0. |
| 1.72 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.07 | +/- | 0. |
| 3.14 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.22 |
| -9999.00 |
120-08-RV
| 4662. | +/- | 4. |
| 4736. | +/- | 84. |
| 2.71 | +/- | 0. |
| 2.39 | +/- | 0. |
| 1.48 | +/- | 0. |
| 1.48 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.17 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.24 |
| -9999.00 |
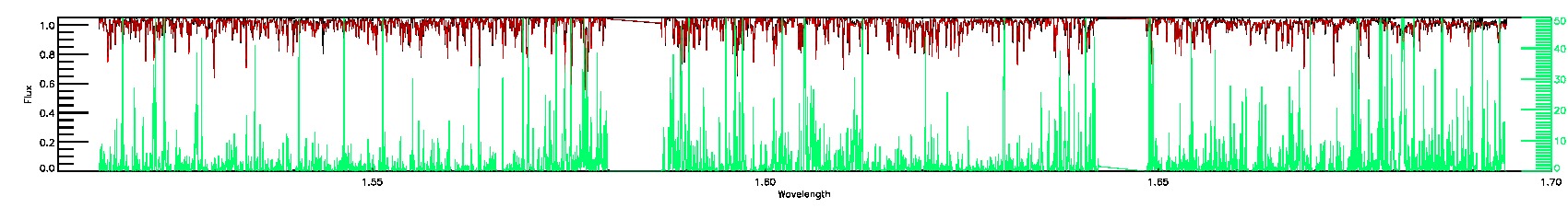
120-08-RV
| 4718. | +/- | 4. |
| 4774. | +/- | 82. |
| 2.83 | +/- | 0. |
| 2.48 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.43 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.67 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| 0.14 |
| -9999.00 |

BRIGHT_NEIGHBOR,SUSPECT_RV_COMBINATION
STAR_WARN,COLORTE_WARN
120-08-RV
| 8223. | +/- | 30. |
| 8102. | +/- | 296. |
| 4.32 | +/- | 0. |
| 4.67 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -1.88 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 53.62 | +/- | 0. |
| 1.73 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.08 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -2.28 |
| -9999.00 |
120-08-RV
| 4795. | +/- | 5. |
| 4862. | +/- | 89. |
| 2.71 | +/- | 0. |
| 2.40 | +/- | 0. |
| 1.60 | +/- | 0. |
| 1.60 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -0.31 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.54 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.73 |
| -9999.00 |

PERSIST_LOW
STAR_WARN,COLORTE_WARN
120-08-RV
| 3581. | +/- | 1. |
| 3686. | +/- | 63. |
| 0.53 | +/- | 0. |
| 0.43 | +/- | 0. |
| 2.01 | +/- | 0. |
| 2.01 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -0.40 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.11 | +/- | 0. |
| 3.73 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.77 |
| -9999.00 |
120-08-RV
| 4400. | +/- | 2. |
| 4493. | +/- | 77. |
| 2.11 | +/- | 0. |
| 1.89 | +/- | 0. |
| 1.63 | +/- | 0. |
| 1.63 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| 0.43 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.00 | +/- | 0. |
| 3.12 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.44 |
| -9999.00 |

SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES
STAR_BAD
120-08-RV
| 7961. | +/- | 16. |
| -10000. | +/- | -NaN |
| 5.21 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.99 | +/- | 2. |
| 0.99 | +/- | -NaN |
| -2.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.41 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.97 | +/- | 0. |
| 0.29 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.18 |
| -9999.00 |
PERSIST_LOW
120-08-RV
| 4158. | +/- | 1. |
| 4267. | +/- | 75. |
| 1.73 | +/- | 0. |
| 1.58 | +/- | 0. |
| 1.70 | +/- | 0. |
| 1.70 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -0.49 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.10 | +/- | 0. |
| 3.94 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.91 |
| -9999.00 |

120-08-RV
| 4185. | +/- | 1. |
| 4287. | +/- | 74. |
| 1.81 | +/- | 0. |
| 1.62 | +/- | 0. |
| 1.80 | +/- | 0. |
| 1.80 | +/- | -NaN |
| -0.32 | +/- | 0. |
| -0.32 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.57 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.95 |
| -9999.00 |
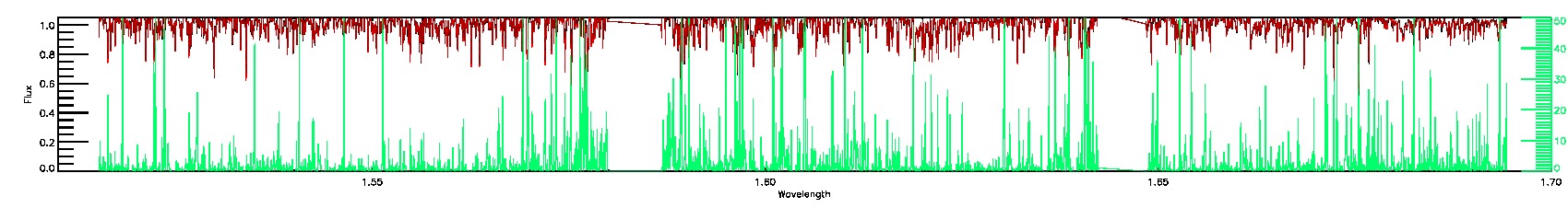
120-08-RV
| 4849. | +/- | 6. |
| 4906. | +/- | 89. |
| 3.18 | +/- | 0. |
| 3.03 | +/- | 0. |
| 1.23 | +/- | 0. |
| 1.23 | +/- | -NaN |
| -0.21 | +/- | 0. |
| -0.21 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.34 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -1.05 |
| -9999.00 |

PERSIST_LOW
120-08-RV
| 4189. | +/- | 1. |
| 4266. | +/- | 70. |
| 2.27 | +/- | 0. |
| 2.00 | +/- | 0. |
| 1.35 | +/- | 0. |
| 1.35 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.52 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 0.32 |
| -9999.00 |

PERSIST_LOW
120-08-RV
| 4678. | +/- | 4. |
| 4740. | +/- | 82. |
| 2.91 | +/- | 0. |
| 2.69 | +/- | 0. |
| 1.32 | +/- | 0. |
| 1.32 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.73 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.01 |
| -9999.00 |
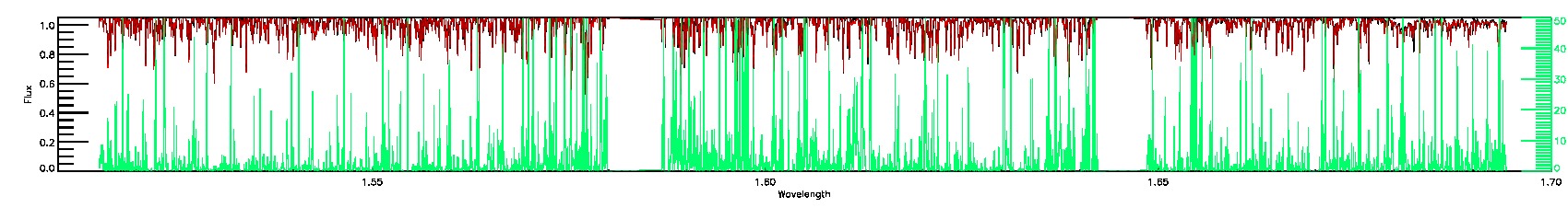
120-08-RV
| 4242. | +/- | 2. |
| 4341. | +/- | 75. |
| 1.98 | +/- | 0. |
| 1.77 | +/- | 0. |
| 1.53 | +/- | 0. |
| 1.53 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.24 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.39 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.18 |
| -9999.00 |

120-08-RV
| 4796. | +/- | 5. |
| 4868. | +/- | 90. |
| 2.65 | +/- | 0. |
| 2.36 | +/- | 0. |
| 1.58 | +/- | 0. |
| 1.58 | +/- | -NaN |
| -0.43 | +/- | 0. |
| -0.43 | +/- | 0. |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.19 | +/- | 0. |
| 3.81 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.85 |
| -9999.00 |

120-08-RV
| 4322. | +/- | 2. |
| 4420. | +/- | 77. |
| 2.26 | +/- | 0. |
| 2.06 | +/- | 0. |
| 1.34 | +/- | 0. |
| 1.34 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.24 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.07 | +/- | 0. |
| 3.40 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.73 |
| -9999.00 |

120-08-RV
| 4214. | +/- | 1. |
| 4325. | +/- | 77. |
| 1.79 | +/- | 0. |
| 1.63 | +/- | 0. |
| 1.62 | +/- | 0. |
| 1.62 | +/- | -NaN |
| -0.51 | +/- | 0. |
| -0.51 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.07 | +/- | 0. |
| 3.98 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.69 |
| -9999.00 |

120-08-RV
| 4576. | +/- | 3. |
| 4644. | +/- | 78. |
| 2.79 | +/- | 0. |
| 2.45 | +/- | 0. |
| 1.25 | +/- | 0. |
| 1.25 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.52 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| 0.29 |
| -9999.00 |

120-08-RV
| 4074. | +/- | 1. |
| 4174. | +/- | 72. |
| 1.83 | +/- | 0. |
| 1.64 | +/- | 0. |
| 1.50 | +/- | 0. |
| 1.50 | +/- | -NaN |
| -0.27 | +/- | 0. |
| -0.27 | +/- | 0. |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.19 | +/- | 0. |
| 3.46 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.81 |
| -9999.00 |

BRIGHT_NEIGHBOR,PERSIST_LOW
120-08-RV
| 4784. | +/- | 5. |
| 4842. | +/- | 86. |
| 2.79 | +/- | 0. |
| 2.45 | +/- | 0. |
| 1.48 | +/- | 0. |
| 1.48 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.05 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.26 |
| -9999.00 |

PERSIST_LOW
120-08-RV
| 4730. | +/- | 4. |
| 4797. | +/- | 85. |
| 2.67 | +/- | 0. |
| 2.37 | +/- | 0. |
| 1.54 | +/- | 0. |
| 1.54 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.13 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.18 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -1.05 |
| -9999.00 |

120-08-RV
| 4862. | +/- | 5. |
| 4916. | +/- | 89. |
| 2.84 | +/- | 0. |
| 2.48 | +/- | 0. |
| 1.47 | +/- | 0. |
| 1.47 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -0.18 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.28 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.06 |
| -9999.00 |

120-08-RV
| 4517. | +/- | 3. |
| 4616. | +/- | 82. |
| 2.59 | +/- | 0. |
| 2.40 | +/- | 0. |
| 1.28 | +/- | 0. |
| 1.28 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -0.29 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.50 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.74 |
| -9999.00 |

120-08-RV
| 4307. | +/- | 2. |
| 4409. | +/- | 77. |
| 2.02 | +/- | 0. |
| 1.83 | +/- | 0. |
| 1.56 | +/- | 0. |
| 1.56 | +/- | -NaN |
| -0.33 | +/- | 0. |
| -0.33 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.06 | +/- | 0. |
| 3.58 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.80 |
| -9999.00 |

120-08-RV
| 4562. | +/- | 3. |
| 4656. | +/- | 83. |
| 2.45 | +/- | 0. |
| 2.25 | +/- | 0. |
| 1.54 | +/- | 0. |
| 1.54 | +/- | -NaN |
| -0.30 | +/- | 0. |
| -0.30 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.52 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.61 |
| -9999.00 |
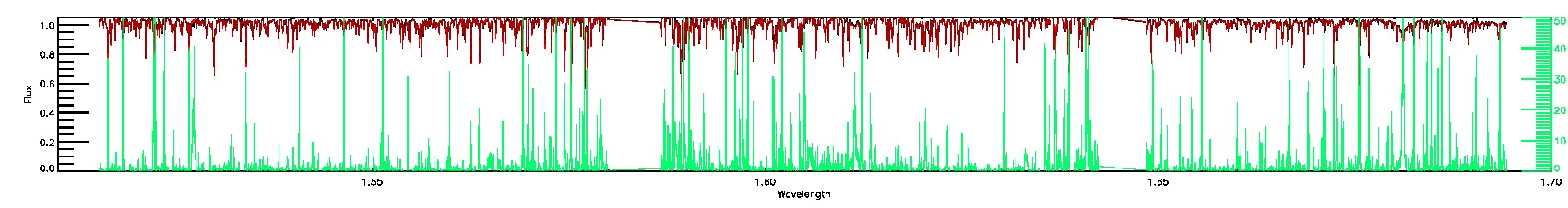
120-08-RV
| 4946. | +/- | 6. |
| 4985. | +/- | 90. |
| 2.98 | +/- | 0. |
| 2.60 | +/- | 0. |
| 1.48 | +/- | 0. |
| 1.48 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.01 | +/- | 0. |
| 3.06 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.44 |
| -9999.00 |

BRIGHT_NEIGHBOR
120-08-RV
| 4854. | +/- | 5. |
| 4902. | +/- | 88. |
| 3.09 | +/- | 0. |
| 2.71 | +/- | 0. |
| 1.20 | +/- | 0. |
| 1.20 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| 3.01 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.04 |
| -9999.00 |
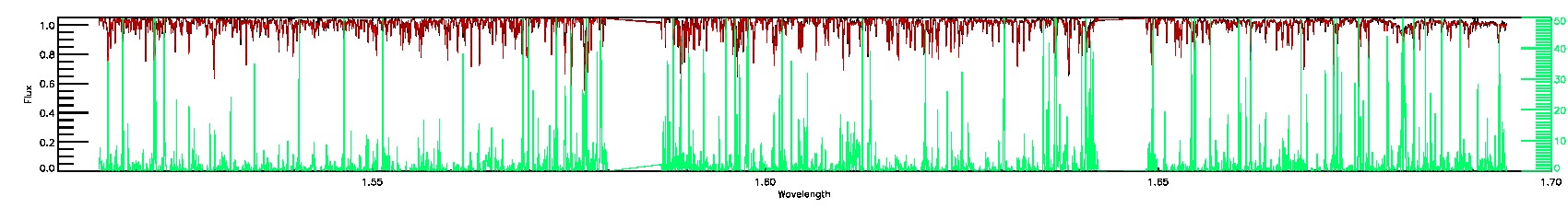
BRIGHT_NEIGHBOR,PERSIST_LOW
120-08-RV
| 4766. | +/- | 5. |
| 4837. | +/- | 88. |
| 2.65 | +/- | 0. |
| 2.36 | +/- | 0. |
| 1.63 | +/- | 0. |
| 1.63 | +/- | -NaN |
| -0.32 | +/- | 0. |
| -0.32 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.06 | +/- | 0. |
| 3.57 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.55 |
| -9999.00 |

120-08-RV
| 4061. | +/- | 1. |
| 4158. | +/- | 71. |
| 1.81 | +/- | 0. |
| 1.62 | +/- | 0. |
| 1.45 | +/- | 0. |
| 1.45 | +/- | -NaN |
| -0.21 | +/- | 0. |
| -0.21 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.34 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.04 |
| -9999.00 |

120-08-RV
| 3868. | +/- | 1. |
| 3952. | +/- | 64. |
| 1.52 | +/- | 0. |
| 1.31 | +/- | 0. |
| 1.55 | +/- | 0. |
| 1.55 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.04 | +/- | 0. |
| 2.80 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.11 |
| -9999.00 |
SUSPECT_RV_COMBINATION
STAR_BAD
120-08-RV
| 7992. | +/- | 13. |
| -10000. | +/- | -NaN |
| 4.45 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| -0.77 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.26 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 55.72 | +/- | 0. |
| 1.75 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.30 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.04 |
| -9999.00 |
BRIGHT_NEIGHBOR,PERSIST_LOW
120-08-RV
| 4535. | +/- | 3. |
| 4629. | +/- | 82. |
| 2.41 | +/- | 0. |
| 2.20 | +/- | 0. |
| 1.45 | +/- | 0. |
| 1.45 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -0.23 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.39 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.72 |
| -9999.00 |

120-08-RV
| 4217. | +/- | 1. |
| 4297. | +/- | 71. |
| 2.16 | +/- | 0. |
| 1.90 | +/- | 0. |
| 1.38 | +/- | 0. |
| 1.38 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.64 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.16 |
| -9999.00 |
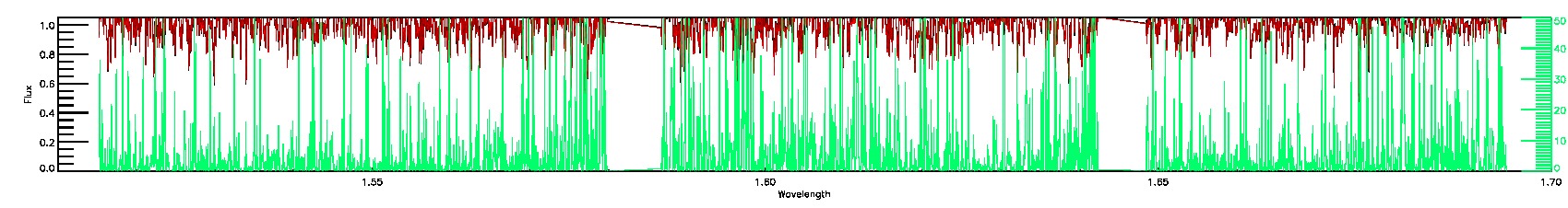
120-08-RV
| 4942. | +/- | 7. |
| 4998. | +/- | 94. |
| 3.11 | +/- | 0. |
| 2.98 | +/- | 0. |
| 1.29 | +/- | 0. |
| 1.29 | +/- | -NaN |
| -0.45 | +/- | 0. |
| -0.45 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.84 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.43 |
| -9999.00 |

PERSIST_LOW
120-08-RV
| 3995. | +/- | 1. |
| 4113. | +/- | 73. |
| 1.30 | +/- | 0. |
| 1.24 | +/- | 0. |
| 1.86 | +/- | 0. |
| 1.86 | +/- | -NaN |
| -0.69 | +/- | 0. |
| -0.69 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.29 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.08 | +/- | 0. |
| 4.42 | +/- | 0. |
| 0.65 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.93 |
| -9999.00 |
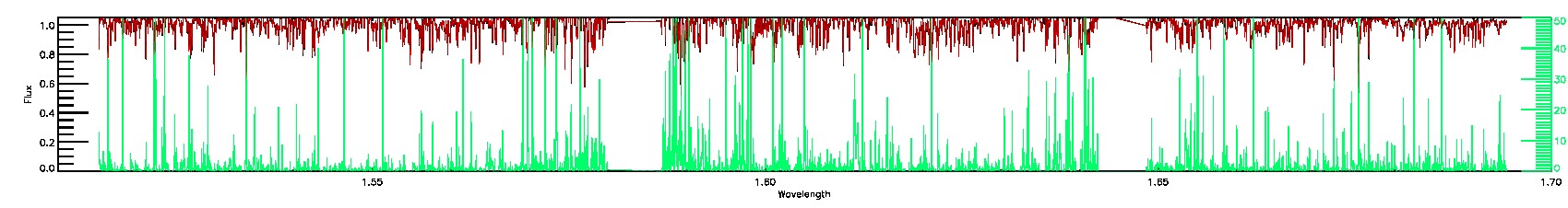
PERSIST_LOW
120-08-RV
| 4054. | +/- | 1. |
| 4149. | +/- | 70. |
| 1.71 | +/- | 0. |
| 1.52 | +/- | 0. |
| 1.58 | +/- | 0. |
| 1.58 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -0.16 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.25 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.49 |
| -9999.00 |

120-08-RV
| 4965. | +/- | 5. |
| 4996. | +/- | 89. |
| 2.87 | +/- | 0. |
| 2.51 | +/- | 0. |
| 1.60 | +/- | 0. |
| 1.60 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.45 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.82 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.23 |
| -9999.00 |

120-08-RV
| 4638. | +/- | 4. |
| 4725. | +/- | 85. |
| 2.70 | +/- | 0. |
| 2.52 | +/- | 0. |
| 1.45 | +/- | 0. |
| 1.45 | +/- | -NaN |
| -0.33 | +/- | 0. |
| -0.33 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.08 | +/- | 0. |
| 3.59 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.56 |
| -9999.00 |

SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD
120-08-RV
| 8619. | +/- | 43. |
| -10000. | +/- | -NaN |
| 4.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -1.60 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 94.38 | +/- | 0. |
| 1.97 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.01 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -1.89 |
| -9999.00 |
PERSIST_LOW
120-08-RV
| 4111. | +/- | 1. |
| 4217. | +/- | 74. |
| 1.65 | +/- | 0. |
| 1.50 | +/- | 0. |
| 1.74 | +/- | 0. |
| 1.74 | +/- | -NaN |
| -0.41 | +/- | 0. |
| -0.41 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.06 | +/- | 0. |
| 3.76 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -1.11 |
| -9999.00 |

SUSPECT_RV_COMBINATION
120-08-RV
| 8086. | +/- | 32. |
| 7985. | +/- | 301. |
| 4.27 | +/- | 0. |
| 4.82 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 71.86 | +/- | 0. |
| 1.86 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.32 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -1.93 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -2.20 |
| -9999.00 |
120-08-RV
| 3738. | +/- | 1. |
| 3852. | +/- | 67. |
| 0.75 | +/- | 0. |
| 0.67 | +/- | 0. |
| 2.15 | +/- | 0. |
| 2.15 | +/- | -NaN |
| -0.60 | +/- | 0. |
| -0.59 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.09 | +/- | 0. |
| 4.19 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.79 |
| -9999.00 |
120-08-RV
| 4624. | +/- | 4. |
| 4716. | +/- | 86. |
| 2.51 | +/- | 0. |
| 2.34 | +/- | 0. |
| 1.41 | +/- | 0. |
| 1.41 | +/- | -NaN |
| -0.42 | +/- | 0. |
| -0.42 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.79 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.72 |
| -9999.00 |
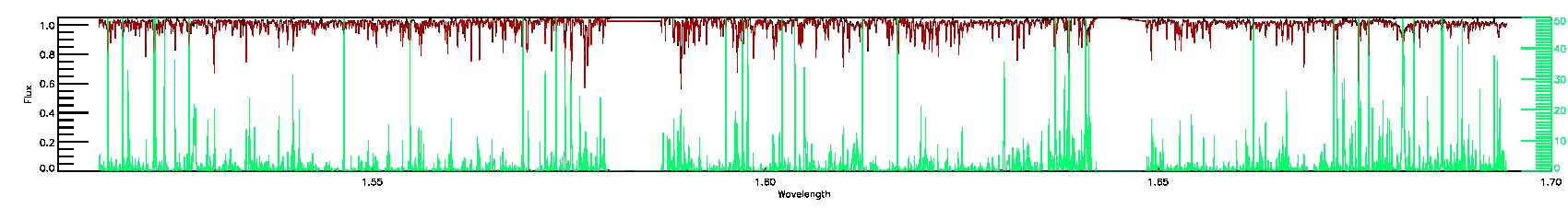
120-08-RV
| 4202. | +/- | 1. |
| 4292. | +/- | 72. |
| 2.01 | +/- | 0. |
| 1.78 | +/- | 0. |
| 1.49 | +/- | 0. |
| 1.49 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.04 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.00 |
| -9999.00 |

SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_WARN,COLORTE_WARN
120-08-RV
| 8266. | +/- | 24. |
| 8158. | +/- | 310. |
| 4.32 | +/- | 0. |
| 4.80 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 27.05 | +/- | 0. |
| 1.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.46 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -1.98 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -1.98 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -1.99 |
| -9999.00 |
120-08-RV
| 4528. | +/- | 3. |
| 4619. | +/- | 81. |
| 2.44 | +/- | 0. |
| 2.23 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.43 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.22 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.47 |
| -9999.00 |

120-08-RV
| 4261. | +/- | 2. |
| 4350. | +/- | 74. |
| 2.19 | +/- | 0. |
| 1.95 | +/- | 0. |
| 1.39 | +/- | 0. |
| 1.39 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.01 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.31 |
| -9999.00 |
120-08-RV
| 4242. | +/- | 1. |
| 4335. | +/- | 74. |
| 1.99 | +/- | 0. |
| 1.77 | +/- | 0. |
| 1.58 | +/- | 0. |
| 1.58 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.16 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.29 |
| -9999.00 |

BRIGHT_NEIGHBOR
120-08-RV
| 4932. | +/- | 6. |
| 4988. | +/- | 94. |
| 2.74 | +/- | 0. |
| 2.41 | +/- | 0. |
| 1.54 | +/- | 0. |
| 1.54 | +/- | -NaN |
| -0.41 | +/- | 0. |
| -0.41 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.06 | +/- | 0. |
| 3.75 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.48 |
| -9999.00 |

BRIGHT_NEIGHBOR
120-08-RV
| 4854. | +/- | 5. |
| 4909. | +/- | 89. |
| 2.82 | +/- | 0. |
| 2.47 | +/- | 0. |
| 1.55 | +/- | 0. |
| 1.55 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -0.20 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.32 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.35 |
| -9999.00 |
120-08-RV
| 4500. | +/- | 3. |
| 4608. | +/- | 83. |
| 2.51 | +/- | 0. |
| 2.34 | +/- | 0. |
| 1.39 | +/- | 0. |
| 1.39 | +/- | -NaN |
| -0.46 | +/- | 0. |
| -0.46 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.20 | +/- | 0. |
| 3.86 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.60 |
| -9999.00 |
BRIGHT_NEIGHBOR
120-08-RV
| 4710. | +/- | 4. |
| 4772. | +/- | 83. |
| 2.80 | +/- | 0. |
| 2.46 | +/- | 0. |
| 1.40 | +/- | 0. |
| 1.40 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.08 | +/- | 0. |
| 2.88 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.10 |
| -9999.00 |

BRIGHT_NEIGHBOR
120-08-RV
| 4852. | +/- | 5. |
| 4906. | +/- | 89. |
| 2.83 | +/- | 0. |
| 2.48 | +/- | 0. |
| 1.54 | +/- | 0. |
| 1.54 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 3.23 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.03 |
| -9999.00 |
120-08-RV
| 4773. | +/- | 5. |
| 4833. | +/- | 86. |
| 2.82 | +/- | 0. |
| 2.47 | +/- | 0. |
| 1.46 | +/- | 0. |
| 1.46 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.07 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.28 |
| -9999.00 |
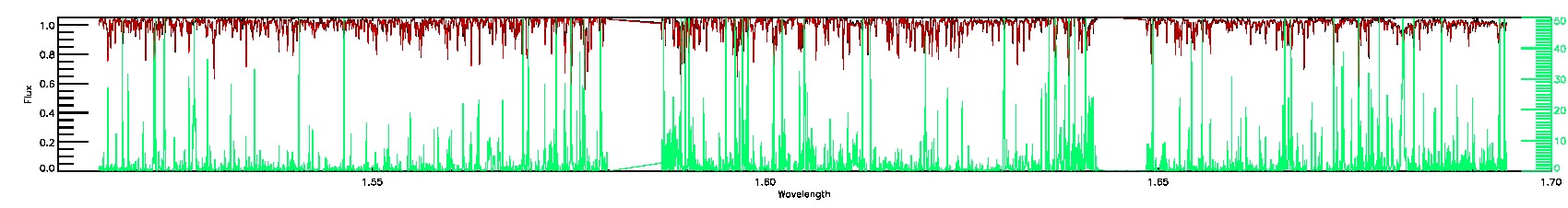
120-08-RV
| 5092. | +/- | 8. |
| 5123. | +/- | 97. |
| 3.29 | +/- | 0. |
| 2.95 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.43 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -0.26 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.45 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.19 |
| -9999.00 |
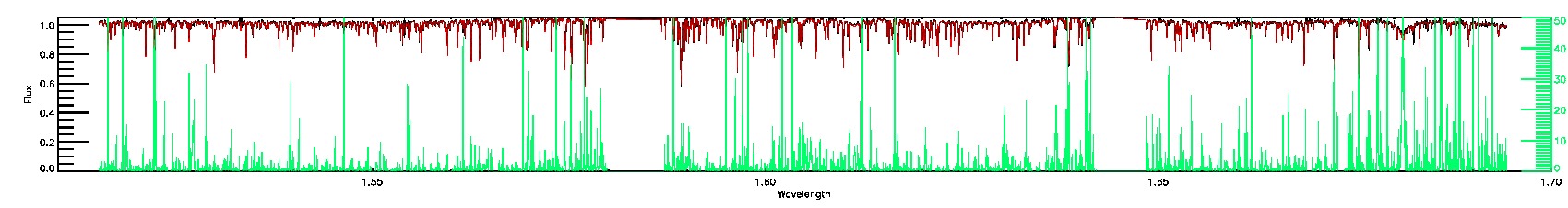
120-08-RV
| 4782. | +/- | 5. |
| 4843. | +/- | 87. |
| 2.70 | +/- | 0. |
| 2.39 | +/- | 0. |
| 1.51 | +/- | 0. |
| 1.51 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.18 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -1.09 |
| -9999.00 |

SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_WARN,COLORTE_WARN
120-08-RV
| 8196. | +/- | 29. |
| 8096. | +/- | 311. |
| 4.37 | +/- | 0. |
| 4.93 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 38.56 | +/- | 0. |
| 1.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.34 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -2.30 |
| -9999.00 |
BRIGHT_NEIGHBOR
120-08-RV
| 5201. | +/- | 7. |
| 5214. | +/- | 98. |
| 3.63 | +/- | 0. |
| 3.55 | +/- | 0. |
| 1.38 | +/- | 0. |
| 1.38 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 5.82 | +/- | 0. |
| 0.76 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.15 |
| -9999.00 |

120-08-RV
| 4106. | +/- | 1. |
| 4204. | +/- | 72. |
| 1.71 | +/- | 0. |
| 1.52 | +/- | 0. |
| 1.60 | +/- | 0. |
| 1.60 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -0.25 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.41 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.03 |
| -9999.00 |

120-08-RV
| 4702. | +/- | 5. |
| 4781. | +/- | 87. |
| 2.62 | +/- | 0. |
| 2.45 | +/- | 0. |
| 1.45 | +/- | 0. |
| 1.45 | +/- | -NaN |
| -0.33 | +/- | 0. |
| -0.33 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.59 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.06 |
| -9999.00 |
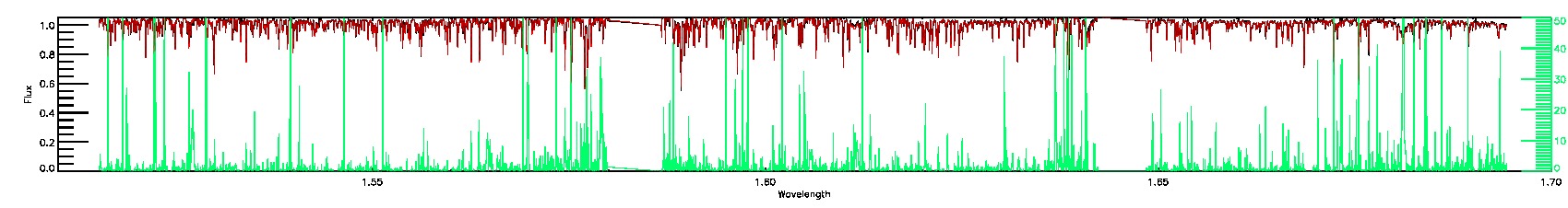
SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES
120-08-RV
| 8494. | +/- | 32. |
| 8383. | +/- | 329. |
| 4.42 | +/- | 0. |
| 4.87 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 29.11 | +/- | 0. |
| 1.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.33 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -1.93 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.12 |
| -9999.00 |
120-08-RV
| 4562. | +/- | 3. |
| 4631. | +/- | 78. |
| 2.88 | +/- | 0. |
| 2.64 | +/- | 0. |
| 1.24 | +/- | 0. |
| 1.24 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.52 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| 0.46 |
| -9999.00 |
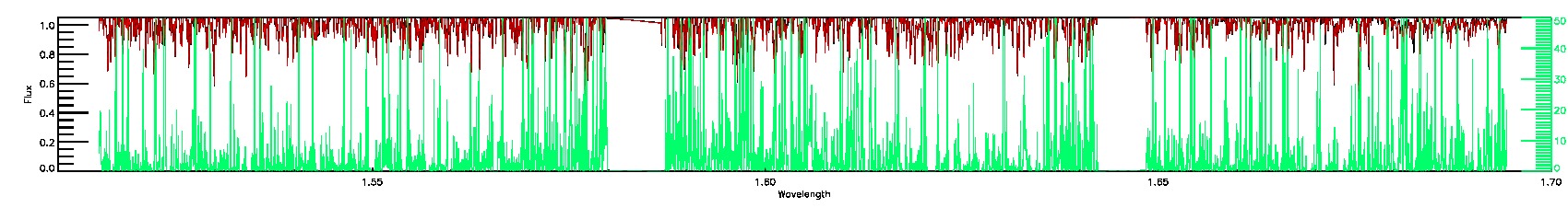
120-08-RV
| 4551. | +/- | 3. |
| 4636. | +/- | 80. |
| 2.51 | +/- | 0. |
| 2.29 | +/- | 0. |
| 1.48 | +/- | 0. |
| 1.48 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.06 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.35 |
| -9999.00 |
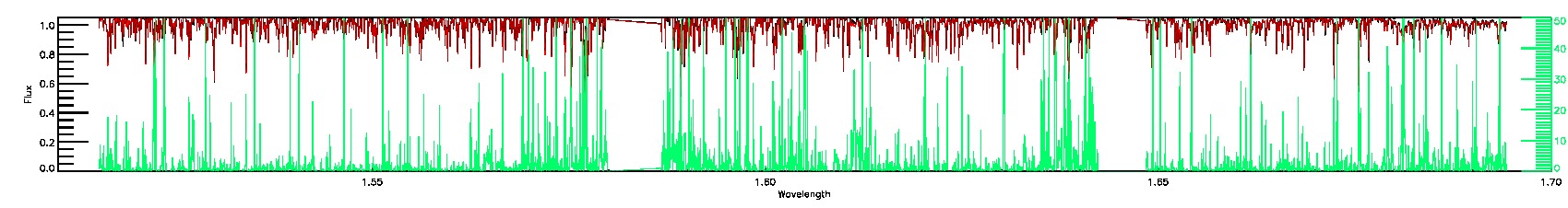
120-08-RV
| 4899. | +/- | 6. |
| 4956. | +/- | 92. |
| 2.77 | +/- | 0. |
| 2.43 | +/- | 0. |
| 1.73 | +/- | 0. |
| 1.73 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -0.35 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.63 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.18 |
| -9999.00 |

120-08-RV
| 4030. | +/- | 1. |
| 4134. | +/- | 71. |
| 1.50 | +/- | 0. |
| 1.36 | +/- | 0. |
| 1.71 | +/- | 0. |
| 1.71 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -0.36 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.06 | +/- | 0. |
| 3.65 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.60 |
| -9999.00 |

PERSIST_LOW
120-08-RV
| 4092. | +/- | 1. |
| 4199. | +/- | 73. |
| 1.64 | +/- | 0. |
| 1.49 | +/- | 0. |
| 1.78 | +/- | 0. |
| 1.78 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -0.44 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.09 | +/- | 0. |
| 3.83 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.77 |
| -9999.00 |

120-08-RV
| 4660. | +/- | 4. |
| 4729. | +/- | 82. |
| 2.73 | +/- | 0. |
| 2.41 | +/- | 0. |
| 1.42 | +/- | 0. |
| 1.42 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.04 | +/- | 0. |
| 2.92 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.22 |
| -9999.00 |
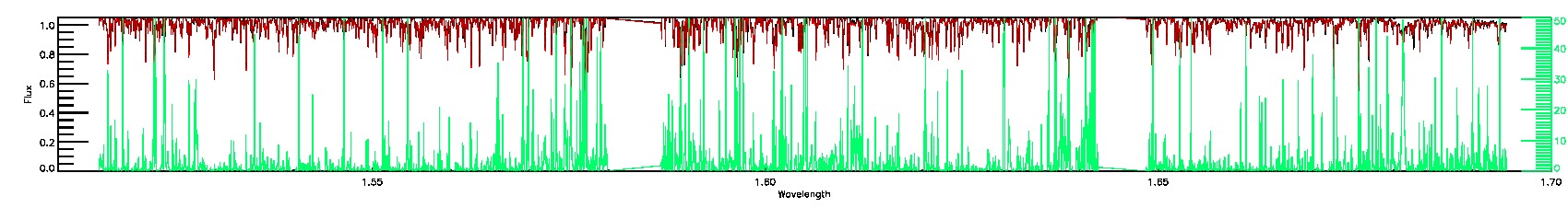
120-08-RV
| 4374. | +/- | 2. |
| 4477. | +/- | 79. |
| 1.99 | +/- | 0. |
| 1.80 | +/- | 0. |
| 1.66 | +/- | 0. |
| 1.66 | +/- | -NaN |
| -0.33 | +/- | 0. |
| -0.33 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.59 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.18 |
| -9999.00 |

120-08-RV
| 3805. | +/- | 1. |
| 3894. | +/- | 64. |
| 1.46 | +/- | 0. |
| 1.28 | +/- | 0. |
| 1.57 | +/- | 0. |
| 1.57 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | 0. |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.20 | +/- | 0. |
| 3.01 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.15 |
| -9999.00 |
120-08-RV
| 4945. | +/- | 5. |
| 4983. | +/- | 90. |
| 2.98 | +/- | 0. |
| 2.60 | +/- | 0. |
| 1.42 | +/- | 0. |
| 1.42 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.43 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.00 | +/- | 0. |
| 3.00 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.19 |
| -9999.00 |

120-08-RV
| 4180. | +/- | 1. |
| 4275. | +/- | 73. |
| 1.90 | +/- | 0. |
| 1.69 | +/- | 0. |
| 1.57 | +/- | 0. |
| 1.57 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -0.14 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.22 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.27 |
| -9999.00 |
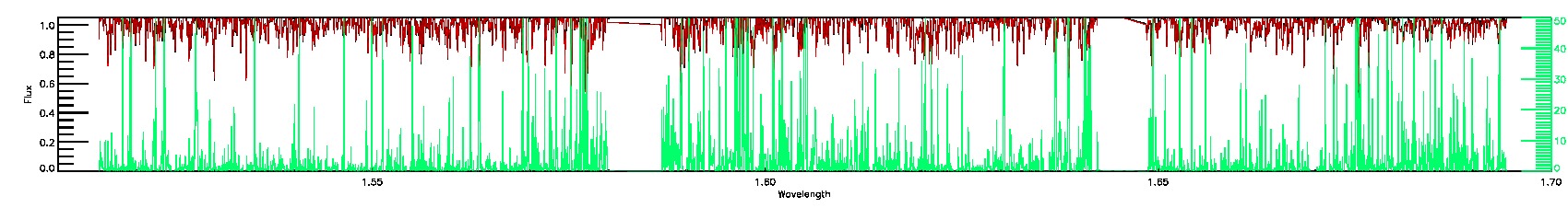
120-08-RV
| 4196. | +/- | 1. |
| 4290. | +/- | 73. |
| 1.91 | +/- | 0. |
| 1.70 | +/- | 0. |
| 1.52 | +/- | 0. |
| 1.52 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -0.14 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.13 | +/- | 0. |
| 3.21 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.13 |
| -9999.00 |

120-08-RV
| 3810. | +/- | 1. |
| 3892. | +/- | 63. |
| 1.42 | +/- | 0. |
| 1.22 | +/- | 0. |
| 1.51 | +/- | 0. |
| 1.51 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.72 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.22 |
| -9999.00 |
120-08-RV
| 4317. | +/- | 2. |
| 4421. | +/- | 78. |
| 2.04 | +/- | 0. |
| 1.86 | +/- | 0. |
| 1.59 | +/- | 0. |
| 1.59 | +/- | -NaN |
| -0.37 | +/- | 0. |
| -0.37 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.06 | +/- | 0. |
| 3.67 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.54 |
| -9999.00 |

BRIGHT_NEIGHBOR
120-08-RV
| 4750. | +/- | 5. |
| 4819. | +/- | 87. |
| 2.67 | +/- | 0. |
| 2.37 | +/- | 0. |
| 1.55 | +/- | 0. |
| 1.55 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -0.23 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.06 | +/- | 0. |
| 3.38 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.34 |
| -9999.00 |
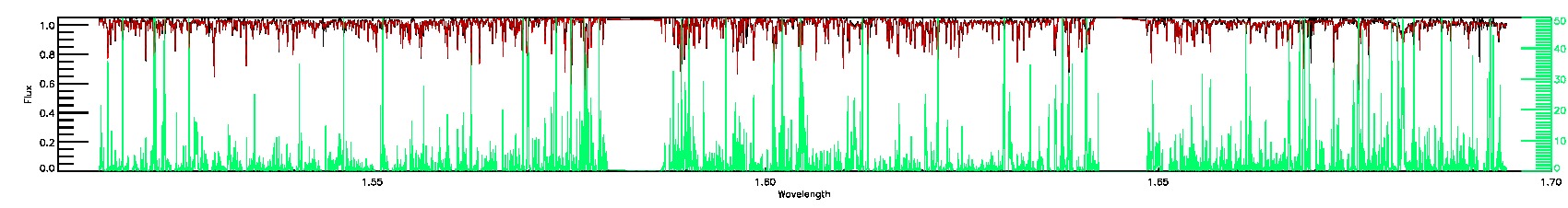
BRIGHT_NEIGHBOR
120-08-RV
| 5014. | +/- | 8. |
| 5052. | +/- | 94. |
| 3.39 | +/- | 0. |
| 3.07 | +/- | 0. |
| 1.20 | +/- | 0. |
| 1.20 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -0.23 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.39 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.85 |
| -9999.00 |
120-08-RV
| 4911. | +/- | 6. |
| 4965. | +/- | 92. |
| 2.82 | +/- | 0. |
| 2.47 | +/- | 0. |
| 1.53 | +/- | 0. |
| 1.53 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -0.31 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.54 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.73 |
| -9999.00 |
120-08-RV
| 4879. | +/- | 6. |
| 4934. | +/- | 91. |
| 2.74 | +/- | 0. |
| 2.41 | +/- | 0. |
| 1.65 | +/- | 0. |
| 1.65 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -0.25 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.41 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.23 |
| -9999.00 |
120-08-RV
| 4652. | +/- | 3. |
| 4738. | +/- | 86. |
| 2.39 | +/- | 0. |
| 2.24 | +/- | 0. |
| 1.53 | +/- | 0. |
| 1.53 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -0.36 | +/- | 0. |
| 0.27 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.66 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.26 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -1.29 |
| -9999.00 |

120-08-RV
| 4180. | +/- | 1. |
| 4260. | +/- | 70. |
| 2.08 | +/- | 0. |
| 1.81 | +/- | 0. |
| 1.36 | +/- | 0. |
| 1.36 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.63 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.22 |
| -9999.00 |

120-08-RV
| 4430. | +/- | 2. |
| 4525. | +/- | 79. |
| 2.19 | +/- | 0. |
| 1.97 | +/- | 0. |
| 1.60 | +/- | 0. |
| 1.60 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -0.18 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.28 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.76 |
| -9999.00 |

SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_WARN,COLORTE_WARN
120-08-RV
| 8503. | +/- | 31. |
| 8395. | +/- | 333. |
| 3.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.20 | +/- | 0. |
| -2.20 | +/- | 1. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 1. |
| 17.17 | +/- | 0. |
| 1.23 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.41 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -1.98 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -1.98 |
| -9999.00 |
120-08-RV
| 4948. | +/- | 6. |
| 4997. | +/- | 93. |
| 2.83 | +/- | 0. |
| 2.48 | +/- | 0. |
| 1.71 | +/- | 0. |
| 1.71 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -0.29 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.50 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.33 |
| -9999.00 |
120-08-RV
| 3935. | +/- | 1. |
| 4038. | +/- | 69. |
| 1.39 | +/- | 0. |
| 1.26 | +/- | 0. |
| 1.70 | +/- | 0. |
| 1.70 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -0.35 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.09 | +/- | 0. |
| 3.64 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -1.03 |
| -9999.00 |
120-08-RV
| 4302. | +/- | 2. |
| 4388. | +/- | 74. |
| 2.36 | +/- | 0. |
| 2.12 | +/- | 0. |
| 1.26 | +/- | 0. |
| 1.26 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.11 | +/- | 0. |
| 2.84 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.40 |
| -9999.00 |

120-08-RV
| 4834. | +/- | 5. |
| 4888. | +/- | 88. |
| 2.97 | +/- | 0. |
| 2.59 | +/- | 0. |
| 1.36 | +/- | 0. |
| 1.36 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 3.14 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.42 |
| -9999.00 |

120-08-RV
| 5069. | +/- | 6. |
| 5097. | +/- | 95. |
| 2.92 | +/- | 0. |
| 2.55 | +/- | 0. |
| 1.76 | +/- | 0. |
| 1.76 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -0.14 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.20 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.65 |
| -9999.00 |

SUSPECT_RV_COMBINATION
STAR_BAD
STAR_WARN,COLORTE_WARN
120-08-RV
| 7992. | +/- | 12. |
| -10000. | +/- | -NaN |
| 4.30 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.13 | +/- | 0. |
| 1.13 | +/- | -NaN |
| -0.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 52.88 | +/- | 0. |
| 1.72 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.30 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.45 |
| -9999.00 |
120-08-RV
| 4466. | +/- | 2. |
| 4548. | +/- | 77. |
| 2.47 | +/- | 0. |
| 2.22 | +/- | 0. |
| 1.39 | +/- | 0. |
| 1.39 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.71 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -0.14 |
| -9999.00 |

