160-45
| 4684. | +/- | 10. |
| 4779. | +/- | 111. |
| 2.64 | +/- | 0. |
| 2.51 | +/- | 0. |
| 1.10 | +/- | 0. |
| 1.10 | +/- | -NaN |
| -0.65 | +/- | 0. |
| -0.65 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.13 | +/- | 0. |
| 4.33 | +/- | 0. |
| 0.64 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -1.39 |
| -9999.00 |

PERSIST_LOW
160-45
| 6804. | +/- | 16. |
| 6644. | +/- | 164. |
| 4.52 | +/- | 0. |
| 4.41 | +/- | 0. |
| 1.52 | +/- | 0. |
| 1.52 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -0.44 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.09 | +/- | 0. |
| 6.25 | +/- | 0. |
| 0.80 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.45 |
| -9999.00 |
BRIGHT_NEIGHBOR
160-45
| 6485. | +/- | 20. |
| 6357. | +/- | 147. |
| 4.19 | +/- | 0. |
| 4.10 | +/- | 0. |
| 1.62 | +/- | 0. |
| 1.62 | +/- | -NaN |
| -0.32 | +/- | 0. |
| -0.32 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.12 | +/- | -NaN |
| 0.66 | +/- | 0. |
| 0.66 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.08 | +/- | 0. |
| 8.82 | +/- | 0. |
| 0.95 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.11 |
| -9999.00 |
PERSIST_LOW
160-45
| 4870. | +/- | 6. |
| 4930. | +/- | 91. |
| 2.77 | +/- | 0. |
| 2.43 | +/- | 0. |
| 1.48 | +/- | 0. |
| 1.48 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -0.35 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.08 | +/- | 0. |
| 3.62 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.34 |
| -9999.00 |

PERSIST_LOW
STAR_WARN,SN_WARN
160-45
| 4537. | +/- | 8. |
| 4623. | +/- | 112. |
| 4.35 | +/- | 0. |
| 4.51 | +/- | 0. |
| 0.42 | +/- | 0. |
| 0.42 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.09 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.12 |
| -9999.00 |

BRIGHT_NEIGHBOR
160-45
| 5662. | +/- | 21. |
| 5625. | +/- | 133. |
| 4.42 | +/- | 0. |
| 4.44 | +/- | 0. |
| 0.76 | +/- | 0. |
| 0.76 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -0.21 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.74 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.39 |
| -9999.00 |

BRIGHT_NEIGHBOR,PERSIST_LOW
STAR_WARN,SN_WARN
160-45
| 5602. | +/- | 41. |
| 5570. | +/- | 157. |
| 4.15 | +/- | 0. |
| 4.16 | +/- | 0. |
| 0.88 | +/- | 0. |
| 0.88 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.17 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 6.16 | +/- | 0. |
| 0.79 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.05 |
| -9999.00 |

PERSIST_LOW
160-45
| 5340. | +/- | 8. |
| 5321. | +/- | 124. |
| 4.35 | +/- | 0. |
| 4.24 | +/- | 0. |
| 0.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.05 | +/- | 0. |
| 2.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.38 |
| -9999.00 |
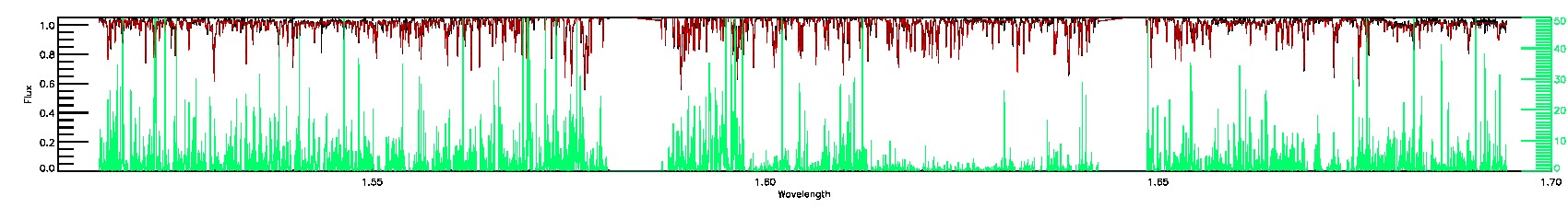
BRIGHT_NEIGHBOR
160-45
| 5722. | +/- | 22. |
| 5663. | +/- | 140. |
| 4.47 | +/- | 0. |
| 4.34 | +/- | 0. |
| 0.97 | +/- | 0. |
| 0.97 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 3.84 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.06 |
| -9999.00 |
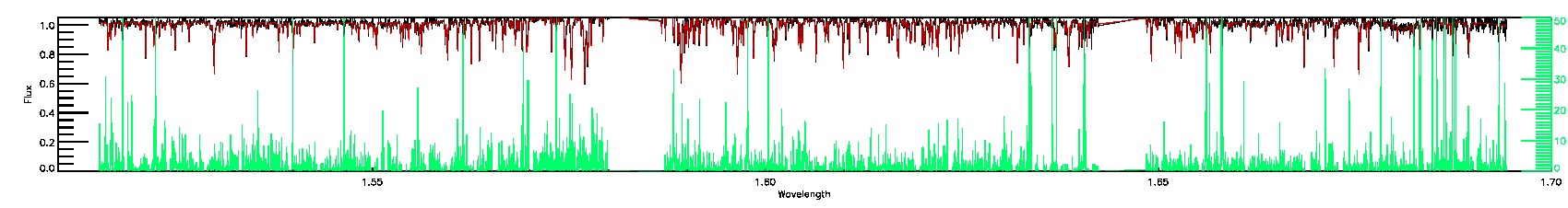
PERSIST_LOW
160-45
| 4850. | +/- | 5. |
| 4913. | +/- | 93. |
| 2.72 | +/- | 0. |
| 2.40 | +/- | 0. |
| 1.55 | +/- | 0. |
| 1.55 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -0.35 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.06 | +/- | 0. |
| 3.62 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.54 |
| -9999.00 |

160-45
| 4593. | +/- | 3. |
| 4676. | +/- | 82. |
| 2.67 | +/- | 0. |
| 2.47 | +/- | 0. |
| 1.24 | +/- | 0. |
| 1.24 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -0.14 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.21 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.28 |
| -9999.00 |

BRIGHT_NEIGHBOR
160-45
| 4928. | +/- | 11. |
| 4980. | +/- | 110. |
| 3.30 | +/- | 0. |
| 3.17 | +/- | 0. |
| 0.72 | +/- | 0. |
| 0.72 | +/- | -NaN |
| -0.33 | +/- | 0. |
| -0.33 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.07 | +/- | 0. |
| 3.58 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.19 |
| -9999.00 |
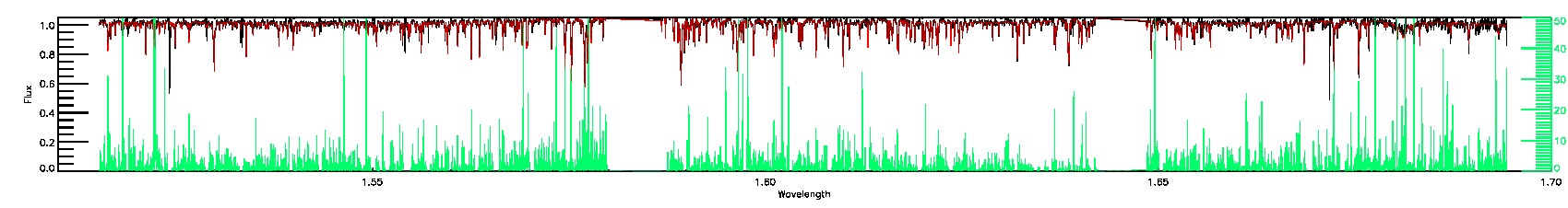
STAR_WARN,COLORTE_WARN
160-45
| 3740. | +/- | 3. |
| 3830. | +/- | 75. |
| 4.45 | +/- | 0. |
| 4.76 | +/- | 0. |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 4.93 | +/- | 0. |
| 0.69 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
160-45
| 3788. | +/- | 4. |
| 3882. | +/- | 79. |
| 4.43 | +/- | 0. |
| 4.76 | +/- | 0. |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.13 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 3.97 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.20 |
| -9999.00 |
PERSIST_LOW
STAR_BAD
STAR_WARN,COLORTE_WARN,SN_WARN
160-45
| 3330. | +/- | 5. |
| -10000. | +/- | -NaN |
| 4.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.83 | +/- | 0. |
| 0.83 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.95 | +/- | 0. |
| 0.77 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.24 |
| -9999.00 |
BRIGHT_NEIGHBOR,PERSIST_LOW,SUSPECT_BROAD_LINES
160-45
| 6610. | +/- | 22. |
| 6465. | +/- | 184. |
| 4.29 | +/- | 0. |
| 4.15 | +/- | 0. |
| 1.39 | +/- | 0. |
| 1.39 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -0.28 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.09 | +/- | 0. |
| 18.95 | +/- | 0. |
| 1.28 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| 0.52 |
| -9999.00 |
PERSIST_LOW
STAR_WARN,SN_WARN
160-45
| 5526. | +/- | 25. |
| 5499. | +/- | 150. |
| 4.39 | +/- | 0. |
| 4.38 | +/- | 0. |
| 1.20 | +/- | 0. |
| 1.20 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 1.67 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 0.23 |
| -9999.00 |

PERSIST_LOW
160-45
| 5187. | +/- | 10. |
| 5194. | +/- | 125. |
| 4.58 | +/- | 0. |
| 4.58 | +/- | 0. |
| 0.83 | +/- | 0. |
| 0.83 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.04 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 1.55 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.19 |
| -9999.00 |
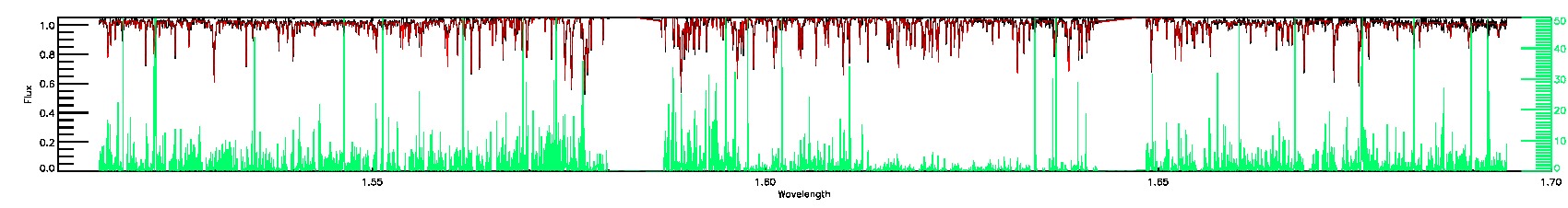
PERSIST_LOW
160-45
| 6483. | +/- | 17. |
| 6342. | +/- | 142. |
| 4.67 | +/- | 0. |
| 4.45 | +/- | 0. |
| 0.77 | +/- | 0. |
| 0.77 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.02 | +/- | 0. |
| 11.09 | +/- | 0. |
| 1.04 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.02 |
| -9999.00 |
PERSIST_LOW,SUSPECT_BROAD_LINES
STAR_BAD
160-45
| 5501. | +/- | 0. |
| -10000. | +/- | -NaN |
| 3.60 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.31 | +/- | 0. |
| 1.31 | +/- | -NaN |
| -0.55 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 84.10 | +/- | 0. |
| 1.92 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.40 |
| -9999.00 |
160-45
| 4432. | +/- | 2. |
| 4533. | +/- | 80. |
| 2.17 | +/- | 0. |
| 1.97 | +/- | 0. |
| 1.57 | +/- | 0. |
| 1.57 | +/- | -NaN |
| -0.30 | +/- | 0. |
| -0.30 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.53 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.97 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,ROTATION_BAD
STAR_WARN,ROTATION_WARN
160-45
| 4569. | +/- | 4. |
| -10000. | +/- | -NaN |
| 3.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| -0.77 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.64 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.35 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.50 |
| -9999.00 |

160-45
| 4647. | +/- | 6. |
| 4712. | +/- | 96. |
| 4.52 | +/- | 0. |
| 4.57 | +/- | 0. |
| 0.50 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.07 | +/- | 0. |
| 4.60 | +/- | 0. |
| 0.66 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.15 |
| -9999.00 |
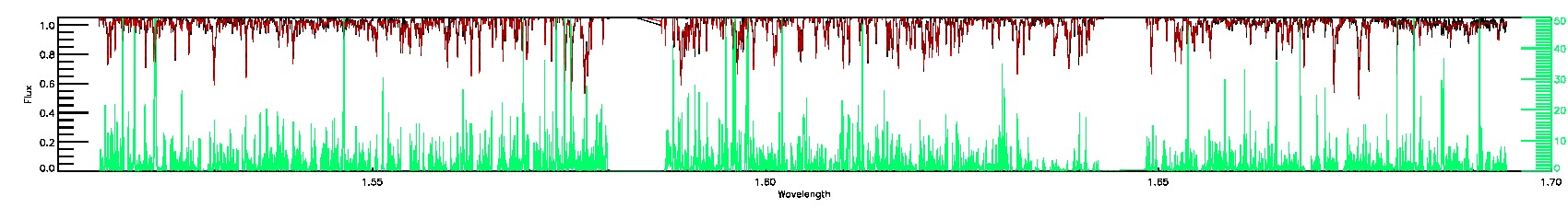
160-45
| 4645. | +/- | 4. |
| 4713. | +/- | 81. |
| 4.37 | +/- | 0. |
| 4.46 | +/- | 0. |
| 0.54 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.08 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.06 | +/- | 0. |
| 3.66 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.17 |
| -9999.00 |

PERSIST_LOW
160-45
| 4424. | +/- | 2. |
| 4502. | +/- | 75. |
| 2.78 | +/- | 0. |
| 2.53 | +/- | 0. |
| 1.13 | +/- | 0. |
| 1.13 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.55 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| 0.30 |
| -9999.00 |

PERSIST_LOW
STAR_WARN,SN_WARN
160-45
| 5632. | +/- | 41. |
| 5609. | +/- | 157. |
| 4.47 | +/- | 0. |
| 4.60 | +/- | 0. |
| 1.01 | +/- | 0. |
| 1.01 | +/- | -NaN |
| -0.46 | +/- | 0. |
| -0.46 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 11.32 | +/- | 0. |
| 1.05 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.45 |
| -9999.00 |

PERSIST_MED,PERSIST_LOW
160-45
| 4427. | +/- | 2. |
| 4532. | +/- | 81. |
| 2.37 | +/- | 0. |
| 2.19 | +/- | 0. |
| 1.38 | +/- | 0. |
| 1.38 | +/- | -NaN |
| -0.39 | +/- | 0. |
| -0.39 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.12 | +/- | 0. |
| 3.71 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.75 |
| -9999.00 |

160-45
| 3849. | +/- | 5. |
| 3942. | +/- | 85. |
| 4.37 | +/- | 0. |
| 4.68 | +/- | 0. |
| 0.60 | +/- | 0. |
| 0.60 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.17 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 5.48 | +/- | 0. |
| 0.74 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.08 |
| -9999.00 |
PERSIST_LOW
160-45
| 4708. | +/- | 6. |
| 4785. | +/- | 104. |
| 2.83 | +/- | 0. |
| 2.66 | +/- | 0. |
| 1.31 | +/- | 0. |
| 1.31 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -0.29 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.07 | +/- | 0. |
| 3.50 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.47 |
| -9999.00 |

160-45
| 4766. | +/- | 4. |
| 4811. | +/- | 82. |
| 3.71 | +/- | 0. |
| 3.60 | +/- | 0. |
| 0.83 | +/- | 0. |
| 0.83 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.51 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.29 |
| -9999.00 |

160-45
| 5714. | +/- | 11. |
| 5657. | +/- | 111. |
| 3.87 | +/- | 0. |
| 3.74 | +/- | 0. |
| 1.35 | +/- | 0. |
| 1.35 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.12 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 6.70 | +/- | 0. |
| 0.83 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.24 |
| -9999.00 |
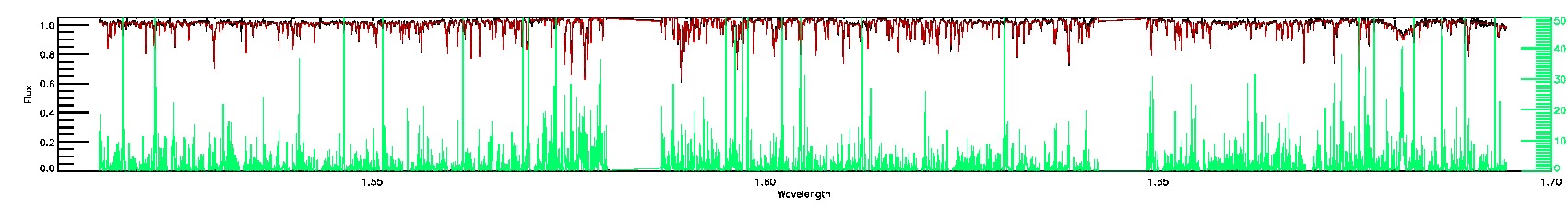
160-45
| 5583. | +/- | 17. |
| 5539. | +/- | 133. |
| 4.46 | +/- | 0. |
| 4.34 | +/- | 0. |
| 0.73 | +/- | 0. |
| 0.73 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.16 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 4.06 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.14 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.14 |
| -9999.00 |
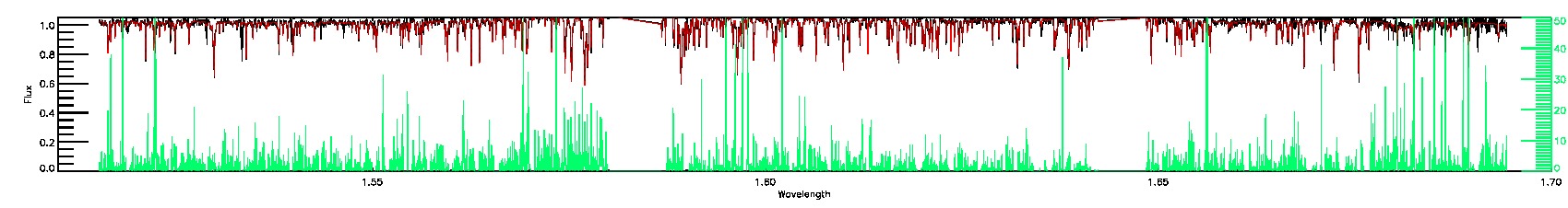
PERSIST_LOW
STAR_WARN,COLORTE_WARN
160-45
| 3687. | +/- | 3. |
| 3778. | +/- | 75. |
| 4.47 | +/- | 0. |
| 4.80 | +/- | 0. |
| 0.47 | +/- | 0. |
| 0.47 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.32 | +/- | 0. |
| -0.32 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.05 | +/- | 0. |
| 2.44 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.41 |
| -9999.00 |
PERSIST_LOW
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
160-45
| 3896. | +/- | 12. |
| -10000. | +/- | -NaN |
| 4.24 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.73 | +/- | 0. |
| 0.73 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.32 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -1.28 |
| -9999.00 |
160-45
| 5966. | +/- | 14. |
| 5882. | +/- | 130. |
| 4.36 | +/- | 0. |
| 4.21 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.43 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.07 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.02 | +/- | 0. |
| 5.45 | +/- | 0. |
| 0.74 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -1.10 |
| -9999.00 |
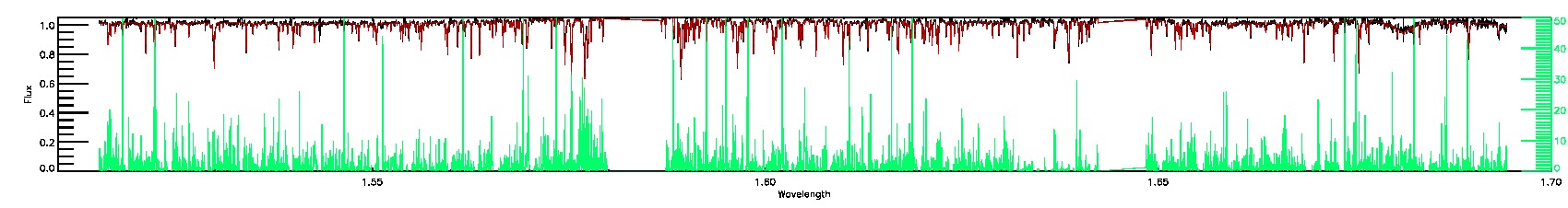
BRIGHT_NEIGHBOR,PERSIST_LOW
160-45
| 4658. | +/- | 5. |
| 4771. | +/- | 105. |
| 2.37 | +/- | 0. |
| 2.27 | +/- | 0. |
| 1.52 | +/- | 0. |
| 1.52 | +/- | -NaN |
| -1.00 | +/- | 0. |
| -1.00 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.28 | +/- | 0. |
| 5.31 | +/- | 0. |
| 0.73 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| -1.82 |
| -9999.00 |

BRIGHT_NEIGHBOR,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES
160-45
| 5930. | +/- | 23. |
| 5863. | +/- | 150. |
| 4.05 | +/- | 0. |
| 4.03 | +/- | 0. |
| 1.70 | +/- | 0. |
| 1.70 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -0.26 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.10 | +/- | 0. |
| 29.80 | +/- | 0. |
| 1.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.06 |
| -9999.00 |
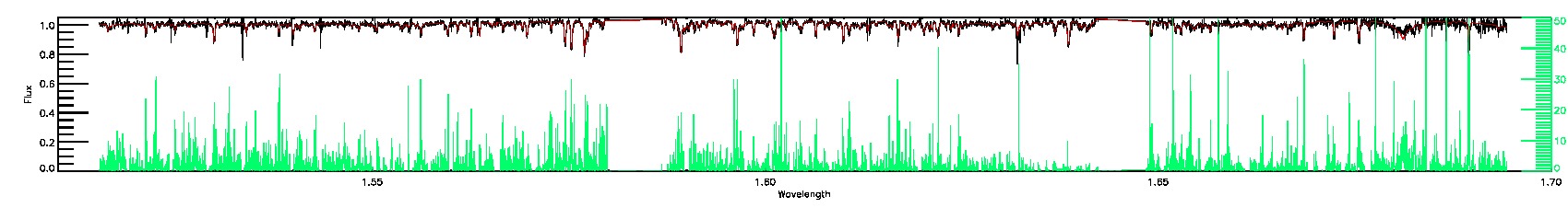
160-45
| 5735. | +/- | 14. |
| 5684. | +/- | 115. |
| 4.52 | +/- | 0. |
| 4.47 | +/- | 0. |
| 0.92 | +/- | 0. |
| 0.92 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.09 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.50 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |

160-45
| 5627. | +/- | 9. |
| 5574. | +/- | 107. |
| 4.17 | +/- | 0. |
| 4.01 | +/- | 0. |
| 0.68 | +/- | 0. |
| 0.68 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 3.86 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.19 |
| -9999.00 |

BRIGHT_NEIGHBOR,PERSIST_MED,PERSIST_LOW
160-45
| 5859. | +/- | 12. |
| 5791. | +/- | 137. |
| 3.99 | +/- | 0. |
| 3.88 | +/- | 0. |
| 1.50 | +/- | 0. |
| 1.50 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 5.25 | +/- | 0. |
| 0.72 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.11 |
| -9999.00 |

160-45
| 5347. | +/- | 9. |
| 5324. | +/- | 111. |
| 4.58 | +/- | 0. |
| 4.44 | +/- | 0. |
| 0.57 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.32 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 3.03 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.46 |
| -9999.00 |

160-45
| 4868. | +/- | 9. |
| 4917. | +/- | 104. |
| 4.53 | +/- | 0. |
| 4.65 | +/- | 0. |
| 1.09 | +/- | 0. |
| 1.09 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 1.88 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.21 |
| -9999.00 |

BRIGHT_NEIGHBOR,PERSIST_MED,PERSIST_LOW
160-45
| 5088. | +/- | 10. |
| 5116. | +/- | 121. |
| 3.36 | +/- | 0. |
| 3.04 | +/- | 0. |
| 1.21 | +/- | 0. |
| 1.21 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -0.18 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.06 | +/- | 0. |
| 3.29 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.36 |
| -9999.00 |
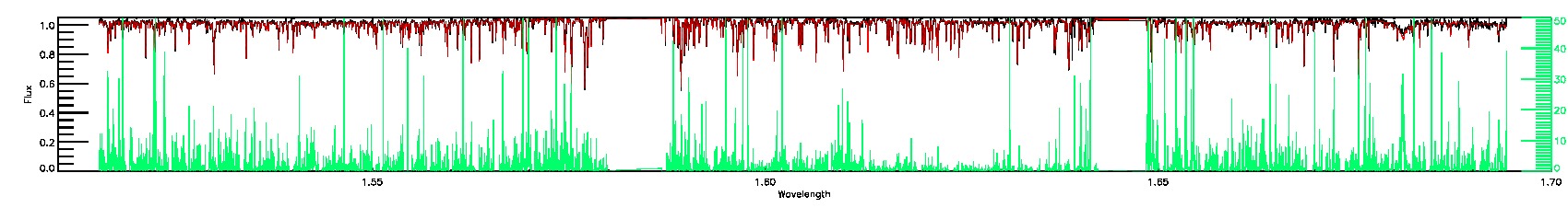
PERSIST_LOW
STAR_WARN,SN_WARN
160-45
| 4733. | +/- | 15. |
| 4813. | +/- | 126. |
| 2.64 | +/- | 0. |
| 2.35 | +/- | 0. |
| 1.38 | +/- | 0. |
| 1.38 | +/- | -NaN |
| -0.45 | +/- | 0. |
| -0.45 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.09 | +/- | 0. |
| 3.84 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.15 |
| -9999.00 |

160-45
| 4895. | +/- | 8. |
| 4940. | +/- | 102. |
| 3.71 | +/- | 0. |
| 3.67 | +/- | 0. |
| 0.64 | +/- | 0. |
| 0.64 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.12 | +/- | 0. |
| 3.05 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.32 |
| -9999.00 |
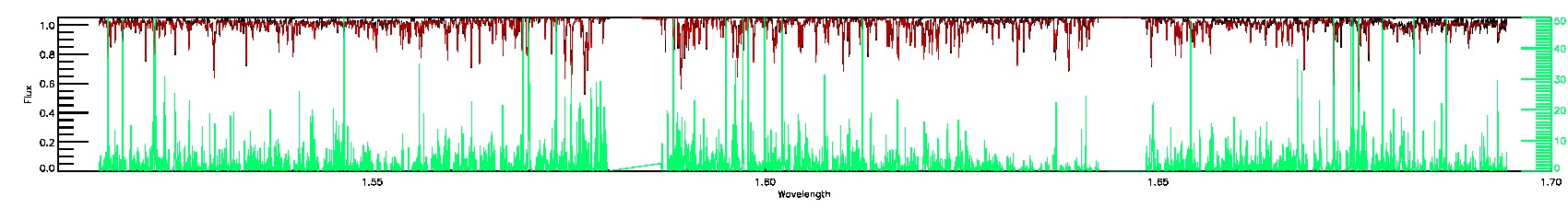
BRIGHT_NEIGHBOR,PERSIST_LOW
160-45
| 5361. | +/- | 9. |
| 5346. | +/- | 108. |
| 4.54 | +/- | 0. |
| 4.49 | +/- | 0. |
| 0.67 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.08 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.07 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.13 |
| -9999.00 |

160-45
| 5958. | +/- | 19. |
| 5889. | +/- | 137. |
| 4.42 | +/- | 0. |
| 4.40 | +/- | 0. |
| 0.88 | +/- | 0. |
| 0.88 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -0.28 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.17 | +/- | 0. |
| 5.03 | +/- | 0. |
| 0.70 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.06 |
| -9999.00 |
160-45
| 5041. | +/- | 9. |
| 5090. | +/- | 112. |
| 2.76 | +/- | 0. |
| 2.43 | +/- | 0. |
| 1.71 | +/- | 0. |
| 1.71 | +/- | -NaN |
| -0.55 | +/- | 0. |
| -0.55 | +/- | 0. |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.09 | +/- | 0. |
| 4.07 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.48 |
| -9999.00 |
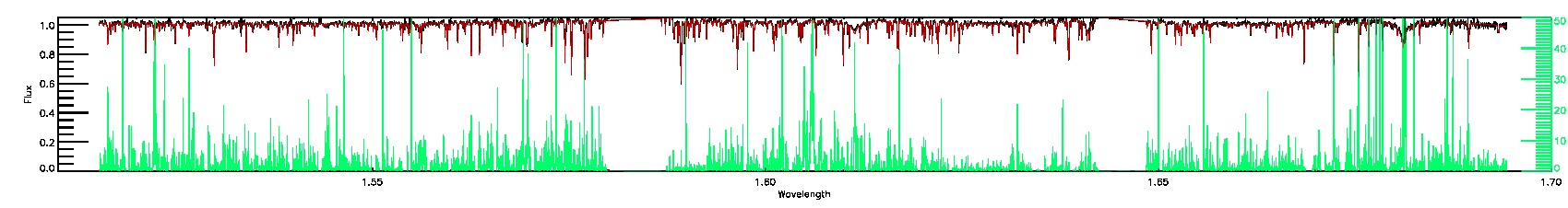
PERSIST_LOW
160-45
| 5807. | +/- | 13. |
| 5753. | +/- | 119. |
| 4.25 | +/- | 0. |
| 4.25 | +/- | 0. |
| 1.18 | +/- | 0. |
| 1.18 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -0.23 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.13 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.01 | +/- | 0. |
| 4.10 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.08 |
| -9999.00 |

160-45
| 5534. | +/- | 12. |
| 5501. | +/- | 114. |
| 4.00 | +/- | 0. |
| 3.94 | +/- | 0. |
| 1.19 | +/- | 0. |
| 1.19 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.03 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.91 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.21 |
| -9999.00 |

SUSPECT_BROAD_LINES
160-45
| 6257. | +/- | 16. |
| 6139. | +/- | 131. |
| 4.07 | +/- | 0. |
| 3.86 | +/- | 0. |
| 1.74 | +/- | 0. |
| 1.74 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.07 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 30.74 | +/- | 0. |
| 1.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.36 |
| -9999.00 |
160-45
| 5657. | +/- | 15. |
| 5618. | +/- | 119. |
| 4.51 | +/- | 0. |
| 4.50 | +/- | 0. |
| 1.36 | +/- | 0. |
| 1.36 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.76 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.01 |
| -9999.00 |
PERSIST_LOW
160-45
| 6104. | +/- | 15. |
| 6003. | +/- | 125. |
| 4.29 | +/- | 0. |
| 4.10 | +/- | 0. |
| 1.69 | +/- | 0. |
| 1.69 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 11.59 | +/- | 0. |
| 1.06 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.63 |
| -9999.00 |
PERSIST_MED,PERSIST_LOW
160-45
| 4826. | +/- | 12. |
| 4903. | +/- | 123. |
| 2.88 | +/- | 0. |
| 2.76 | +/- | 0. |
| 1.50 | +/- | 0. |
| 1.50 | +/- | -NaN |
| -0.62 | +/- | 0. |
| -0.62 | +/- | 0. |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.24 | +/- | 0. |
| 4.25 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -1.54 |
| -9999.00 |

SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES
160-45
| 6476. | +/- | 17. |
| 6332. | +/- | 140. |
| 3.99 | +/- | 0. |
| 3.73 | +/- | 0. |
| 0.54 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.15 | +/- | 0. |
| 67.17 | +/- | 0. |
| 1.83 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.20 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -2.14 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.42 |
| -9999.00 |
160-45
| 5947. | +/- | 14. |
| 5872. | +/- | 128. |
| 4.28 | +/- | 0. |
| 4.20 | +/- | 0. |
| 1.45 | +/- | 0. |
| 1.45 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.00 | +/- | 0. |
| 5.12 | +/- | 0. |
| 0.71 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.19 |
| -9999.00 |
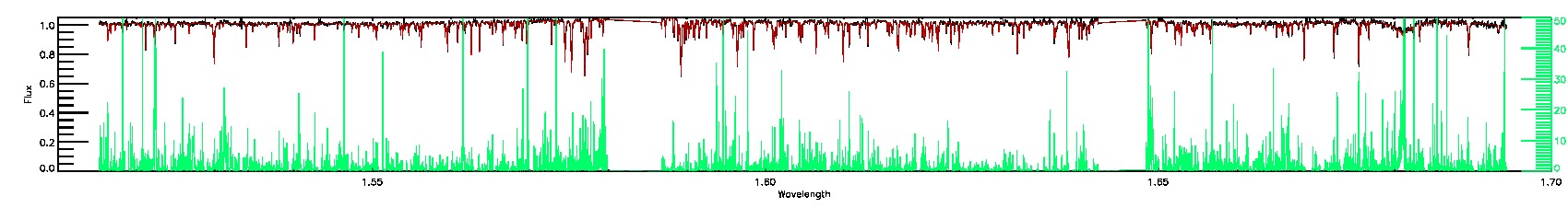
160-45
| 4699. | +/- | 9. |
| 4760. | +/- | 107. |
| 4.40 | +/- | 0. |
| 4.47 | +/- | 0. |
| 0.49 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 3.41 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -2.44 |
| -9999.00 |
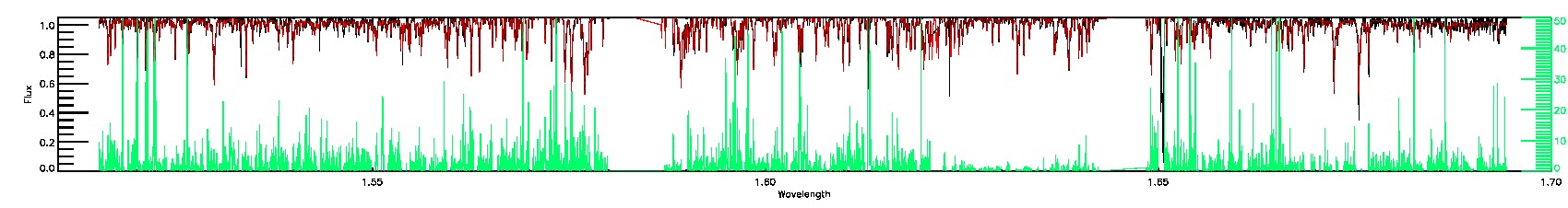
160-45
| 5318. | +/- | 23. |
| 5322. | +/- | 134. |
| 4.48 | +/- | 0. |
| 4.58 | +/- | 0. |
| 0.84 | +/- | 0. |
| 0.84 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -0.26 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.85 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.13 |
| -9999.00 |
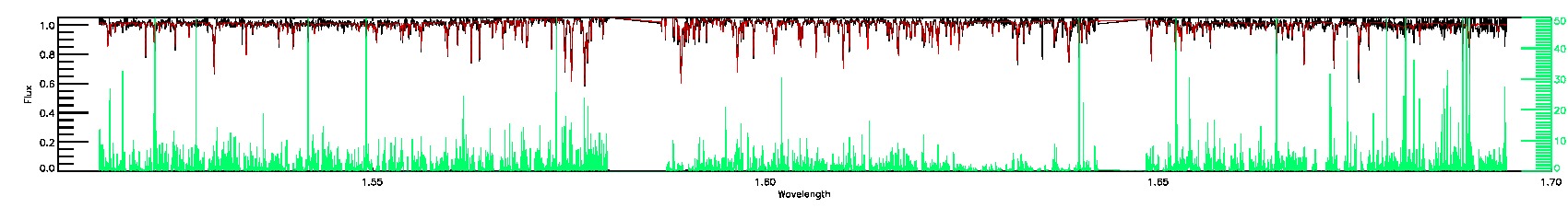
160-45
| 5755. | +/- | 19. |
| 5719. | +/- | 123. |
| 4.47 | +/- | 0. |
| 4.58 | +/- | 0. |
| 0.90 | +/- | 0. |
| 0.90 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -0.49 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 4.30 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -0.32 |
| -9999.00 |

160-45
| 5523. | +/- | 15. |
| 5507. | +/- | 111. |
| 4.50 | +/- | 0. |
| 4.59 | +/- | 0. |
| 1.41 | +/- | 0. |
| 1.41 | +/- | -NaN |
| -0.34 | +/- | 0. |
| -0.33 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 4.23 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.31 |
| -9999.00 |

160-45
| 5680. | +/- | 21. |
| 5627. | +/- | 140. |
| 4.47 | +/- | 0. |
| 4.35 | +/- | 0. |
| 0.71 | +/- | 0. |
| 0.71 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | 0. |
| -0.14 | +/- | 0. |
| -0.14 | +/- | -NaN |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.79 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |

160-45
| 6288. | +/- | 15. |
| 6165. | +/- | 132. |
| 4.27 | +/- | 0. |
| 4.04 | +/- | 0. |
| 1.41 | +/- | 0. |
| 1.41 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -0.14 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 11.14 | +/- | 0. |
| 1.05 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.50 |
| -9999.00 |
PERSIST_MED
160-45
| 4739. | +/- | 7. |
| 4808. | +/- | 112. |
| 4.33 | +/- | 0. |
| 4.51 | +/- | 0. |
| 0.45 | +/- | 0. |
| 0.45 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -0.19 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 2.46 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.32 |
| -9999.00 |

PERSIST_MED,SUSPECT_BROAD_LINES
160-45
| 6965. | +/- | 18. |
| 6772. | +/- | 166. |
| 4.71 | +/- | 0. |
| 4.44 | +/- | 0. |
| 0.82 | +/- | 0. |
| 0.82 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.12 | +/- | 0. |
| 26.57 | +/- | 0. |
| 1.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.16 |
| -9999.00 |
160-45
| 4410. | +/- | 3. |
| 4503. | +/- | 83. |
| 4.32 | +/- | 0. |
| 4.53 | +/- | 0. |
| 0.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 6.44 | +/- | 0. |
| 0.81 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
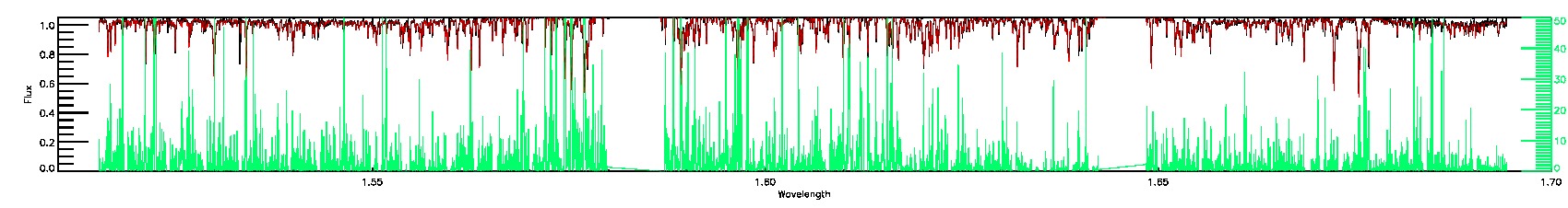
BRIGHT_NEIGHBOR,PERSIST_MED,PERSIST_LOW
160-45
| 5410. | +/- | 11. |
| 5386. | +/- | 125. |
| 4.40 | +/- | 0. |
| 4.31 | +/- | 0. |
| 0.44 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.15 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 3.07 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.23 |
| -9999.00 |

PERSIST_MED
160-45
| 4589. | +/- | 3. |
| 4667. | +/- | 81. |
| 2.85 | +/- | 0. |
| 2.64 | +/- | 0. |
| 1.18 | +/- | 0. |
| 1.18 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.96 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| -0.21 |
| -9999.00 |
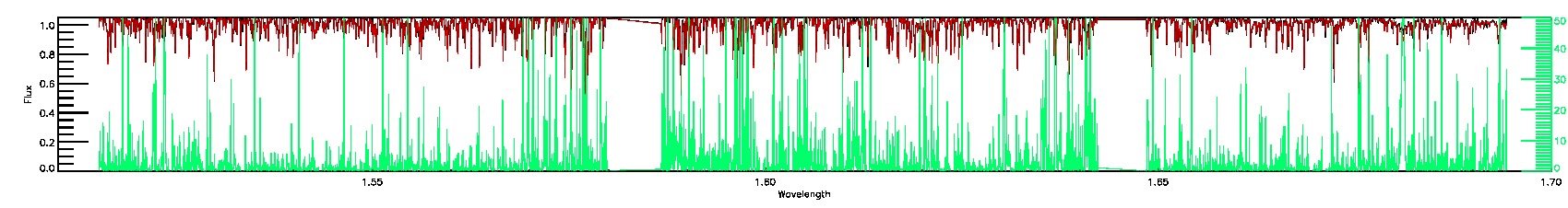
160-45
| 5405. | +/- | 14. |
| 5401. | +/- | 111. |
| 4.39 | +/- | 0. |
| 4.49 | +/- | 0. |
| 0.85 | +/- | 0. |
| 0.85 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -0.29 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.58 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.28 |
| -9999.00 |
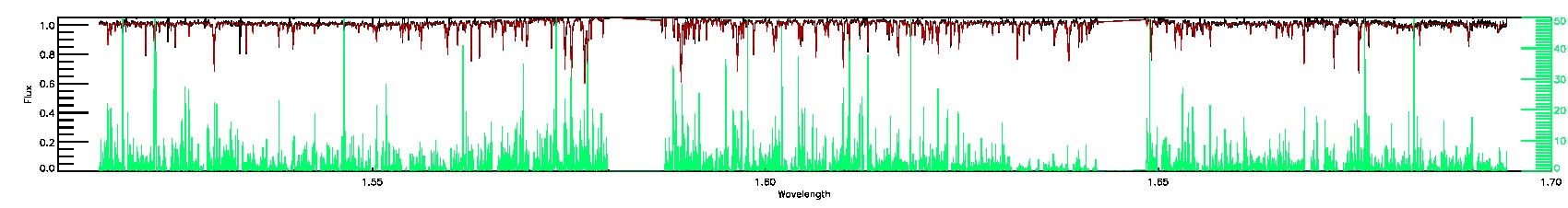
160-45
| 5594. | +/- | 12. |
| 5566. | +/- | 120. |
| 4.50 | +/- | 0. |
| 4.55 | +/- | 0. |
| 0.47 | +/- | 0. |
| 0.47 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -0.25 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 1.72 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.64 |
| -9999.00 |

160-45
| 4772. | +/- | 4. |
| 4824. | +/- | 84. |
| 4.37 | +/- | 0. |
| 4.42 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.80 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.06 |
| -9999.00 |

PERSIST_LOW
160-45
| 5154. | +/- | 9. |
| 5178. | +/- | 104. |
| 3.74 | +/- | 0. |
| 3.75 | +/- | 0. |
| 1.35 | +/- | 0. |
| 1.35 | +/- | -NaN |
| -0.27 | +/- | 0. |
| -0.27 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.46 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.10 |
| -9999.00 |

160-45
| 5485. | +/- | 16. |
| 5460. | +/- | 136. |
| 4.39 | +/- | 0. |
| 4.36 | +/- | 0. |
| 0.43 | +/- | 0. |
| 0.43 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 5.33 | +/- | 0. |
| 0.73 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.31 |
| -9999.00 |
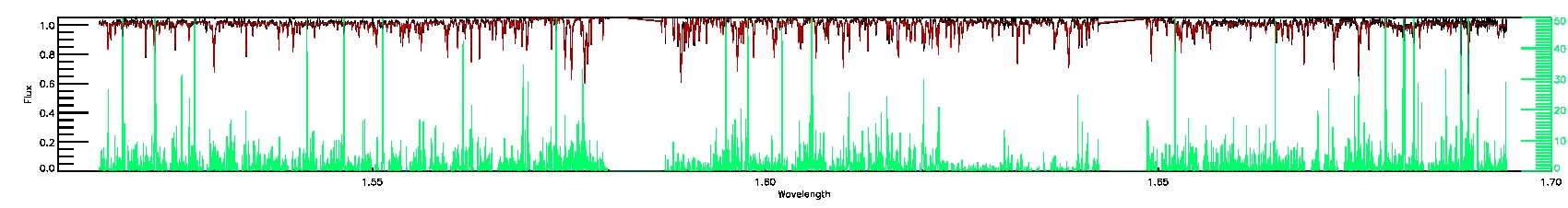
160-45
| 3967. | +/- | 2. |
| 4058. | +/- | 71. |
| 4.46 | +/- | 0. |
| 4.72 | +/- | 0. |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.17 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 3.05 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.13 |
| -9999.00 |
PERSIST_LOW
160-45
| 5035. | +/- | 7. |
| 5066. | +/- | 93. |
| 3.69 | +/- | 0. |
| 3.64 | +/- | 0. |
| 1.16 | +/- | 0. |
| 1.16 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.14 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.21 |
| -9999.00 |

PERSIST_LOW
160-45
| 5406. | +/- | 8. |
| 5383. | +/- | 111. |
| 4.37 | +/- | 0. |
| 4.28 | +/- | 0. |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.15 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 3.66 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.22 |
| -9999.00 |

160-45
| 5129. | +/- | 11. |
| 5148. | +/- | 113. |
| 4.44 | +/- | 0. |
| 4.50 | +/- | 0. |
| 0.47 | +/- | 0. |
| 0.47 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.08 | +/- | 0. |
| 1.89 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.01 |
| -9999.00 |
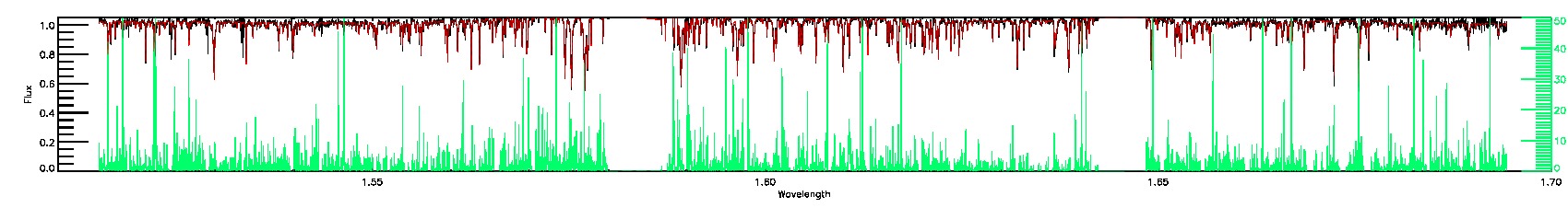
PERSIST_MED
160-45
| 5634. | +/- | 11. |
| 5593. | +/- | 111. |
| 4.53 | +/- | 0. |
| 4.49 | +/- | 0. |
| 0.67 | +/- | 0. |
| 0.67 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 7.35 | +/- | 0. |
| 0.87 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.20 |
| -9999.00 |
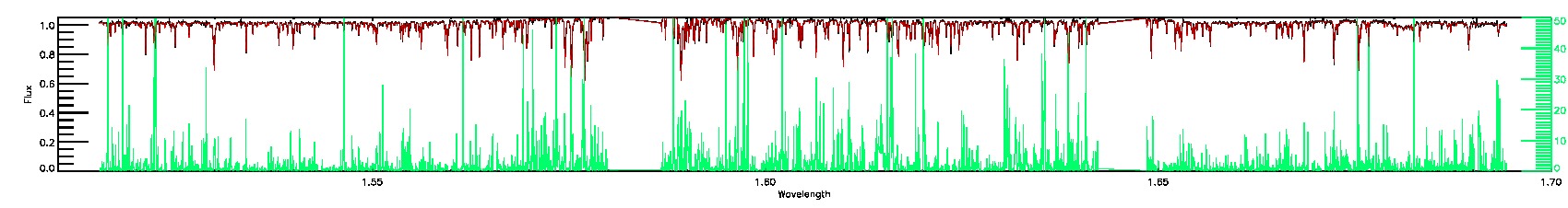
PERSIST_MED
160-45
| 5329. | +/- | 13. |
| 5328. | +/- | 125. |
| 3.78 | +/- | 0. |
| 3.75 | +/- | 0. |
| 1.45 | +/- | 0. |
| 1.45 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 3.23 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.08 |
| -9999.00 |
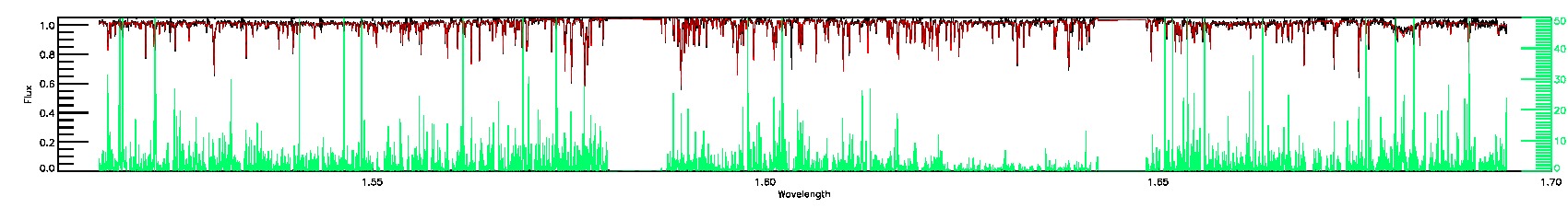
160-45
| 4435. | +/- | 6. |
| 4526. | +/- | 101. |
| 4.37 | +/- | 0. |
| 4.56 | +/- | 0. |
| 0.61 | +/- | 0. |
| 0.61 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.25 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.35 |
| -9999.00 |

PERSIST_LOW
160-45
| 4771. | +/- | 5. |
| 4847. | +/- | 97. |
| 2.78 | +/- | 0. |
| 2.44 | +/- | 0. |
| 1.27 | +/- | 0. |
| 1.27 | +/- | -NaN |
| -0.43 | +/- | 0. |
| -0.43 | +/- | 0. |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.19 | +/- | 0. |
| 3.81 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.20 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.78 |
| -9999.00 |
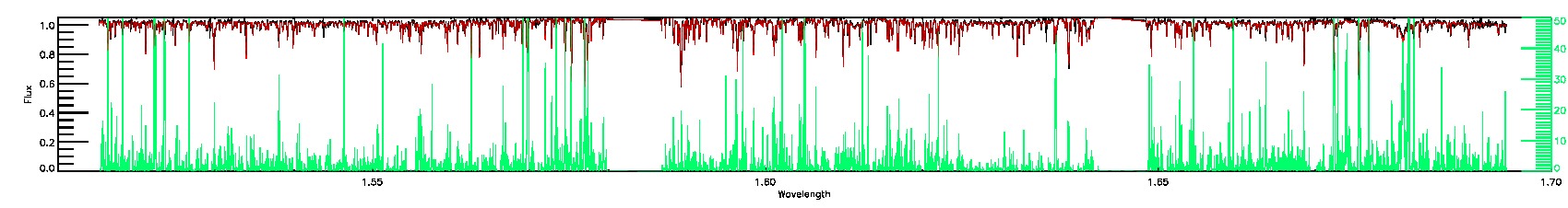
160-45
| 5856. | +/- | 16. |
| 5798. | +/- | 121. |
| 4.35 | +/- | 0. |
| 4.35 | +/- | 0. |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| -0.27 | +/- | 0. |
| -0.27 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 2.71 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.32 |
| -9999.00 |
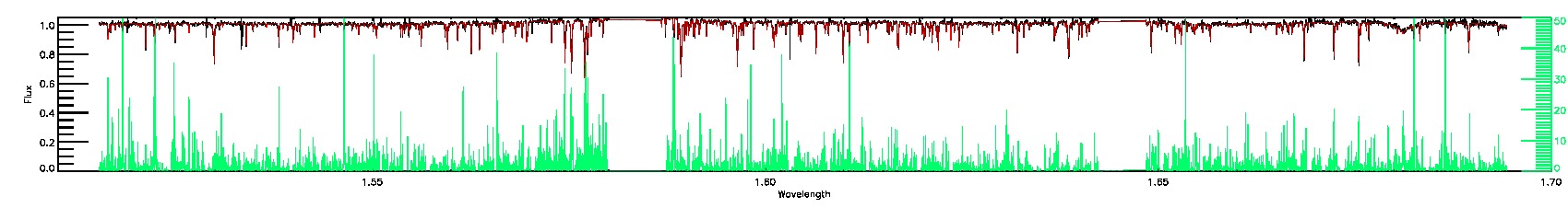
PERSIST_LOW
160-45
| 5754. | +/- | 13. |
| 5696. | +/- | 137. |
| 4.48 | +/- | 0. |
| 4.38 | +/- | 0. |
| 0.94 | +/- | 0. |
| 0.94 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.13 | +/- | 0. |
| 0.33 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.00 |
| -9999.00 |
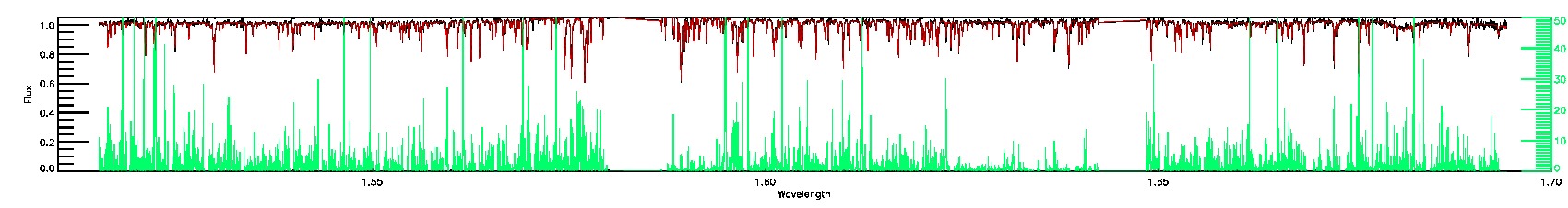
BRIGHT_NEIGHBOR
160-45
| 4764. | +/- | 7. |
| 4844. | +/- | 108. |
| 2.57 | +/- | 0. |
| 2.31 | +/- | 0. |
| 1.58 | +/- | 0. |
| 1.58 | +/- | -NaN |
| -0.52 | +/- | 0. |
| -0.52 | +/- | 0. |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.25 | +/- | 0. |
| 4.00 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.18 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD
160-45
| 7410. | +/- | 10. |
| -10000. | +/- | -NaN |
| 3.98 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.09 | +/- | 0. |
| 4.09 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.45 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 29.42 | +/- | 0. |
| 1.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -1.99 |
| -9999.00 |
SUSPECT_BROAD_LINES
160-45
| 6931. | +/- | 15. |
| 6738. | +/- | 162. |
| 4.52 | +/- | 0. |
| 4.22 | +/- | 0. |
| 0.69 | +/- | 0. |
| 0.69 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.06 | +/- | 0. |
| 27.68 | +/- | 0. |
| 1.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.57 |
| -9999.00 |
PERSIST_LOW
160-45
| 4110. | +/- | 3. |
| 4201. | +/- | 92. |
| 4.41 | +/- | 0. |
| 4.65 | +/- | 0. |
| 0.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.05 | +/- | 0. |
| 3.93 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.11 |
| -9999.00 |
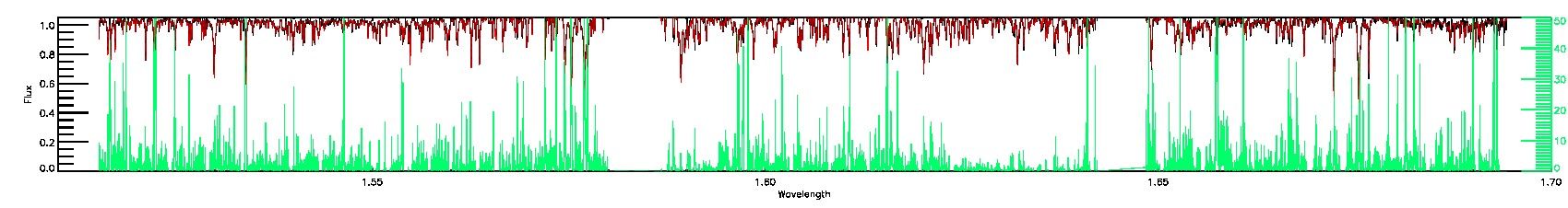
PERSIST_LOW
160-45
| 5843. | +/- | 19. |
| 5775. | +/- | 151. |
| 4.34 | +/- | 0. |
| 4.23 | +/- | 0. |
| 1.15 | +/- | 0. |
| 1.15 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.37 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 4.06 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.99 |
| -9999.00 |

BRIGHT_NEIGHBOR
160-45
| 5767. | +/- | 12. |
| 5701. | +/- | 112. |
| 4.24 | +/- | 0. |
| 4.08 | +/- | 0. |
| 0.98 | +/- | 0. |
| 0.98 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.69 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.19 |
| -9999.00 |
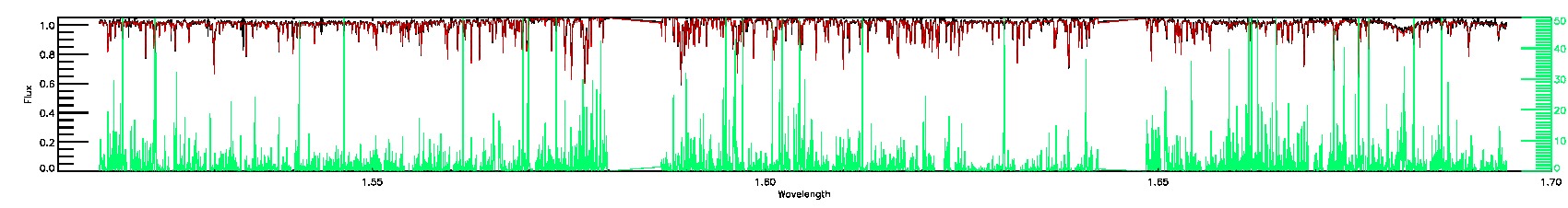
160-45
| 6927. | +/- | 16. |
| 6739. | +/- | 164. |
| 4.53 | +/- | 0. |
| 4.26 | +/- | 0. |
| 1.60 | +/- | 0. |
| 1.60 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.10 | +/- | 0. |
| 7.49 | +/- | 0. |
| 0.87 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.00 |
| -9999.00 |
160-45
| 5898. | +/- | 16. |
| 5822. | +/- | 136. |
| 4.56 | +/- | 0. |
| 4.42 | +/- | 0. |
| 0.89 | +/- | 0. |
| 0.89 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.21 | +/- | 0. |
| -0.21 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 3.01 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.01 |
| -9999.00 |

160-45
| 5736. | +/- | 18. |
| 5688. | +/- | 127. |
| 4.44 | +/- | 0. |
| 4.42 | +/- | 0. |
| 1.26 | +/- | 0. |
| 1.26 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.17 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.15 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.45 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.06 |
| -9999.00 |

160-45
| 6556. | +/- | 17. |
| 6420. | +/- | 150. |
| 4.32 | +/- | 0. |
| 4.21 | +/- | 0. |
| 0.82 | +/- | 0. |
| 0.82 | +/- | -NaN |
| -0.33 | +/- | 0. |
| -0.32 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.08 | +/- | 0. |
| 12.88 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.36 |
| -9999.00 |
BRIGHT_NEIGHBOR
160-45
| 5866. | +/- | 21. |
| 5807. | +/- | 145. |
| 4.44 | +/- | 0. |
| 4.43 | +/- | 0. |
| 0.94 | +/- | 0. |
| 0.94 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -0.26 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 3.10 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| -0.19 |
| -9999.00 |
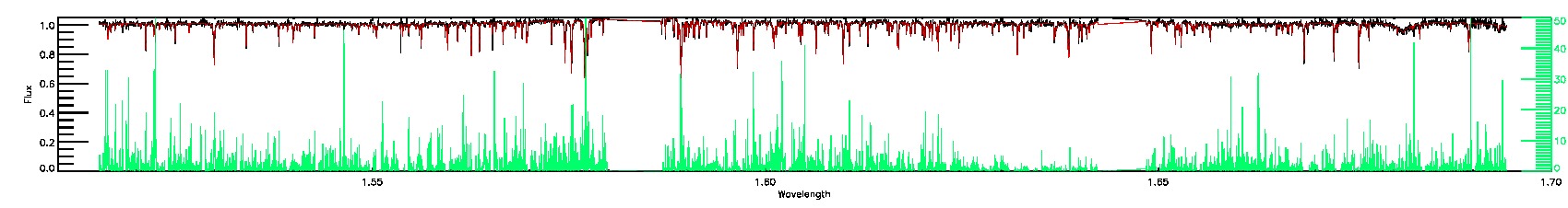
BRIGHT_NEIGHBOR
160-45
| 3940. | +/- | 2. |
| 4031. | +/- | 68. |
| 4.48 | +/- | 0. |
| 4.76 | +/- | 0. |
| 0.41 | +/- | 0. |
| 0.41 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.24 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 7.74 | +/- | 0. |
| 0.89 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
160-45
| 6112. | +/- | 31. |
| 6025. | +/- | 163. |
| 4.49 | +/- | 0. |
| 4.44 | +/- | 0. |
| 0.50 | +/- | 0. |
| 0.50 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -0.27 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 4.55 | +/- | 0. |
| 0.66 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -1.01 |
| -9999.00 |
PERSIST_MED
160-45
| 4986. | +/- | 9. |
| 5015. | +/- | 111. |
| 3.85 | +/- | 0. |
| 3.83 | +/- | 0. |
| 0.67 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.06 | +/- | 0. |
| 2.85 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.15 |
| -9999.00 |

PERSIST_LOW
160-45
| 4522. | +/- | 3. |
| 4627. | +/- | 88. |
| 2.45 | +/- | 0. |
| 2.28 | +/- | 0. |
| 1.46 | +/- | 0. |
| 1.46 | +/- | -NaN |
| -0.45 | +/- | 0. |
| -0.45 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.07 | +/- | 0. |
| 3.84 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.42 |
| -9999.00 |
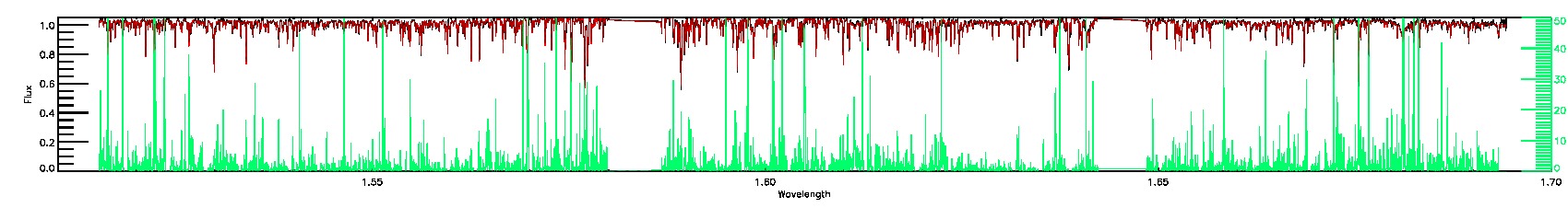
SUSPECT_BROAD_LINES
160-45
| 3932. | +/- | 3. |
| 4021. | +/- | 82. |
| 4.38 | +/- | 0. |
| 4.64 | +/- | 0. |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.07 | +/- | 0. |
| 16.11 | +/- | 0. |
| 1.21 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.00 |
| -9999.00 |
SUSPECT_BROAD_LINES
160-45
| 6622. | +/- | 22. |
| 6468. | +/- | 149. |
| 4.36 | +/- | 0. |
| 4.14 | +/- | 0. |
| 2.04 | +/- | 0. |
| 2.04 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.08 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.07 | +/- | 0. |
| 40.36 | +/- | 0. |
| 1.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.56 |
| -9999.00 |
160-45
| 3607. | +/- | 1. |
| 3737. | +/- | 68. |
| 0.00 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.85 | +/- | 0. |
| 2.85 | +/- | -NaN |
| -0.98 | +/- | 0. |
| -0.97 | +/- | 0. |
| -0.45 | +/- | 0. |
| -0.45 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.07 | +/- | 0. |
| 5.23 | +/- | 0. |
| 0.72 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.45 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -1.96 |
| -9999.00 |
160-45
| 5141. | +/- | 9. |
| 5151. | +/- | 110. |
| 4.49 | +/- | 0. |
| 4.47 | +/- | 0. |
| 0.90 | +/- | 0. |
| 0.90 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.05 | +/- | 0. |
| 1.69 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |

160-45
| 4825. | +/- | 7. |
| 4872. | +/- | 98. |
| 4.53 | +/- | 0. |
| 4.59 | +/- | 0. |
| 1.06 | +/- | 0. |
| 1.06 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 1.51 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.13 |
| -9999.00 |

PERSIST_LOW
160-45
| 4898. | +/- | 7. |
| 4947. | +/- | 108. |
| 4.26 | +/- | 0. |
| 4.41 | +/- | 0. |
| 0.49 | +/- | 0. |
| 0.49 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -0.18 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.17 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 7.73 | +/- | 0. |
| 0.89 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.00 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
160-45
| 4625. | +/- | 6. |
| 4729. | +/- | 97. |
| 3.89 | +/- | 0. |
| 4.21 | +/- | 0. |
| 0.45 | +/- | 0. |
| 0.45 | +/- | -NaN |
| -0.70 | +/- | 0. |
| -0.70 | +/- | 0. |
| -0.26 | +/- | 0. |
| -0.26 | +/- | -NaN |
| -0.33 | +/- | 0. |
| -0.33 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.12 | +/- | 0. |
| 6.45 | +/- | 0. |
| 0.81 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.20 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.43 |
| -9999.00 |

160-45
| 5052. | +/- | 8. |
| 5082. | +/- | 102. |
| 3.54 | +/- | 0. |
| 3.42 | +/- | 0. |
| 1.08 | +/- | 0. |
| 1.08 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.04 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.28 |
| -9999.00 |
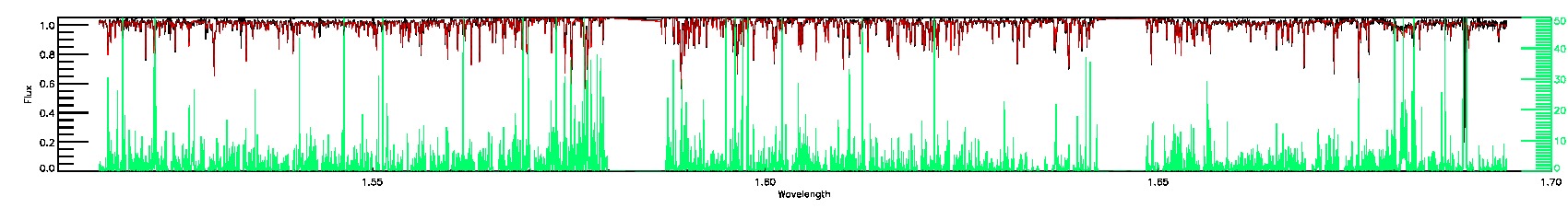
PERSIST_LOW,SUSPECT_BROAD_LINES
160-45
| 4295. | +/- | 4. |
| 4378. | +/- | 94. |
| 4.41 | +/- | 0. |
| 4.54 | +/- | 0. |
| 0.58 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.07 | +/- | 0. |
| 15.05 | +/- | 0. |
| 1.18 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.29 |
| -9999.00 |

160-45
| 5608. | +/- | 16. |
| 5594. | +/- | 119. |
| 3.98 | +/- | 0. |
| 4.16 | +/- | 0. |
| 0.78 | +/- | 0. |
| 0.78 | +/- | -NaN |
| -0.60 | +/- | 0. |
| -0.60 | +/- | 0. |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -0.20 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.14 | +/- | 0. |
| 4.21 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -0.41 |
| -9999.00 |

160-45
| 4733. | +/- | 6. |
| 4825. | +/- | 92. |
| 2.72 | +/- | 0. |
| 2.60 | +/- | 0. |
| 1.50 | +/- | 0. |
| 1.50 | +/- | -NaN |
| -0.73 | +/- | 0. |
| -0.73 | +/- | 0. |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.24 | +/- | 0. |
| 4.53 | +/- | 0. |
| 0.66 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.87 |
| -9999.00 |
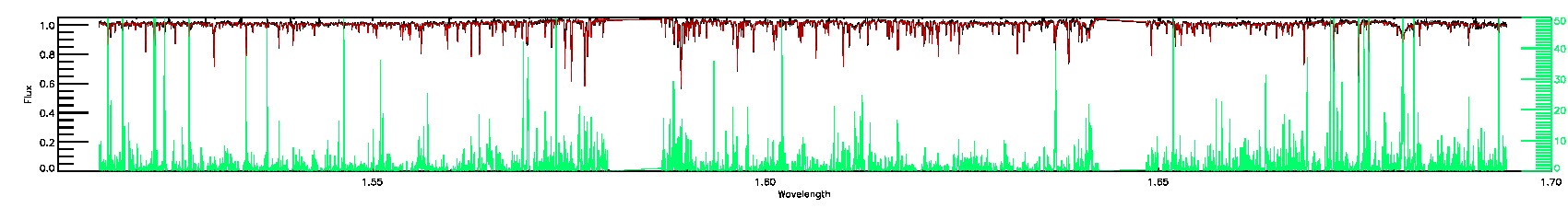
PERSIST_LOW
STAR_BAD
STAR_WARN,COLORTE_WARN
160-45
| 3737. | +/- | 3. |
| -10000. | +/- | -NaN |
| 4.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.81 | +/- | 0. |
| 0.81 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.01 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
160-45
| 5519. | +/- | 30. |
| 5501. | +/- | 142. |
| 4.43 | +/- | 0. |
| 4.49 | +/- | 0. |
| 0.52 | +/- | 0. |
| 0.52 | +/- | -NaN |
| -0.27 | +/- | 0. |
| -0.27 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 4.92 | +/- | 0. |
| 0.69 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.16 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.46 |
| -9999.00 |
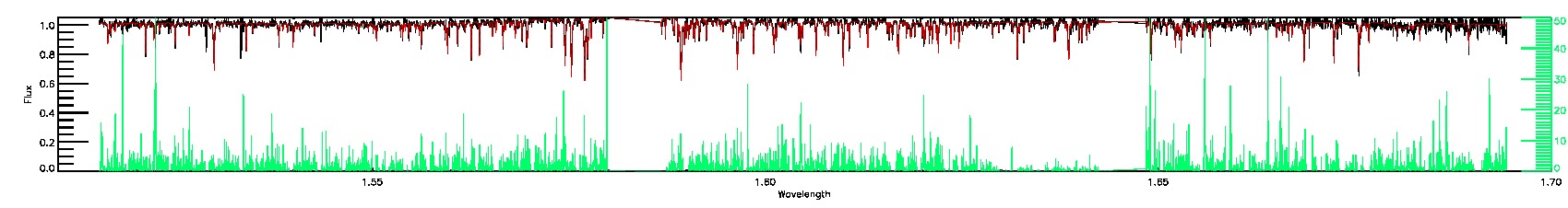
PERSIST_LOW
STAR_WARN,SN_WARN
160-45
| 6113. | +/- | 34. |
| 6023. | +/- | 175. |
| 4.34 | +/- | 0. |
| 4.28 | +/- | 0. |
| 1.00 | +/- | 0. |
| 1.00 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| -0.27 | +/- | 0. |
| -0.27 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 6.30 | +/- | 0. |
| 0.80 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.40 |
| -9999.00 |
SUSPECT_BROAD_LINES
160-45
| 6612. | +/- | 16. |
| 6465. | +/- | 151. |
| 4.22 | +/- | 0. |
| 4.06 | +/- | 0. |
| 2.06 | +/- | 0. |
| 2.06 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.08 | +/- | 0. |
| 15.06 | +/- | 0. |
| 1.18 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.28 |
| -9999.00 |
SUSPECT_BROAD_LINES
160-45
| 5137. | +/- | 10. |
| 5159. | +/- | 97. |
| 4.46 | +/- | 0. |
| 4.56 | +/- | 0. |
| 2.14 | +/- | 0. |
| 2.14 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -0.18 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.01 | +/- | 0. |
| 24.59 | +/- | 0. |
| 1.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.08 |
| -9999.00 |

PERSIST_LOW
160-45
| 5702. | +/- | 17. |
| 5651. | +/- | 146. |
| 4.22 | +/- | 0. |
| 4.14 | +/- | 0. |
| 1.00 | +/- | 0. |
| 1.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.01 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.08 | +/- | 0. |
| 3.56 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.32 |
| -9999.00 |
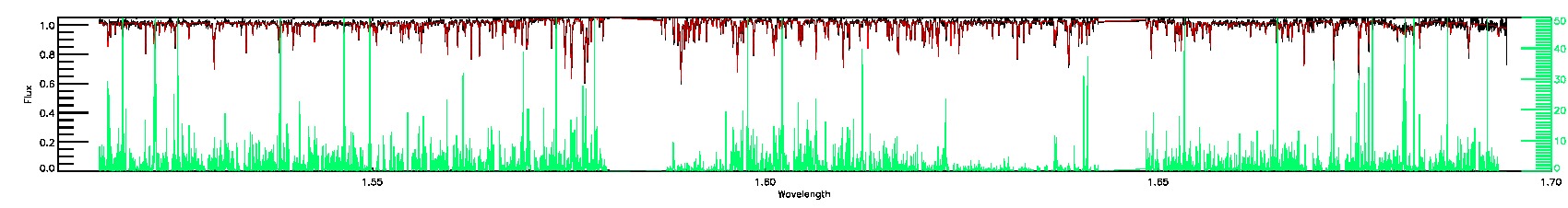
SUSPECT_BROAD_LINES
160-45
| 6578. | +/- | 18. |
| 6434. | +/- | 149. |
| 4.48 | +/- | 0. |
| 4.32 | +/- | 0. |
| 0.88 | +/- | 0. |
| 0.88 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.06 | +/- | 0. |
| 19.25 | +/- | 0. |
| 1.28 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.15 |
| -9999.00 |
160-45
| 5453. | +/- | 14. |
| 5445. | +/- | 118. |
| 3.87 | +/- | 0. |
| 3.93 | +/- | 0. |
| 0.82 | +/- | 0. |
| 0.82 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -0.35 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.06 | +/- | 0. |
| 3.62 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.37 |
| -9999.00 |

160-45
| 5064. | +/- | 15. |
| 5125. | +/- | 116. |
| 3.17 | +/- | 0. |
| 2.80 | +/- | 0. |
| 1.15 | +/- | 0. |
| 1.15 | +/- | -NaN |
| -0.88 | +/- | 0. |
| -0.88 | +/- | 0. |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| -0.33 | +/- | 0. |
| -0.33 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.14 | +/- | 0. |
| 4.94 | +/- | 0. |
| 0.69 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -1.98 |
| -9999.00 |
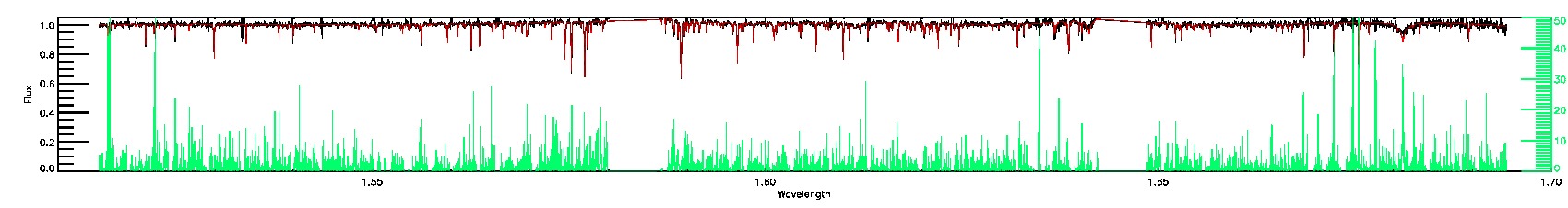
160-45
| 4325. | +/- | 4. |
| 4482. | +/- | 93. |
| 1.16 | +/- | 0. |
| 1.22 | +/- | 0. |
| 2.06 | +/- | 0. |
| 2.06 | +/- | -NaN |
| -1.60 | +/- | 0. |
| -1.60 | +/- | 0. |
| -0.32 | +/- | 0. |
| -0.32 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.19 | +/- | 0. |
| 7.56 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.32 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -2.14 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -1.15 |
| -9999.00 |

160-45
| 4379. | +/- | 6. |
| 4483. | +/- | 98. |
| 4.50 | +/- | 0. |
| 4.82 | +/- | 0. |
| 0.88 | +/- | 0. |
| 0.88 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -0.36 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.08 | +/- | 0. |
| 1.62 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.09 |
| -9999.00 |
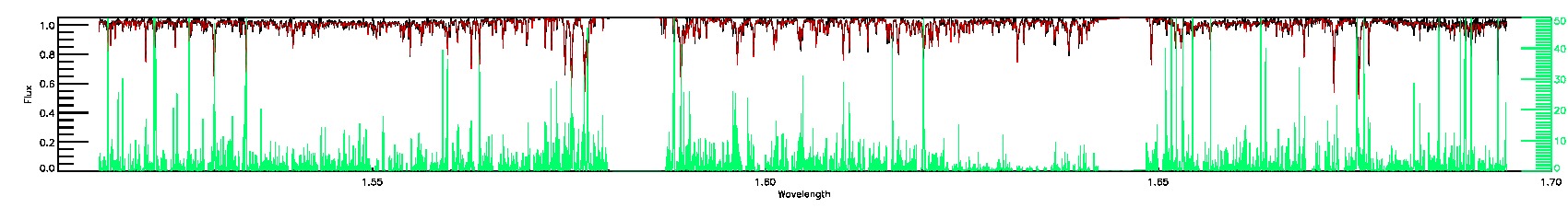
PERSIST_LOW
STAR_WARN,COLORTE_WARN
160-45
| 6253. | +/- | 22. |
| 6145. | +/- | 168. |
| 4.36 | +/- | 0. |
| 4.23 | +/- | 0. |
| 1.58 | +/- | 0. |
| 1.58 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.13 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 6.71 | +/- | 0. |
| 0.83 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.27 |
| -9999.00 |
PERSIST_LOW
STAR_WARN,SN_WARN
160-45
| 5558. | +/- | 16. |
| 5521. | +/- | 142. |
| 4.14 | +/- | 0. |
| 4.06 | +/- | 0. |
| 0.89 | +/- | 0. |
| 0.89 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.05 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.07 |
| -9999.00 |
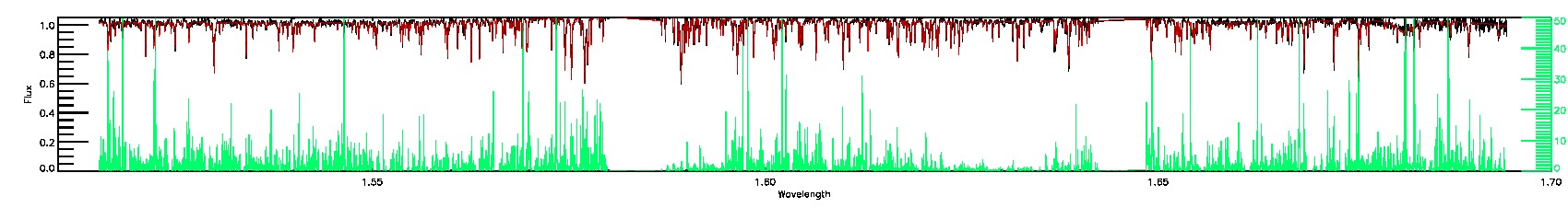
160-45
| 4070. | +/- | 3. |
| 4163. | +/- | 82. |
| 4.42 | +/- | 0. |
| 4.69 | +/- | 0. |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -0.14 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 3.12 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.04 |
| -9999.00 |

160-45
| 6385. | +/- | 15. |
| 6257. | +/- | 139. |
| 4.03 | +/- | 0. |
| 3.85 | +/- | 0. |
| 1.90 | +/- | 0. |
| 1.90 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.17 | +/- | 0. |
| -0.17 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 7.19 | +/- | 0. |
| 0.86 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.08 |
| -9999.00 |
160-45
| 5218. | +/- | 7. |
| 5213. | +/- | 106. |
| 4.47 | +/- | 0. |
| 4.38 | +/- | 0. |
| 0.46 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 1.74 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.30 |
| -9999.00 |

BRIGHT_NEIGHBOR
160-45
| 6042. | +/- | 22. |
| 5968. | +/- | 130. |
| 4.42 | +/- | 0. |
| 4.44 | +/- | 0. |
| 0.90 | +/- | 0. |
| 0.90 | +/- | -NaN |
| -0.38 | +/- | 0. |
| -0.38 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 4.51 | +/- | 0. |
| 0.65 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.22 |
| -9999.00 |
160-45
| 5260. | +/- | 19. |
| 5339. | +/- | 128. |
| 2.76 | +/- | 0. |
| 2.79 | +/- | 0. |
| 1.71 | +/- | 0. |
| 1.71 | +/- | -NaN |
| -1.83 | +/- | 0. |
| -1.83 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -0.26 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.26 | +/- | 0. |
| 8.63 | +/- | 0. |
| 0.94 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -2.11 |
| -9999.00 |
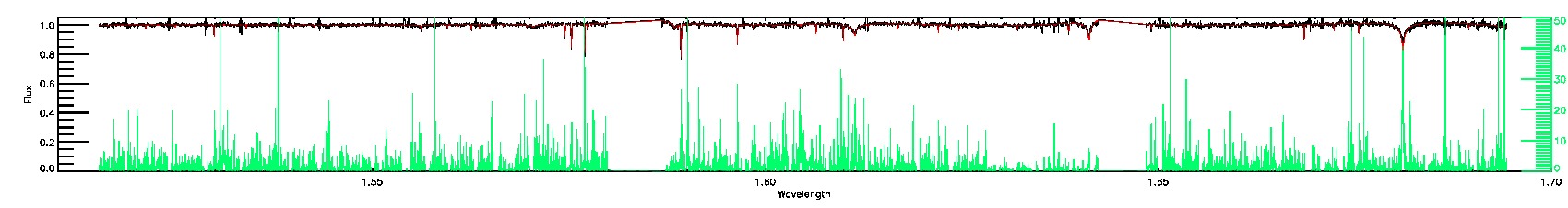
160-45
| 4360. | +/- | 2. |
| 4445. | +/- | 75. |
| 2.60 | +/- | 0. |
| 2.36 | +/- | 0. |
| 1.11 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.09 | +/- | 0. |
| 2.85 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.24 |
| -9999.00 |
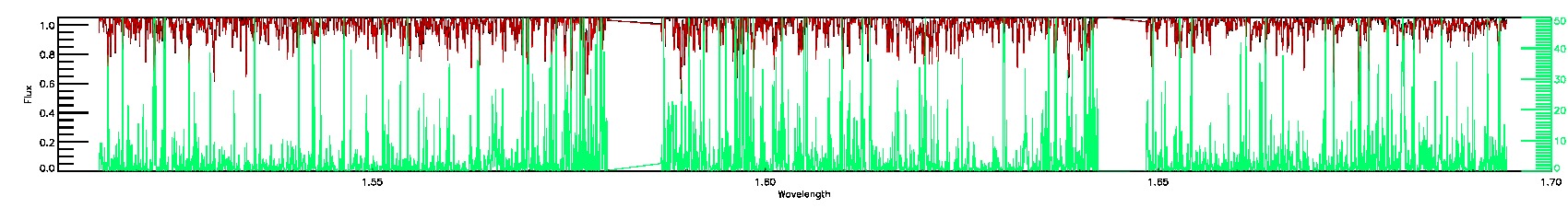
BRIGHT_NEIGHBOR,PERSIST_LOW,SUSPECT_BROAD_LINES
160-45
| 6624. | +/- | 18. |
| 6464. | +/- | 147. |
| 4.62 | +/- | 0. |
| 4.35 | +/- | 0. |
| 1.02 | +/- | 0. |
| 1.02 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.06 | +/- | 0. |
| -0.13 | +/- | 0. |
| -0.13 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -0.25 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 13.02 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.43 |
| -9999.00 |
PERSIST_LOW,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_WARN,COLORTE_WARN
160-45
| 8230. | +/- | 22. |
| 8116. | +/- | 302. |
| 4.53 | +/- | 0. |
| 4.95 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 26.01 | +/- | 0. |
| 1.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.97 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -1.76 |
| -9999.00 |
160-45
| 4265. | +/- | 2. |
| 4363. | +/- | 75. |
| 2.11 | +/- | 0. |
| 1.90 | +/- | 0. |
| 1.39 | +/- | 0. |
| 1.39 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -0.23 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.07 | +/- | 0. |
| 3.37 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.58 |
| -9999.00 |

160-45
| 6029. | +/- | 29. |
| 5953. | +/- | 155. |
| 4.29 | +/- | 0. |
| 4.28 | +/- | 0. |
| 0.78 | +/- | 0. |
| 0.78 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -0.30 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.06 | +/- | 0. |
| 4.44 | +/- | 0. |
| 0.65 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -1.76 |
| -9999.00 |
PERSIST_LOW
160-45
| 5780. | +/- | 26. |
| 5736. | +/- | 146. |
| 4.40 | +/- | 0. |
| 4.46 | +/- | 0. |
| 0.93 | +/- | 0. |
| 0.93 | +/- | -NaN |
| -0.37 | +/- | 0. |
| -0.37 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.18 | +/- | 0. |
| 2.05 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -2.30 |
| -9999.00 |

PERSIST_LOW
160-45
| 4165. | +/- | 4. |
| 4250. | +/- | 88. |
| 4.30 | +/- | 0. |
| 4.48 | +/- | 0. |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.08 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.07 | +/- | 0. |
| 5.00 | +/- | 0. |
| 0.70 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.16 |
| -9999.00 |

160-45
| 5451. | +/- | 12. |
| 5429. | +/- | 119. |
| 4.62 | +/- | 0. |
| 4.59 | +/- | 0. |
| 1.51 | +/- | 0. |
| 1.51 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.34 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |

SUSPECT_BROAD_LINES
160-45
| 6577. | +/- | 18. |
| 6426. | +/- | 146. |
| 4.43 | +/- | 0. |
| 4.20 | +/- | 0. |
| 0.90 | +/- | 0. |
| 0.90 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 33.03 | +/- | 0. |
| 1.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.05 |
| -9999.00 |
160-45
| 5966. | +/- | 13. |
| 5886. | +/- | 122. |
| 4.08 | +/- | 0. |
| 3.97 | +/- | 0. |
| 1.38 | +/- | 0. |
| 1.38 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 4.69 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.11 |
| -9999.00 |
PERSIST_LOW
160-45
| 3936. | +/- | 4. |
| 4028. | +/- | 88. |
| 4.38 | +/- | 0. |
| 4.67 | +/- | 0. |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -0.20 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.07 | +/- | 0. |
| 4.94 | +/- | 0. |
| 0.69 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.01 |
| -9999.00 |
160-45
| 4745. | +/- | 5. |
| 4823. | +/- | 92. |
| 2.55 | +/- | 0. |
| 2.30 | +/- | 0. |
| 1.58 | +/- | 0. |
| 1.58 | +/- | -NaN |
| -0.43 | +/- | 0. |
| -0.43 | +/- | 0. |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.23 | +/- | 0. |
| 3.80 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.85 |
| -9999.00 |

160-45
| 4561. | +/- | 5. |
| 4652. | +/- | 102. |
| 4.39 | +/- | 0. |
| 4.62 | +/- | 0. |
| 0.66 | +/- | 0. |
| 0.66 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.24 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.10 | +/- | 0. |
| 1.81 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.21 |
| -9999.00 |
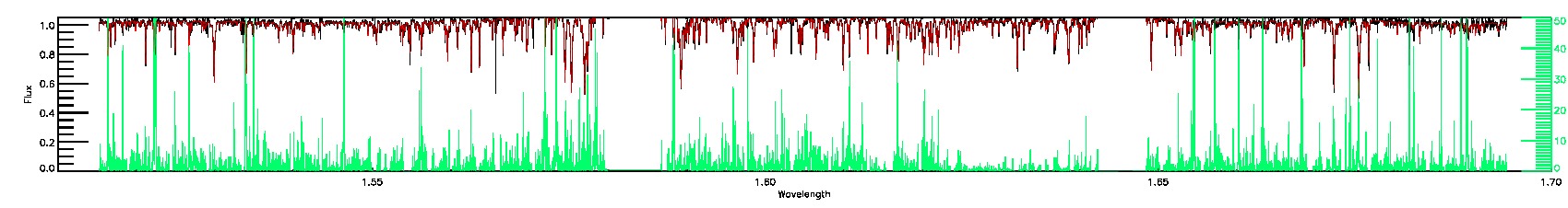
160-45
| 4936. | +/- | 6. |
| 4996. | +/- | 95. |
| 2.74 | +/- | 0. |
| 2.41 | +/- | 0. |
| 1.68 | +/- | 0. |
| 1.68 | +/- | -NaN |
| -0.52 | +/- | 0. |
| -0.52 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.08 | +/- | 0. |
| 4.01 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.47 |
| -9999.00 |

STAR_WARN,COLORTE_WARN,SN_WARN
160-45
| 3542. | +/- | 7. |
| 3641. | +/- | 85. |
| 4.40 | +/- | 0. |
| 4.83 | +/- | 0. |
| 0.60 | +/- | 0. |
| 0.60 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -0.25 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.13 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 5.66 | +/- | 0. |
| 0.75 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.14 |
| -9999.00 |
160-45
| 4824. | +/- | 5. |
| 4882. | +/- | 88. |
| 3.07 | +/- | 0. |
| 2.90 | +/- | 0. |
| 1.22 | +/- | 0. |
| 1.22 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.22 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.45 |
| -9999.00 |
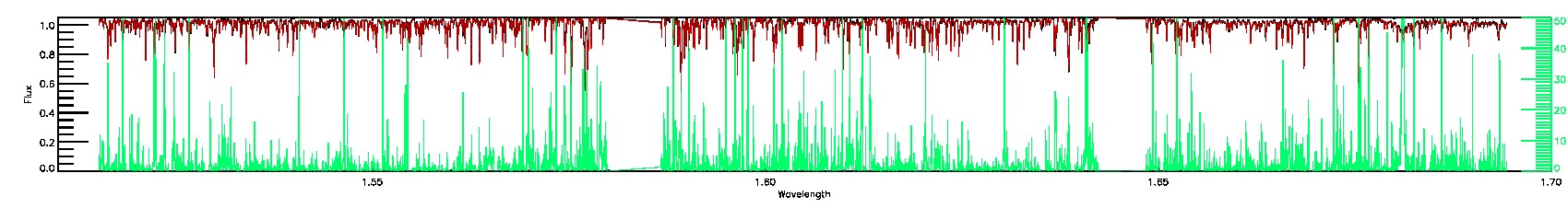
BRIGHT_NEIGHBOR
160-45
| 5804. | +/- | 22. |
| 5742. | +/- | 148. |
| 4.40 | +/- | 0. |
| 4.31 | +/- | 0. |
| 0.91 | +/- | 0. |
| 0.91 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.34 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.36 |
| -9999.00 |
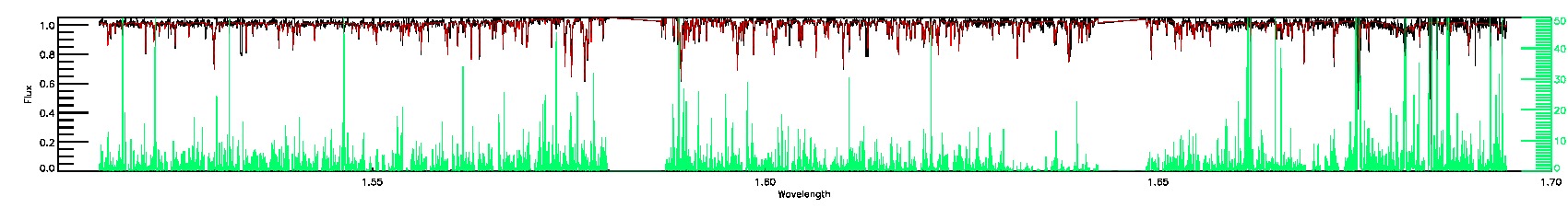
160-45
| 5833. | +/- | 20. |
| 5775. | +/- | 141. |
| 4.38 | +/- | 0. |
| 4.36 | +/- | 0. |
| 0.80 | +/- | 0. |
| 0.80 | +/- | -NaN |
| -0.21 | +/- | 0. |
| -0.21 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.15 | +/- | 0. |
| 1.91 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.13 |
| -9999.00 |

160-45
| 6464. | +/- | 18. |
| 6329. | +/- | 143. |
| 4.61 | +/- | 0. |
| 4.43 | +/- | 0. |
| 0.90 | +/- | 0. |
| 0.90 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -0.19 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 9.63 | +/- | 0. |
| 0.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.20 |
| -9999.00 |
160-45
| 5647. | +/- | 9. |
| 5595. | +/- | 108. |
| 4.45 | +/- | 0. |
| 4.30 | +/- | 0. |
| 0.37 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.19 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.10 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.15 |
| -9999.00 |
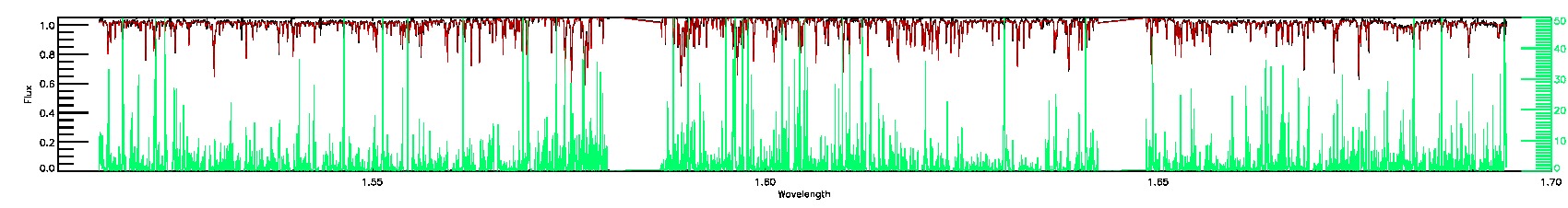
160-45
| 5095. | +/- | 9. |
| 5125. | +/- | 105. |
| 4.40 | +/- | 0. |
| 4.54 | +/- | 0. |
| 0.93 | +/- | 0. |
| 0.93 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -0.25 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.77 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.13 |
| -9999.00 |
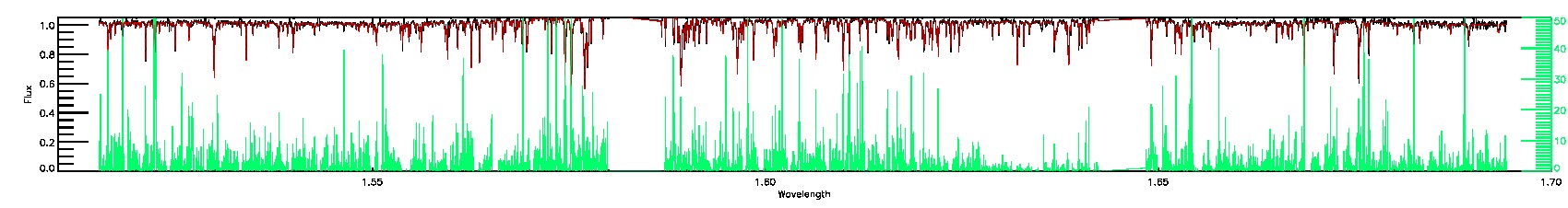
BRIGHT_NEIGHBOR
160-45
| 4556. | +/- | 4. |
| 4644. | +/- | 90. |
| 4.42 | +/- | 0. |
| 4.61 | +/- | 0. |
| 1.11 | +/- | 0. |
| 1.11 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 4.82 | +/- | 0. |
| 0.68 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.03 |
| -9999.00 |

160-45
| 5408. | +/- | 11. |
| 5400. | +/- | 117. |
| 4.40 | +/- | 0. |
| 4.47 | +/- | 0. |
| 0.47 | +/- | 0. |
| 0.47 | +/- | -NaN |
| -0.21 | +/- | 0. |
| -0.21 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.07 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.26 |
| -9999.00 |

SUSPECT_BROAD_LINES
160-45
| 5065. | +/- | 11. |
| 5082. | +/- | 107. |
| 3.68 | +/- | 0. |
| 3.56 | +/- | 0. |
| 2.68 | +/- | 0. |
| 2.68 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | 0. |
| -0.24 | +/- | 0. |
| -0.24 | +/- | -NaN |
| 0.37 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 74.83 | +/- | 0. |
| 1.87 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.27 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.29 |
| -9999.00 |

160-45
| 5945. | +/- | 13. |
| 5863. | +/- | 134. |
| 4.61 | +/- | 0. |
| 4.46 | +/- | 0. |
| 1.04 | +/- | 0. |
| 1.04 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.07 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.05 | +/- | 0. |
| 5.01 | +/- | 0. |
| 0.70 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.27 |
| -9999.00 |

PERSIST_LOW
160-45
| 5263. | +/- | 7. |
| 5259. | +/- | 100. |
| 4.65 | +/- | 0. |
| 4.61 | +/- | 0. |
| 1.13 | +/- | 0. |
| 1.13 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 2.75 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.17 |
| -9999.00 |
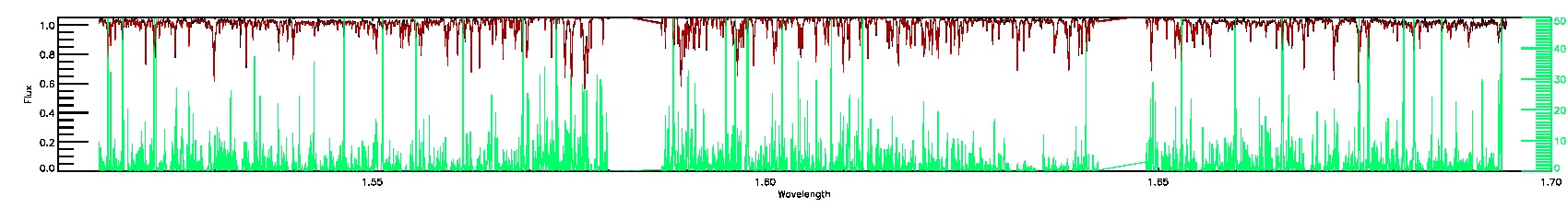
160-45
| 6107. | +/- | 17. |
| 6020. | +/- | 130. |
| 4.28 | +/- | 0. |
| 4.22 | +/- | 0. |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -0.25 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.15 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 6.23 | +/- | 0. |
| 0.79 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| 0.03 |
| -9999.00 |
BRIGHT_NEIGHBOR
160-45
| 4870. | +/- | 7. |
| 4914. | +/- | 101. |
| 4.63 | +/- | 0. |
| 4.70 | +/- | 0. |
| 0.90 | +/- | 0. |
| 0.90 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.03 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 1.55 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.12 |
| -9999.00 |
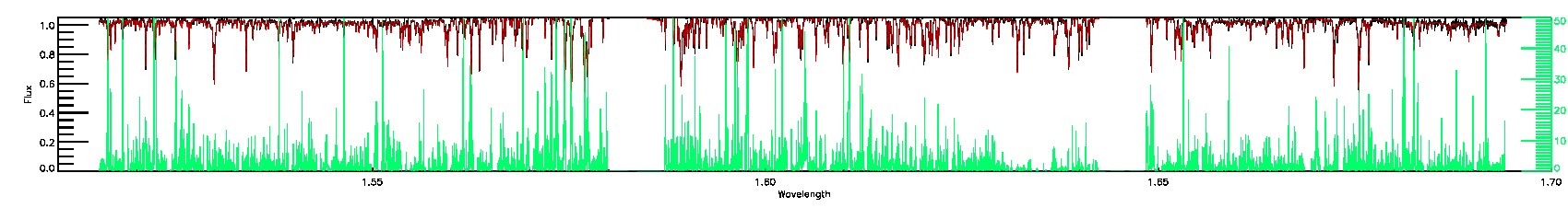
SUSPECT_BROAD_LINES
160-45
| 6517. | +/- | 18. |
| 6378. | +/- | 146. |
| 4.35 | +/- | 0. |
| 4.17 | +/- | 0. |
| 1.25 | +/- | 0. |
| 1.25 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 29.21 | +/- | 0. |
| 1.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.06 |
| -9999.00 |
160-45
| 3780. | +/- | 1. |
| 3914. | +/- | 82. |
| 0.49 | +/- | 0. |
| 0.49 | +/- | 0. |
| 2.39 | +/- | 0. |
| 2.39 | +/- | -NaN |
| -1.08 | +/- | 0. |
| -1.08 | +/- | 0. |
| -0.42 | +/- | 0. |
| -0.42 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 5.58 | +/- | 0. |
| 0.75 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.43 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -2.07 |
| -9999.00 |
160-45
| 6134. | +/- | 18. |
| 6037. | +/- | 133. |
| 4.43 | +/- | 0. |
| 4.31 | +/- | 0. |
| 1.30 | +/- | 0. |
| 1.30 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 5.83 | +/- | 0. |
| 0.77 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.65 |
| -9999.00 |
160-45
| 5170. | +/- | 9. |
| 5172. | +/- | 115. |
| 3.91 | +/- | 0. |
| 3.83 | +/- | 0. |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.04 | +/- | 0. |
| 2.64 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.26 |
| -9999.00 |

BRIGHT_NEIGHBOR,PERSIST_LOW
160-45
| 6529. | +/- | 23. |
| 6401. | +/- | 167. |
| 4.25 | +/- | 0. |
| 4.20 | +/- | 0. |
| 1.33 | +/- | 0. |
| 1.33 | +/- | -NaN |
| -0.46 | +/- | 0. |
| -0.46 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.14 | +/- | 0. |
| 13.48 | +/- | 0. |
| 1.13 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.12 |
| -9999.00 |
160-45
| 4315. | +/- | 3. |
| 4412. | +/- | 95. |
| 4.32 | +/- | 0. |
| 4.58 | +/- | 0. |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -0.20 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 4.09 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.10 |
| -9999.00 |

SUSPECT_BROAD_LINES
160-45
| 6393. | +/- | 16. |
| 6264. | +/- | 139. |
| 4.08 | +/- | 0. |
| 3.90 | +/- | 0. |
| 1.78 | +/- | 0. |
| 1.78 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 42.10 | +/- | 0. |
| 1.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.15 |
| -9999.00 |
160-45
| 5942. | +/- | 20. |
| 5873. | +/- | 140. |
| 4.22 | +/- | 0. |
| 4.18 | +/- | 0. |
| 1.49 | +/- | 0. |
| 1.49 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 4.71 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.11 |
| -9999.00 |
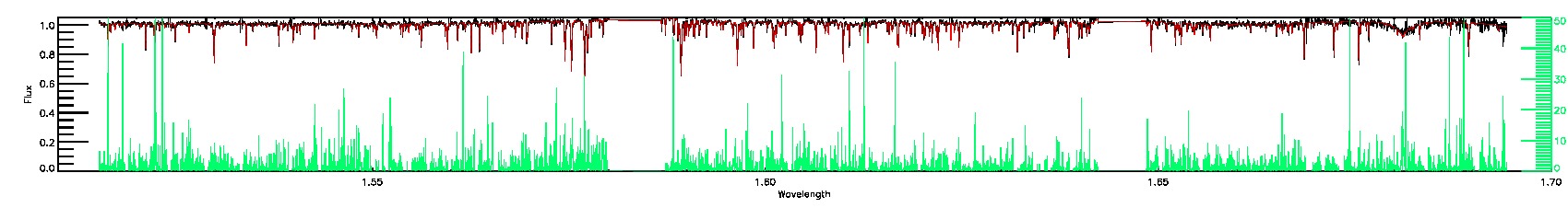
160-45
| 4155. | +/- | 2. |
| 4265. | +/- | 94. |
| 1.85 | +/- | 0. |
| 1.69 | +/- | 0. |
| 1.52 | +/- | 0. |
| 1.52 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -0.50 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.20 | +/- | 0. |
| 3.97 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.03 |
| -9999.00 |

160-45
| 4933. | +/- | 9. |
| 4970. | +/- | 107. |
| 4.55 | +/- | 0. |
| 4.60 | +/- | 0. |
| 0.88 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 1.59 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.08 |
| -9999.00 |

BRIGHT_NEIGHBOR
160-45
| 5658. | +/- | 11. |
| 5604. | +/- | 122. |
| 4.41 | +/- | 0. |
| 4.27 | +/- | 0. |
| 0.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 1.85 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.34 |
| -9999.00 |
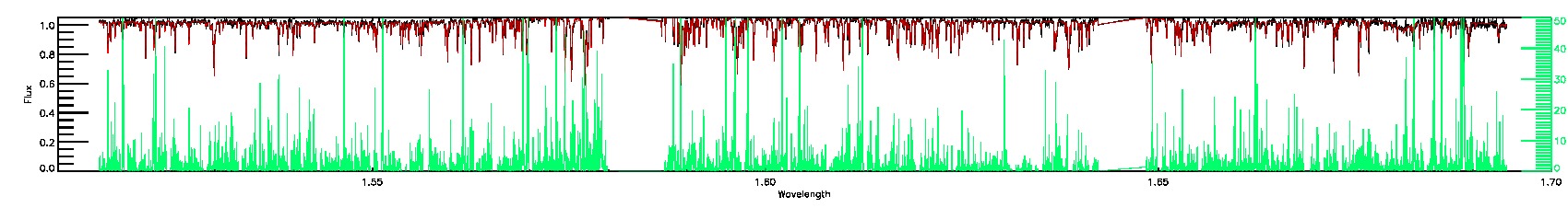
160-45
| 5788. | +/- | 19. |
| 5734. | +/- | 146. |
| 4.32 | +/- | 0. |
| 4.30 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.43 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.17 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.10 | +/- | 0. |
| 2.11 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 1.00 |
| -9999.00 |

160-45
| 4832. | +/- | 7. |
| 4869. | +/- | 97. |
| 4.54 | +/- | 0. |
| 4.51 | +/- | 0. |
| 0.74 | +/- | 0. |
| 0.74 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.29 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.12 | +/- | 0. |
| 2.37 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.45 |
| -9999.00 |

160-45
| 5141. | +/- | 10. |
| 5158. | +/- | 111. |
| 4.54 | +/- | 0. |
| 4.60 | +/- | 0. |
| 1.27 | +/- | 0. |
| 1.27 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.02 | +/- | 0. |
| 1.60 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.21 |
| -9999.00 |
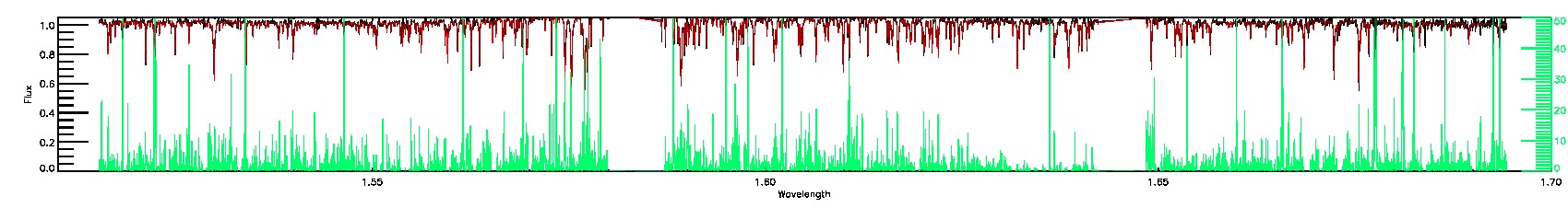
PERSIST_LOW
160-45
| 6404. | +/- | 19. |
| 6274. | +/- | 144. |
| 4.28 | +/- | 0. |
| 4.08 | +/- | 0. |
| 0.67 | +/- | 0. |
| 0.67 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.14 | +/- | 0. |
| -0.14 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 12.46 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.16 |
| -9999.00 |
PERSIST_LOW
160-45
| 6052. | +/- | 19. |
| 5955. | +/- | 144. |
| 4.35 | +/- | 0. |
| 4.16 | +/- | 0. |
| 1.18 | +/- | 0. |
| 1.18 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.11 | +/- | 0. |
| -0.13 | +/- | 0. |
| -0.13 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.29 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 4.43 | +/- | 0. |
| 0.65 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.01 |
| -9999.00 |
160-45
| 6145. | +/- | 20. |
| 6054. | +/- | 132. |
| 4.06 | +/- | 0. |
| 4.01 | +/- | 0. |
| 1.62 | +/- | 0. |
| 1.62 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -0.27 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 6.32 | +/- | 0. |
| 0.80 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.61 |
| -9999.00 |
PERSIST_LOW,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES
160-45
| 5703. | +/- | 23. |
| 5645. | +/- | 141. |
| 4.45 | +/- | 0. |
| 4.30 | +/- | 0. |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.00 | +/- | 0. |
| 17.02 | +/- | 0. |
| 1.23 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 0.51 |
| -9999.00 |

160-45
| 4584. | +/- | 3. |
| 4658. | +/- | 80. |
| 2.76 | +/- | 0. |
| 2.53 | +/- | 0. |
| 1.24 | +/- | 0. |
| 1.24 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.79 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.00 |
| -9999.00 |
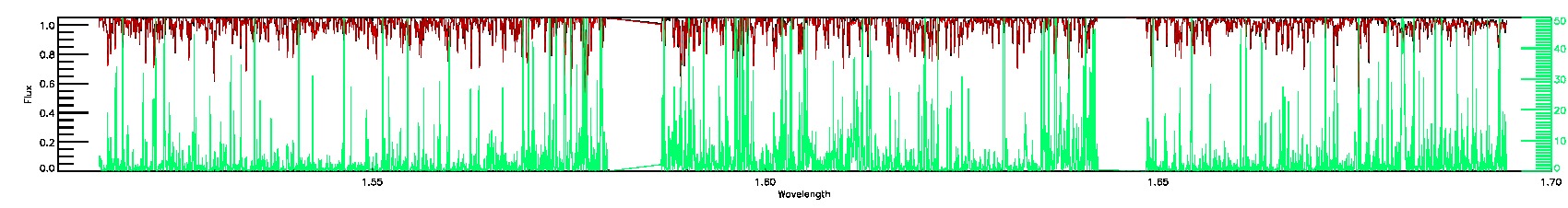
BRIGHT_NEIGHBOR
160-45
| 5344. | +/- | 9. |
| 5333. | +/- | 109. |
| 4.60 | +/- | 0. |
| 4.58 | +/- | 0. |
| 1.08 | +/- | 0. |
| 1.08 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.03 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 1.75 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.10 |
| -9999.00 |
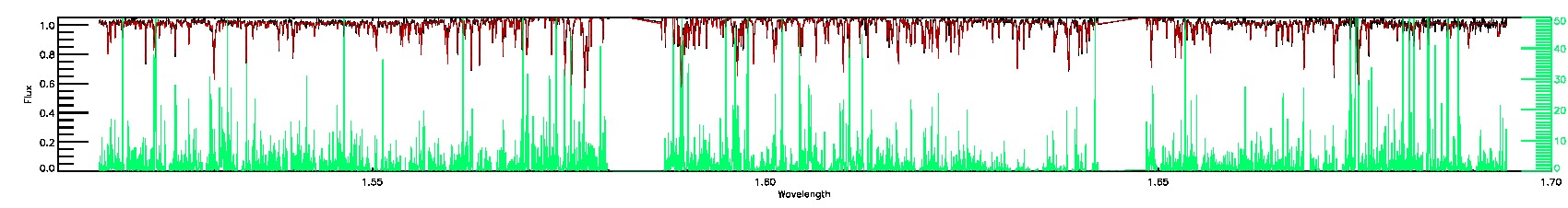
160-45
| 5682. | +/- | 26. |
| 5644. | +/- | 137. |
| 3.95 | +/- | 0. |
| 3.97 | +/- | 0. |
| 1.52 | +/- | 0. |
| 1.52 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -0.26 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 6.20 | +/- | 0. |
| 0.79 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.08 |
| -9999.00 |
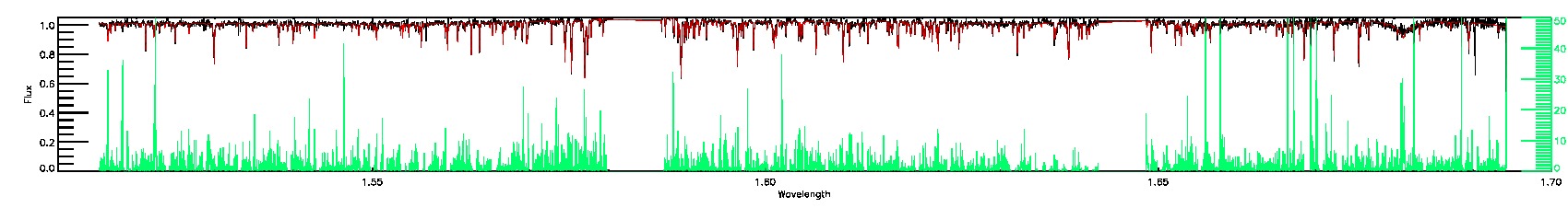
160-45
| 4494. | +/- | 3. |
| 4585. | +/- | 79. |
| 4.35 | +/- | 0. |
| 4.52 | +/- | 0. |
| 0.46 | +/- | 0. |
| 0.46 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 8.41 | +/- | 0. |
| 0.92 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.06 |
| -9999.00 |

BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_WARN,COLORTE_WARN
160-45
| 3503. | +/- | 3. |
| 3593. | +/- | 65. |
| 4.51 | +/- | 0. |
| 4.86 | +/- | 0. |
| 1.22 | +/- | 0. |
| 1.22 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 18.24 | +/- | 0. |
| 1.26 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.56 |
| -9999.00 |
BRIGHT_NEIGHBOR
160-45
| 5746. | +/- | 13. |
| 5689. | +/- | 115. |
| 4.68 | +/- | 0. |
| 4.58 | +/- | 0. |
| 1.00 | +/- | 0. |
| 1.00 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.03 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 6.45 | +/- | 0. |
| 0.81 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.01 |
| -9999.00 |

STAR_BAD
160-45
| 6772. | +/- | 15. |
| -10000. | +/- | -NaN |
| 3.99 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.75 | +/- | 0. |
| 0.75 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.90 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -0.31 |
| -9999.00 |
160-45
| 5774. | +/- | 12. |
| 5713. | +/- | 114. |
| 4.11 | +/- | 0. |
| 4.00 | +/- | 0. |
| 1.27 | +/- | 0. |
| 1.27 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.87 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.09 |
| -9999.00 |

160-45
| 5864. | +/- | 17. |
| 5805. | +/- | 131. |
| 4.45 | +/- | 0. |
| 4.45 | +/- | 0. |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -0.26 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.54 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 1.00 |
| -9999.00 |
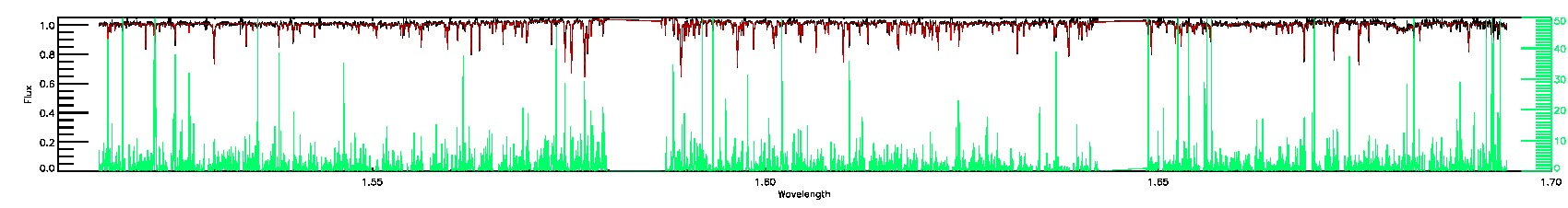
160-45
| 5584. | +/- | 9. |
| 5548. | +/- | 109. |
| 4.53 | +/- | 0. |
| 4.48 | +/- | 0. |
| 0.65 | +/- | 0. |
| 0.65 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.03 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 1.72 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.04 |
| -9999.00 |

BRIGHT_NEIGHBOR
160-45
| 3936. | +/- | 3. |
| 4027. | +/- | 83. |
| 4.43 | +/- | 0. |
| 4.72 | +/- | 0. |
| 0.56 | +/- | 0. |
| 0.56 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 4.41 | +/- | 0. |
| 0.64 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.04 |
| -9999.00 |
160-45
| 5539. | +/- | 13. |
| 5512. | +/- | 115. |
| 3.90 | +/- | 0. |
| 3.87 | +/- | 0. |
| 1.36 | +/- | 0. |
| 1.36 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.35 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.16 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
160-45
| 3941. | +/- | 15. |
| -10000. | +/- | -NaN |
| 4.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| -0.73 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.11 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.82 |
| -9999.00 |
BRIGHT_NEIGHBOR
160-45
| 4671. | +/- | 7. |
| 4748. | +/- | 102. |
| 4.47 | +/- | 0. |
| 4.66 | +/- | 0. |
| 0.75 | +/- | 0. |
| 0.75 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -0.19 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.35 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.03 |
| -9999.00 |

PERSIST_LOW
160-45
| 5274. | +/- | 10. |
| 5279. | +/- | 112. |
| 3.84 | +/- | 0. |
| 3.84 | +/- | 0. |
| 1.40 | +/- | 0. |
| 1.40 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.14 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 1.73 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.24 |
| -9999.00 |

160-45
| 5905. | +/- | 9. |
| 5825. | +/- | 118. |
| 4.31 | +/- | 0. |
| 4.14 | +/- | 0. |
| 0.93 | +/- | 0. |
| 0.93 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 5.48 | +/- | 0. |
| 0.74 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.23 |
| -9999.00 |
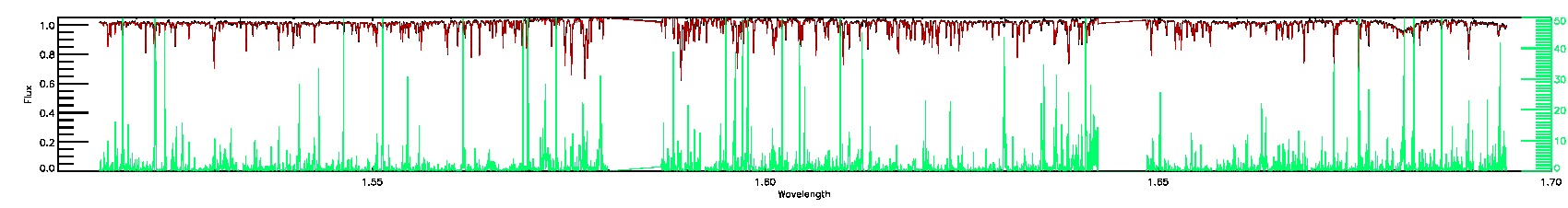
BRIGHT_NEIGHBOR
160-45
| 4689. | +/- | 6. |
| 4759. | +/- | 93. |
| 4.50 | +/- | 0. |
| 4.64 | +/- | 0. |
| 0.66 | +/- | 0. |
| 0.66 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.00 | +/- | 0. |
| 9.02 | +/- | 0. |
| 0.96 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.00 |
| -9999.00 |
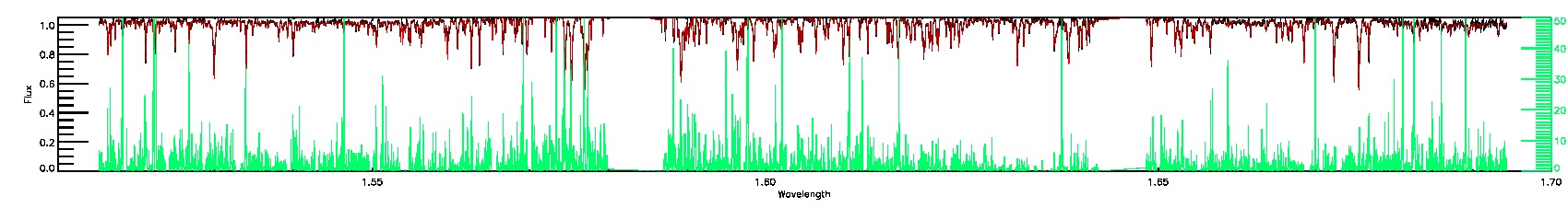
160-45
| 4470. | +/- | 2. |
| 4567. | +/- | 80. |
| 2.73 | +/- | 0. |
| 2.54 | +/- | 0. |
| 1.07 | +/- | 0. |
| 1.07 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -0.19 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.14 | +/- | 0. |
| 3.30 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.29 |
| -9999.00 |
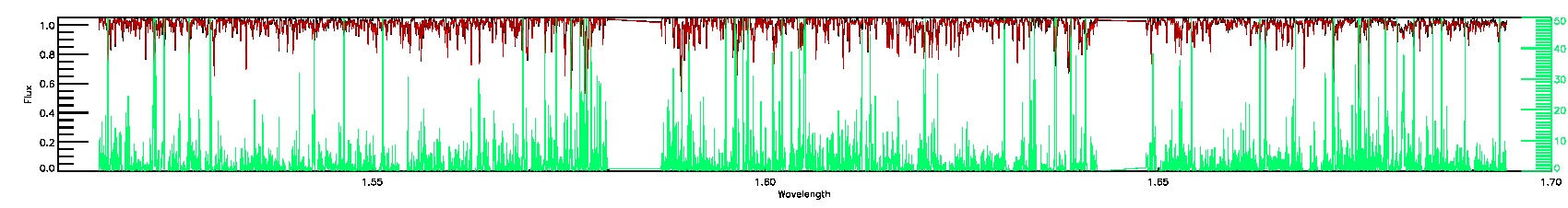
PERSIST_LOW
160-45
| 5535. | +/- | 14. |
| 5499. | +/- | 130. |
| 4.54 | +/- | 0. |
| 4.46 | +/- | 0. |
| 1.22 | +/- | 0. |
| 1.22 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 1.75 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.09 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD
160-45
| 6800. | +/- | 12. |
| -10000. | +/- | -NaN |
| 3.67 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.84 | +/- | 0. |
| 0.84 | +/- | -NaN |
| -0.52 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.92 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -0.17 |
| -9999.00 |
160-45
| 6372. | +/- | 18. |
| 6269. | +/- | 147. |
| 4.24 | +/- | 0. |
| 4.28 | +/- | 0. |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -0.60 | +/- | 0. |
| -0.59 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.13 | +/- | 0. |
| 8.54 | +/- | 0. |
| 0.93 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.21 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -0.67 |
| -9999.00 |
160-45
| 6241. | +/- | 43. |
| 6136. | +/- | 173. |
| 4.15 | +/- | 0. |
| 4.05 | +/- | 0. |
| 1.68 | +/- | 0. |
| 1.68 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -0.20 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.54 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.09 | +/- | 0. |
| 7.23 | +/- | 0. |
| 0.86 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.61 |
| -9999.00 |
160-45
| 5839. | +/- | 25. |
| 5767. | +/- | 144. |
| 4.20 | +/- | 0. |
| 4.05 | +/- | 0. |
| 1.21 | +/- | 0. |
| 1.21 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.43 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.77 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.23 |
| -9999.00 |
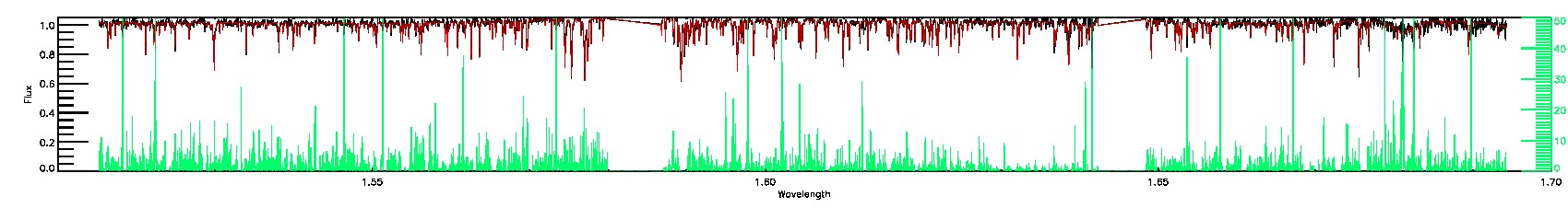
PERSIST_LOW
160-45
| 6429. | +/- | 16. |
| 6298. | +/- | 141. |
| 4.39 | +/- | 0. |
| 4.22 | +/- | 0. |
| 1.82 | +/- | 0. |
| 1.82 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.13 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 6.65 | +/- | 0. |
| 0.82 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.08 |
| -9999.00 |
160-45
| 5001. | +/- | 7. |
| 5042. | +/- | 94. |
| 3.28 | +/- | 0. |
| 3.13 | +/- | 0. |
| 1.31 | +/- | 0. |
| 1.31 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.23 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.39 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.30 |
| -9999.00 |

160-45
| 5603. | +/- | 13. |
| 5575. | +/- | 112. |
| 4.44 | +/- | 0. |
| 4.49 | +/- | 0. |
| 1.13 | +/- | 0. |
| 1.13 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -0.26 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.71 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.34 |
| -9999.00 |
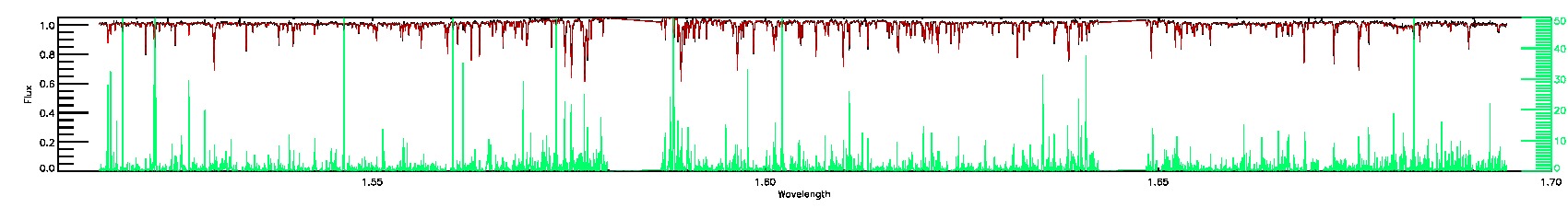
SUSPECT_BROAD_LINES
160-45
| 6716. | +/- | 17. |
| 6551. | +/- | 153. |
| 4.43 | +/- | 0. |
| 4.19 | +/- | 0. |
| 1.52 | +/- | 0. |
| 1.52 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -0.19 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.06 | +/- | 0. |
| 34.87 | +/- | 0. |
| 1.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.32 |
| -9999.00 |
PERSIST_LOW
160-45
| 4902. | +/- | 4. |
| 4942. | +/- | 88. |
| 2.84 | +/- | 0. |
| 2.48 | +/- | 0. |
| 1.64 | +/- | 0. |
| 1.64 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.87 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.31 |
| -9999.00 |
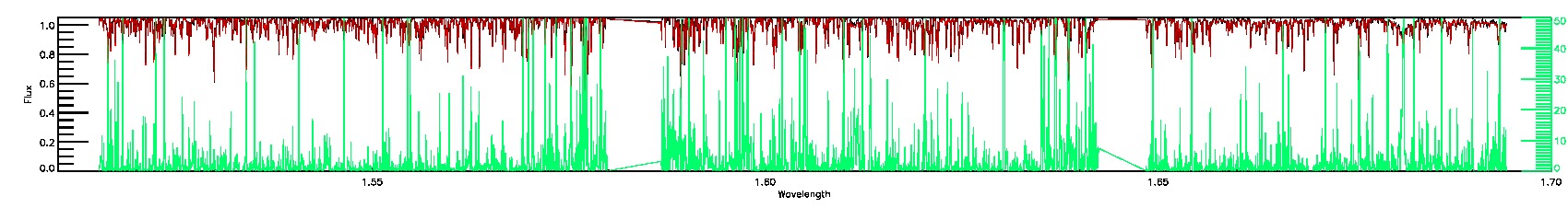
STAR_WARN,COLORTE_WARN
160-45
| 3697. | +/- | 4. |
| 3790. | +/- | 77. |
| 4.38 | +/- | 0. |
| 4.72 | +/- | 0. |
| 0.56 | +/- | 0. |
| 0.56 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -0.28 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 5.77 | +/- | 0. |
| 0.76 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.10 |
| -9999.00 |
160-45
| 5419. | +/- | 9. |
| 5406. | +/- | 105. |
| 4.50 | +/- | 0. |
| 4.51 | +/- | 0. |
| 1.26 | +/- | 0. |
| 1.26 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 1.64 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.13 |
| -9999.00 |
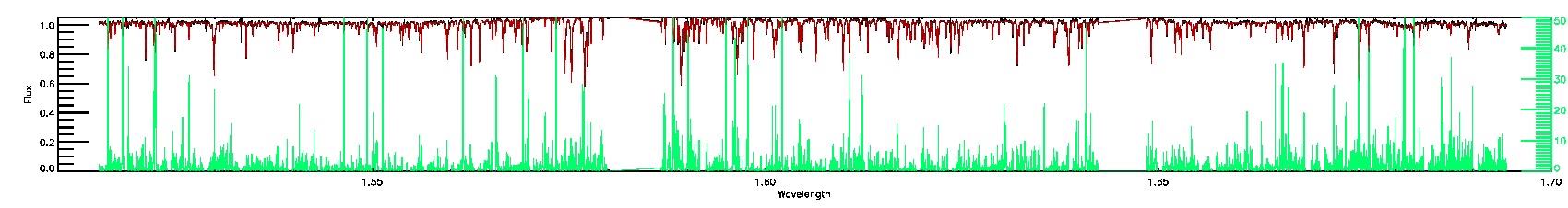
SUSPECT_BROAD_LINES
STAR_WARN,COLORTE_WARN
160-45
| 3630. | +/- | 3. |
| 3726. | +/- | 74. |
| 4.28 | +/- | 0. |
| 4.67 | +/- | 0. |
| 0.59 | +/- | 0. |
| 0.59 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -0.19 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -0.22 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.05 | +/- | 0. |
| 13.73 | +/- | 0. |
| 1.14 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.08 |
| -9999.00 |
BRIGHT_NEIGHBOR
160-45
| 5997. | +/- | 29. |
| 5918. | +/- | 152. |
| 4.53 | +/- | 0. |
| 4.45 | +/- | 0. |
| 1.13 | +/- | 0. |
| 1.13 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -0.14 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 3.95 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -2.23 |
| -9999.00 |
160-45
| 5287. | +/- | 23. |
| 5300. | +/- | 130. |
| 4.24 | +/- | 0. |
| 4.39 | +/- | 0. |
| 0.79 | +/- | 0. |
| 0.79 | +/- | -NaN |
| -0.37 | +/- | 0. |
| -0.36 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.24 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.04 | +/- | 0. |
| 6.08 | +/- | 0. |
| 0.78 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
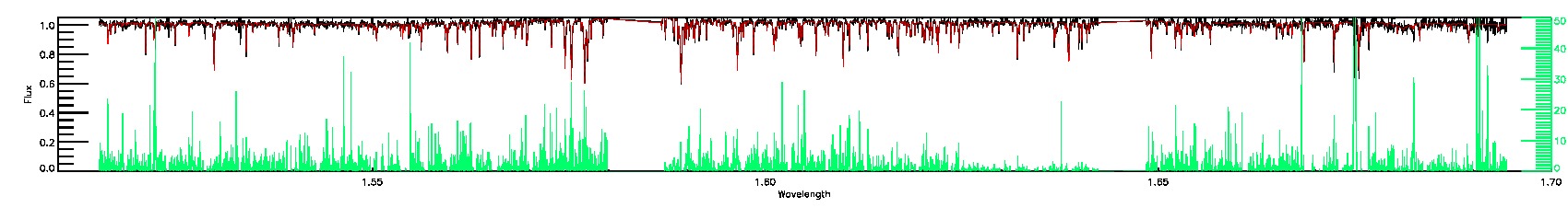
160-45
| 4460. | +/- | 3. |
| 4561. | +/- | 81. |
| 2.37 | +/- | 0. |
| 2.18 | +/- | 0. |
| 1.36 | +/- | 0. |
| 1.36 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -0.31 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.54 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.66 |
| -9999.00 |

PERSIST_LOW
160-45
| 5794. | +/- | 20. |
| 5740. | +/- | 144. |
| 4.37 | +/- | 0. |
| 4.35 | +/- | 0. |
| 0.59 | +/- | 0. |
| 0.59 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -0.19 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.30 | +/- | 0. |
| -0.30 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 1.56 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.53 |
| -9999.00 |

BRIGHT_NEIGHBOR
160-45
| 5664. | +/- | 17. |
| 5616. | +/- | 128. |
| 4.38 | +/- | 0. |
| 4.30 | +/- | 0. |
| 1.01 | +/- | 0. |
| 1.01 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.03 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.00 | +/- | 0. |
| 5.17 | +/- | 0. |
| 0.71 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.00 |
| -9999.00 |
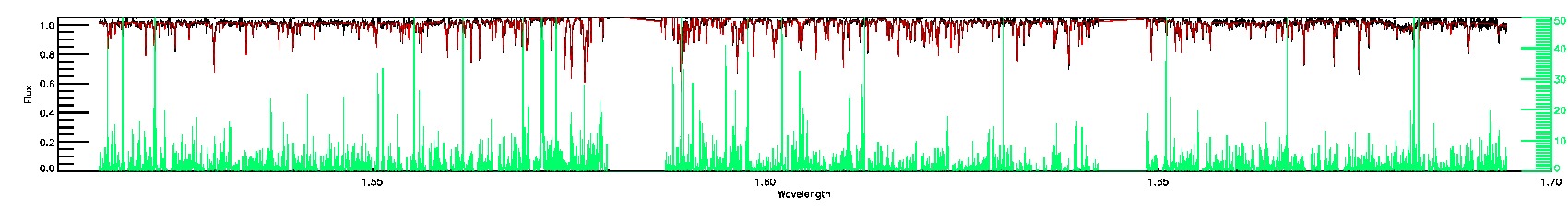
160-45
| 5671. | +/- | 26. |
| 5644. | +/- | 142. |
| 3.87 | +/- | 0. |
| 3.94 | +/- | 0. |
| 1.27 | +/- | 0. |
| 1.27 | +/- | -NaN |
| -0.48 | +/- | 0. |
| -0.48 | +/- | 0. |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.09 | +/- | 0. |
| 3.91 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.21 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -1.06 |
| -9999.00 |
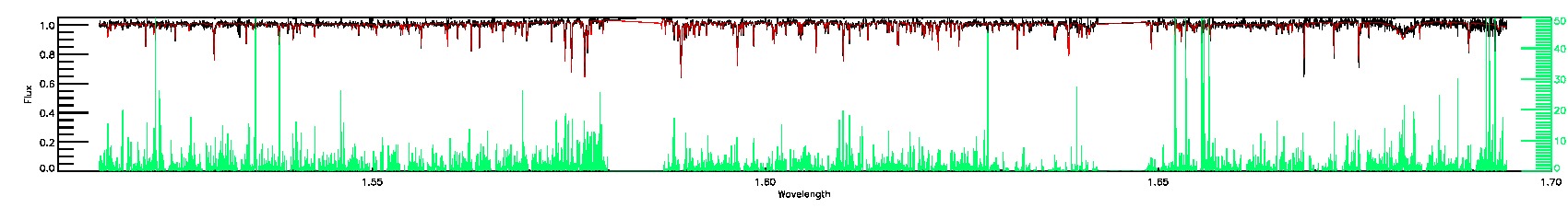
160-45
| 3780. | +/- | 1. |
| 3895. | +/- | 83. |
| 0.65 | +/- | 0. |
| 0.58 | +/- | 0. |
| 2.31 | +/- | 0. |
| 2.31 | +/- | -NaN |
| -0.62 | +/- | 0. |
| -0.62 | +/- | 0. |
| -0.35 | +/- | 0. |
| -0.35 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.03 | +/- | 0. |
| 4.24 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.36 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -1.73 |
| -9999.00 |
PERSIST_LOW
160-45
| 4675. | +/- | 10. |
| 4754. | +/- | 107. |
| 3.95 | +/- | 0. |
| 4.13 | +/- | 0. |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.24 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 7.86 | +/- | 0. |
| 0.90 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.06 |
| -9999.00 |
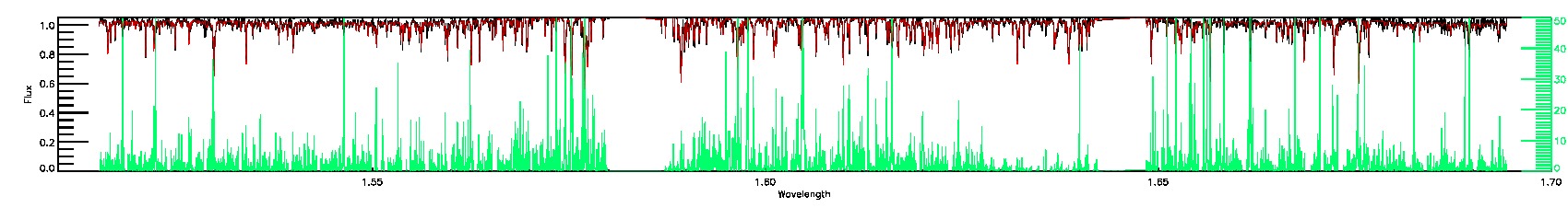
160-45
| 5381. | +/- | 10. |
| 5379. | +/- | 105. |
| 3.96 | +/- | 0. |
| 4.04 | +/- | 0. |
| 1.24 | +/- | 0. |
| 1.24 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -0.26 | +/- | 0. |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.13 | +/- | 0. |
| 3.45 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.55 |
| -9999.00 |

BRIGHT_NEIGHBOR
STAR_BAD
160-45
| 5666. | +/- | 25. |
| -10000. | +/- | -NaN |
| 4.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| -0.57 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.46 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.65 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.91 |
| -9999.00 |
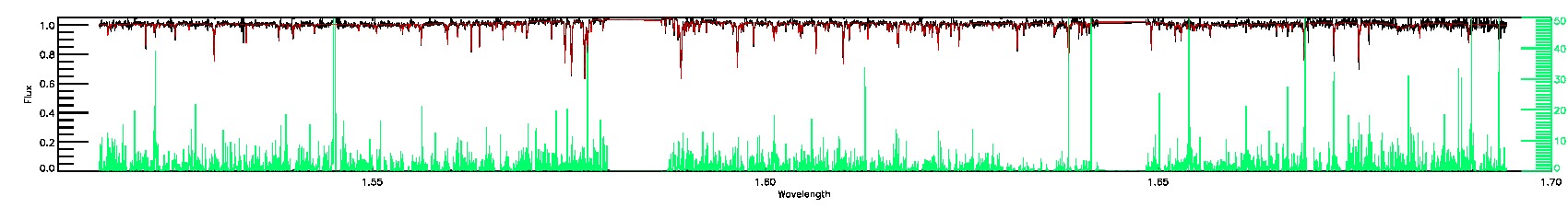
160-45
| 5476. | +/- | 15. |
| 5474. | +/- | 112. |
| 4.44 | +/- | 0. |
| 4.62 | +/- | 0. |
| 0.49 | +/- | 0. |
| 0.49 | +/- | -NaN |
| -0.53 | +/- | 0. |
| -0.53 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -0.26 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 1.59 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.84 |
| -9999.00 |

160-45
| 6084. | +/- | 19. |
| 5989. | +/- | 127. |
| 4.48 | +/- | 0. |
| 4.34 | +/- | 0. |
| 1.21 | +/- | 0. |
| 1.21 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 10.82 | +/- | 0. |
| 1.03 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.99 |
| -9999.00 |
SUSPECT_RV_COMBINATION
STAR_BAD
STAR_WARN,COLORTE_WARN
160-45
| 5694. | +/- | 13. |
| -10000. | +/- | -NaN |
| 3.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.77 | +/- | 0. |
| 0.77 | +/- | -NaN |
| -1.35 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.21 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.94 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -1.78 |
| -9999.00 |
BRIGHT_NEIGHBOR,PERSIST_LOW
160-45
| 5866. | +/- | 12. |
| 5797. | +/- | 129. |
| 4.44 | +/- | 0. |
| 4.34 | +/- | 0. |
| 1.31 | +/- | 0. |
| 1.31 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.27 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.08 |
| -9999.00 |

BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_WARN,COLORTE_WARN
160-45
| 3912. | +/- | 3. |
| 4003. | +/- | 81. |
| 4.41 | +/- | 0. |
| 4.69 | +/- | 0. |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -0.31 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.05 | +/- | 0. |
| 18.67 | +/- | 0. |
| 1.27 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.13 |
| -9999.00 |
PERSIST_LOW
160-45
| 5494. | +/- | 13. |
| 5471. | +/- | 122. |
| 3.95 | +/- | 0. |
| 3.94 | +/- | 0. |
| 1.35 | +/- | 0. |
| 1.35 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.13 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.70 |
| -9999.00 |

160-45
| 5462. | +/- | 17. |
| 5444. | +/- | 126. |
| 4.61 | +/- | 0. |
| 4.62 | +/- | 0. |
| 1.48 | +/- | 0. |
| 1.48 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 3.15 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.19 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
160-45
| 4833. | +/- | 15. |
| 4923. | +/- | 118. |
| 3.94 | +/- | 0. |
| 4.35 | +/- | 0. |
| 0.47 | +/- | 0. |
| 0.47 | +/- | -NaN |
| -0.94 | +/- | 0. |
| -0.94 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.23 | +/- | 0. |
| 14.45 | +/- | 0. |
| 1.16 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.17 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -1.31 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
160-45
| 7301. | +/- | 16. |
| 7104. | +/- | 186. |
| 4.54 | +/- | 0. |
| 4.21 | +/- | 0. |
| 4.14 | +/- | 0. |
| 4.14 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.37 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 8.78 | +/- | 0. |
| 0.94 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.32 |
| -9999.00 |
STAR_WARN,SN_WARN
160-45
| 4010. | +/- | 7. |
| 4111. | +/- | 95. |
| 4.45 | +/- | 0. |
| 4.81 | +/- | 0. |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| -0.30 | +/- | 0. |
| -0.30 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -0.16 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.85 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.79 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
160-45
| 6526. | +/- | 17. |
| 6385. | +/- | 146. |
| 4.34 | +/- | 0. |
| 4.17 | +/- | 0. |
| 1.74 | +/- | 0. |
| 1.74 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.06 | +/- | 0. |
| 8.72 | +/- | 0. |
| 0.94 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.03 |
| -9999.00 |
160-45
| 4182. | +/- | 4. |
| 4296. | +/- | 96. |
| 4.28 | +/- | 0. |
| 4.73 | +/- | 0. |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -0.61 | +/- | 0. |
| -0.60 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.09 | +/- | 0. |
| 2.53 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.38 |
| -9999.00 |

BRIGHT_NEIGHBOR
160-45
| 4942. | +/- | 14. |
| 4992. | +/- | 119. |
| 3.26 | +/- | 0. |
| 3.13 | +/- | 0. |
| 0.69 | +/- | 0. |
| 0.69 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -0.31 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.55 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.43 |
| -9999.00 |

160-45
| 5648. | +/- | 16. |
| 5632. | +/- | 129. |
| 4.04 | +/- | 0. |
| 4.25 | +/- | 0. |
| 0.46 | +/- | 0. |
| 0.46 | +/- | -NaN |
| -0.67 | +/- | 0. |
| -0.67 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| -0.33 | +/- | 0. |
| -0.33 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.24 | +/- | 0. |
| 4.39 | +/- | 0. |
| 0.64 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.80 |
| -9999.00 |

160-45
| 4811. | +/- | 8. |
| 4855. | +/- | 106. |
| 4.61 | +/- | 0. |
| 4.62 | +/- | 0. |
| 1.39 | +/- | 0. |
| 1.39 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 1.72 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.38 |
| -9999.00 |

160-45
| 7525. | +/- | 16. |
| 7340. | +/- | 202. |
| 4.63 | +/- | 0. |
| 4.37 | +/- | 0. |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| -0.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.43 | +/- | 0. |
| -0.43 | +/- | -NaN |
| -0.41 | +/- | 0. |
| -0.41 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 15.79 | +/- | 0. |
| 1.20 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.43 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.42 |
| -9999.00 |
160-45
| 5406. | +/- | 17. |
| 5399. | +/- | 129. |
| 4.41 | +/- | 0. |
| 4.48 | +/- | 0. |
| 0.43 | +/- | 0. |
| 0.43 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -0.22 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.15 | +/- | 0. |
| 1.79 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.21 |
| -9999.00 |
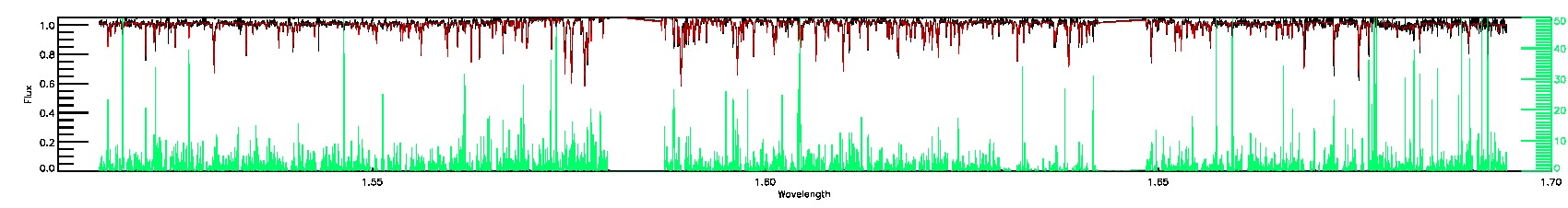
160-45
| 4266. | +/- | 5. |
| 4357. | +/- | 93. |
| 4.41 | +/- | 0. |
| 4.62 | +/- | 0. |
| 0.50 | +/- | 0. |
| 0.50 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.02 | +/- | 0. |
| 4.54 | +/- | 0. |
| 0.66 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.00 |
| -9999.00 |
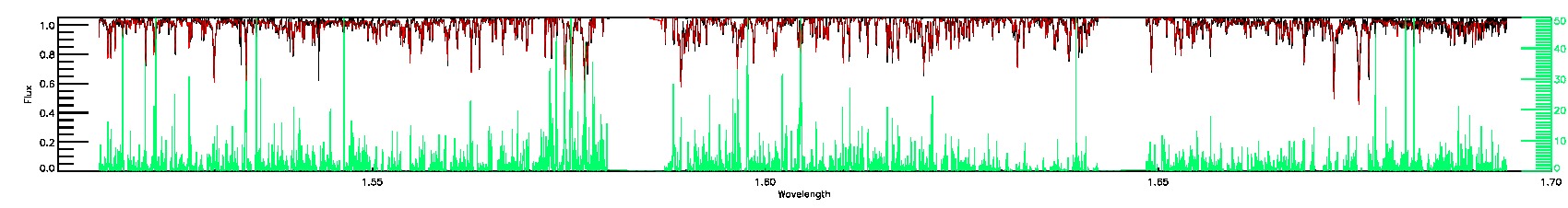
160-45
| 5964. | +/- | 28. |
| 5889. | +/- | 151. |
| 4.40 | +/- | 0. |
| 4.33 | +/- | 0. |
| 1.02 | +/- | 0. |
| 1.02 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 9.12 | +/- | 0. |
| 0.96 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.38 |
| -9999.00 |
160-45
| 5894. | +/- | 19. |
| 5839. | +/- | 129. |
| 4.07 | +/- | 0. |
| 4.13 | +/- | 0. |
| 0.79 | +/- | 0. |
| 0.79 | +/- | -NaN |
| -0.43 | +/- | 0. |
| -0.42 | +/- | 0. |
| -0.13 | +/- | 0. |
| -0.13 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.09 | +/- | 0. |
| 4.78 | +/- | 0. |
| 0.68 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.67 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_WARN,COLORTE_WARN
160-45
| 3633. | +/- | 3. |
| 3728. | +/- | 68. |
| 4.40 | +/- | 0. |
| 4.77 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.42 | +/- | 0. |
| -0.42 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 13.00 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.20 |
| -9999.00 |
160-45
| 5401. | +/- | 34. |
| 5419. | +/- | 140. |
| 4.36 | +/- | 0. |
| 4.67 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| -0.82 | +/- | 0. |
| -0.81 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.23 | +/- | 0. |
| 2.67 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.31 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.63 |
| -9999.00 |
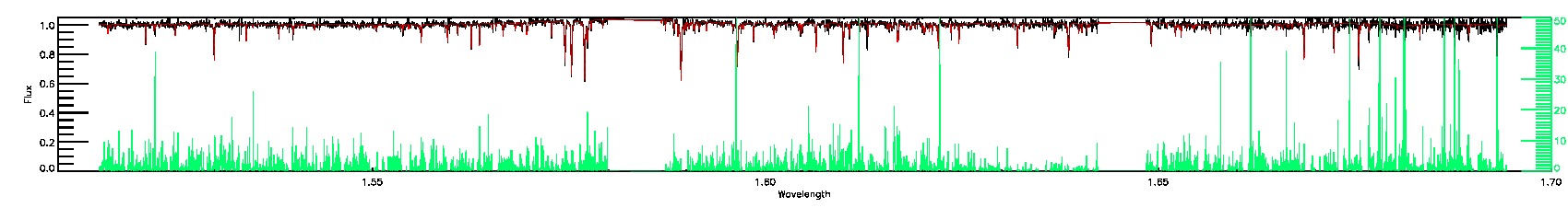
160-45
| 5617. | +/- | 16. |
| 5572. | +/- | 133. |
| 4.07 | +/- | 0. |
| 3.97 | +/- | 0. |
| 1.22 | +/- | 0. |
| 1.22 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.80 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.27 |
| -9999.00 |

160-45
| 4860. | +/- | 7. |
| 4901. | +/- | 102. |
| 4.37 | +/- | 0. |
| 4.40 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.14 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.09 | +/- | 0. |
| 3.85 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.19 |
| -9999.00 |
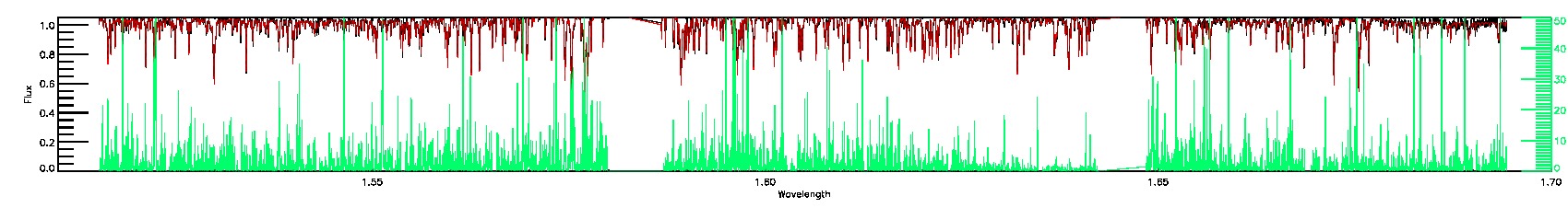
SUSPECT_BROAD_LINES
STAR_WARN,ROTATION_WARN
160-45
| 4128. | +/- | 2. |
| 4249. | +/- | 77. |
| 2.07 | +/- | 0. |
| 1.94 | +/- | 0. |
| 1.23 | +/- | 0. |
| 1.23 | +/- | -NaN |
| -0.75 | +/- | 0. |
| -0.75 | +/- | 0. |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.13 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.21 | +/- | 0. |
| 4.58 | +/- | 0. |
| 0.66 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.24 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.85 |
| -9999.00 |

160-45
| 5594. | +/- | 10. |
| 5554. | +/- | 109. |
| 4.45 | +/- | 0. |
| 4.39 | +/- | 0. |
| 1.05 | +/- | 0. |
| 1.05 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.02 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 3.25 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.05 |
| -9999.00 |

160-45
| 6263. | +/- | 19. |
| 6153. | +/- | 135. |
| 4.43 | +/- | 0. |
| 4.31 | +/- | 0. |
| 1.25 | +/- | 0. |
| 1.25 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 4.19 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.07 |
| -9999.00 |
160-45
| 5522. | +/- | 13. |
| 5490. | +/- | 125. |
| 4.53 | +/- | 0. |
| 4.46 | +/- | 0. |
| 1.14 | +/- | 0. |
| 1.14 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.12 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 1.98 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.00 |
| -9999.00 |
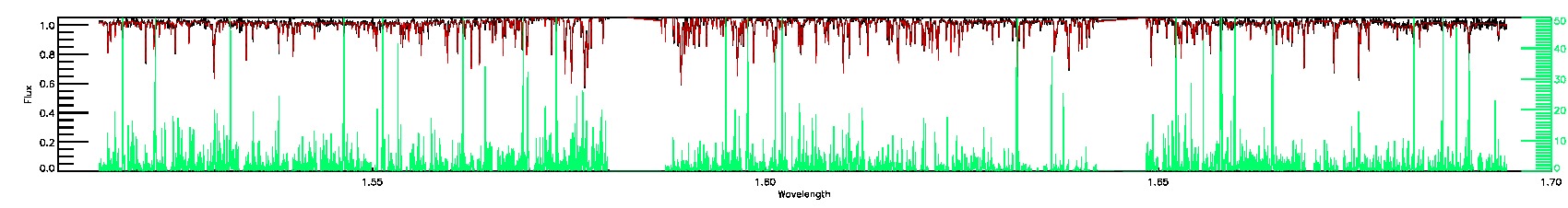
STAR_WARN,COLORTE_WARN
160-45
| 3464. | +/- | 5. |
| 3570. | +/- | 77. |
| 4.34 | +/- | 0. |
| 4.85 | +/- | 0. |
| 1.21 | +/- | 0. |
| 1.21 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -0.40 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.05 | +/- | 0. |
| 6.68 | +/- | 0. |
| 0.82 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.56 |
| -9999.00 |
160-45
| 6134. | +/- | 26. |
| 6052. | +/- | 149. |
| 3.94 | +/- | 0. |
| 3.95 | +/- | 0. |
| 1.37 | +/- | 0. |
| 1.37 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -0.44 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.09 | +/- | 0. |
| 6.03 | +/- | 0. |
| 0.78 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.37 |
| -9999.00 |
160-45
| 4942. | +/- | 6. |
| 4990. | +/- | 92. |
| 2.87 | +/- | 0. |
| 2.51 | +/- | 0. |
| 1.60 | +/- | 0. |
| 1.60 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -0.26 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.07 | +/- | 0. |
| 3.45 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.49 |
| -9999.00 |

160-45
| 5474. | +/- | 23. |
| 5451. | +/- | 137. |
| 4.46 | +/- | 0. |
| 4.44 | +/- | 0. |
| 0.80 | +/- | 0. |
| 0.80 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.04 | +/- | 0. |
| 2.27 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -1.93 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| 0.27 |
| -9999.00 |
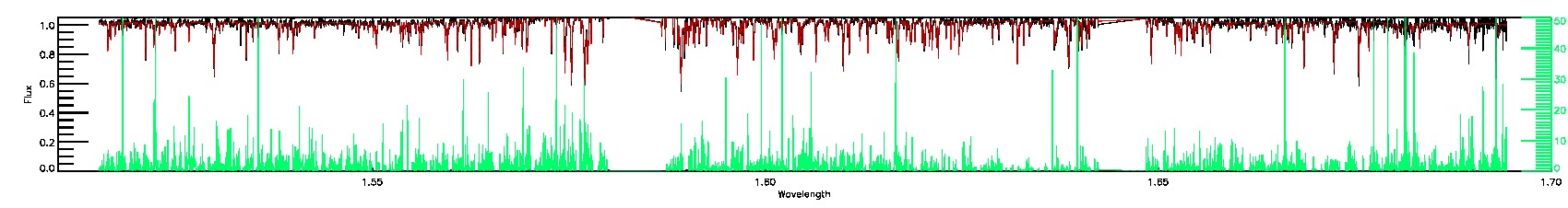
160-45
| 4293. | +/- | 2. |
| 4393. | +/- | 76. |
| 2.16 | +/- | 0. |
| 1.96 | +/- | 0. |
| 1.32 | +/- | 0. |
| 1.32 | +/- | -NaN |
| -0.27 | +/- | 0. |
| -0.27 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.08 | +/- | 0. |
| 3.46 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.68 |
| -9999.00 |
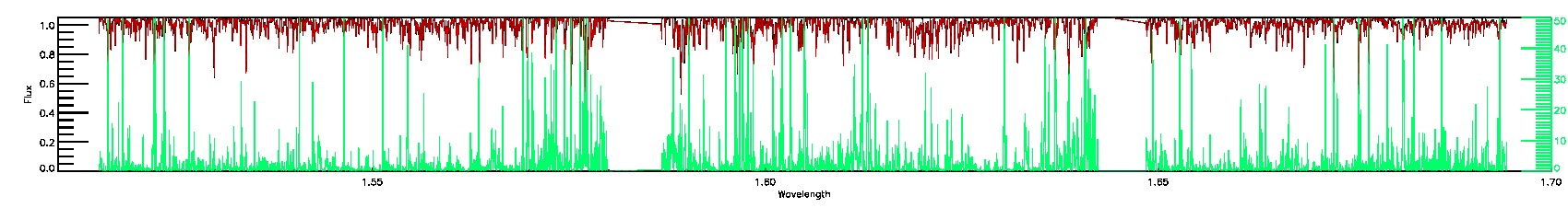
STAR_WARN,COLORTE_WARN
160-45
| 6565. | +/- | 19. |
| 6427. | +/- | 150. |
| 4.73 | +/- | 0. |
| 4.62 | +/- | 0. |
| 0.41 | +/- | 0. |
| 0.41 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -0.31 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 6.35 | +/- | 0. |
| 0.80 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
160-45
| 6944. | +/- | 12. |
| 6753. | +/- | 164. |
| 4.34 | +/- | 0. |
| 4.07 | +/- | 0. |
| 2.75 | +/- | 0. |
| 2.75 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.10 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.08 | +/- | 0. |
| 9.26 | +/- | 0. |
| 0.97 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.84 |
| -9999.00 |
160-45
| 5397. | +/- | 11. |
| 5390. | +/- | 110. |
| 3.85 | +/- | 0. |
| 3.86 | +/- | 0. |
| 1.08 | +/- | 0. |
| 1.08 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -0.22 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.35 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.40 |
| -9999.00 |

160-45
| 4840. | +/- | 6. |
| 4873. | +/- | 93. |
| 4.52 | +/- | 0. |
| 4.46 | +/- | 0. |
| 0.95 | +/- | 0. |
| 0.95 | +/- | -NaN |
| 0.37 | +/- | 0. |
| 0.37 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.10 | +/- | 0. |
| 1.56 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.43 |
| -9999.00 |

160-45
| 5173. | +/- | 9. |
| 5197. | +/- | 100. |
| 3.56 | +/- | 0. |
| 3.48 | +/- | 0. |
| 1.45 | +/- | 0. |
| 1.45 | +/- | -NaN |
| -0.33 | +/- | 0. |
| -0.33 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.58 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.33 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
160-45
| 4677. | +/- | 5. |
| 4758. | +/- | 89. |
| 4.15 | +/- | 0. |
| 4.38 | +/- | 0. |
| 0.81 | +/- | 0. |
| 0.81 | +/- | -NaN |
| -0.30 | +/- | 0. |
| -0.29 | +/- | 0. |
| -0.36 | +/- | 0. |
| -0.36 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.29 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 5.24 | +/- | 0. |
| 0.72 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.34 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.28 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
160-45
| 4542. | +/- | 3. |
| 4652. | +/- | 86. |
| 4.10 | +/- | 0. |
| 4.49 | +/- | 0. |
| 0.97 | +/- | 0. |
| 0.97 | +/- | -NaN |
| -0.62 | +/- | 0. |
| -0.62 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.15 | +/- | 0. |
| 5.32 | +/- | 0. |
| 0.73 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.87 |
| -9999.00 |
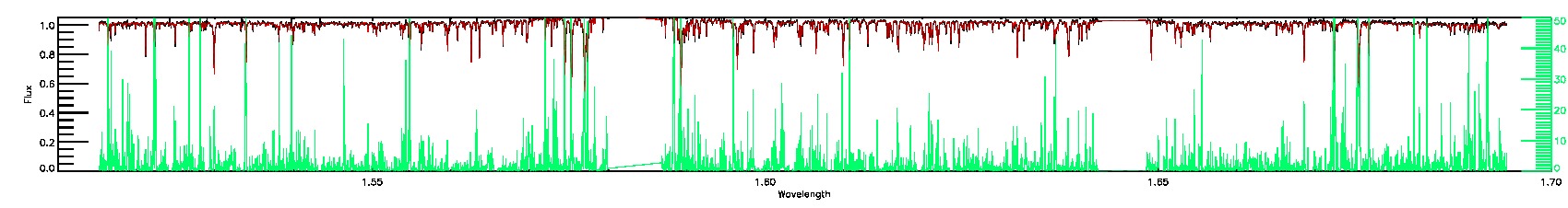
STAR_WARN,COLORTE_WARN
160-45
| 3520. | +/- | 3. |
| 3615. | +/- | 74. |
| 4.48 | +/- | 0. |
| 4.87 | +/- | 0. |
| 1.09 | +/- | 0. |
| 1.09 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.42 | +/- | 0. |
| -0.42 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 5.14 | +/- | 0. |
| 0.71 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.03 |
| -9999.00 |
BRIGHT_NEIGHBOR
160-45
| 4928. | +/- | 7. |
| 4983. | +/- | 93. |
| 3.62 | +/- | 0. |
| 3.59 | +/- | 0. |
| 1.03 | +/- | 0. |
| 1.03 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -0.36 | +/- | 0. |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.20 | +/- | 0. |
| 3.65 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -1.17 |
| -9999.00 |

160-45
| 5953. | +/- | 16. |
| 5884. | +/- | 129. |
| 4.02 | +/- | 0. |
| 4.00 | +/- | 0. |
| 1.23 | +/- | 0. |
| 1.23 | +/- | -NaN |
| -0.27 | +/- | 0. |
| -0.27 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 4.66 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -2.14 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.13 |
| -9999.00 |
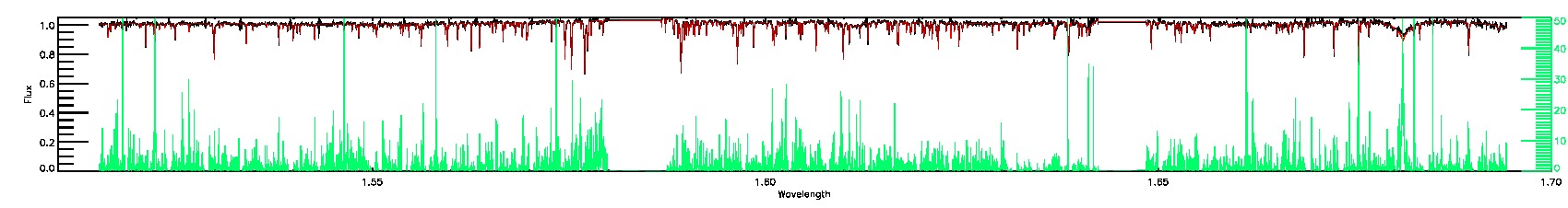
STAR_WARN,COLORTE_WARN
160-45
| 3973. | +/- | 2. |
| 4092. | +/- | 89. |
| 4.16 | +/- | 0. |
| 4.70 | +/- | 0. |
| 0.67 | +/- | 0. |
| 0.67 | +/- | -NaN |
| -0.72 | +/- | 0. |
| -0.72 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -0.31 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.09 | +/- | 0. |
| 6.34 | +/- | 0. |
| 0.80 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.53 |
| -9999.00 |
160-45
| 5703. | +/- | 14. |
| 5657. | +/- | 114. |
| 4.32 | +/- | 0. |
| 4.29 | +/- | 0. |
| 0.87 | +/- | 0. |
| 0.87 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 14.69 | +/- | 0. |
| 1.17 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
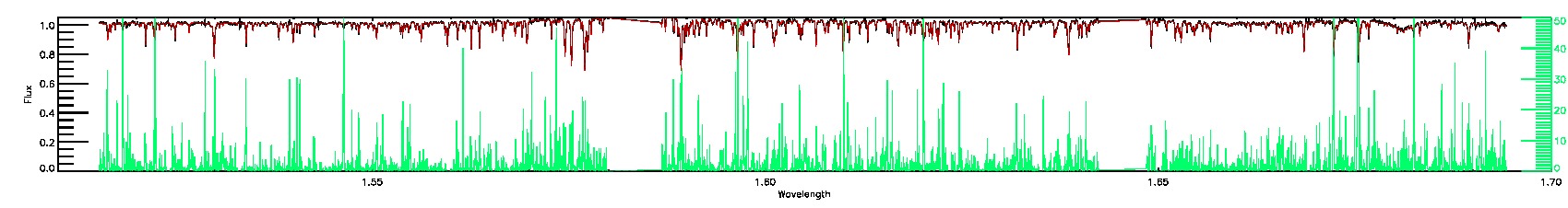
160-45
| 5508. | +/- | 9. |
| 5477. | +/- | 105. |
| 4.56 | +/- | 0. |
| 4.49 | +/- | 0. |
| 1.17 | +/- | 0. |
| 1.17 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.52 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.15 |
| -9999.00 |

STAR_WARN,SN_WARN
160-45
| 3907. | +/- | 7. |
| 4000. | +/- | 93. |
| 4.23 | +/- | 0. |
| 4.54 | +/- | 0. |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.24 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.05 | +/- | 0. |
| 4.99 | +/- | 0. |
| 0.70 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.11 |
| -9999.00 |
160-45
| 5770. | +/- | 12. |
| 5712. | +/- | 122. |
| 4.08 | +/- | 0. |
| 3.98 | +/- | 0. |
| 1.32 | +/- | 0. |
| 1.32 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.97 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.19 |
| -9999.00 |

160-45
| 5100. | +/- | 8. |
| 5122. | +/- | 101. |
| 2.97 | +/- | 0. |
| 2.60 | +/- | 0. |
| 1.52 | +/- | 0. |
| 1.52 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.40 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.01 | +/- | 0. |
| 3.09 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.33 |
| -9999.00 |
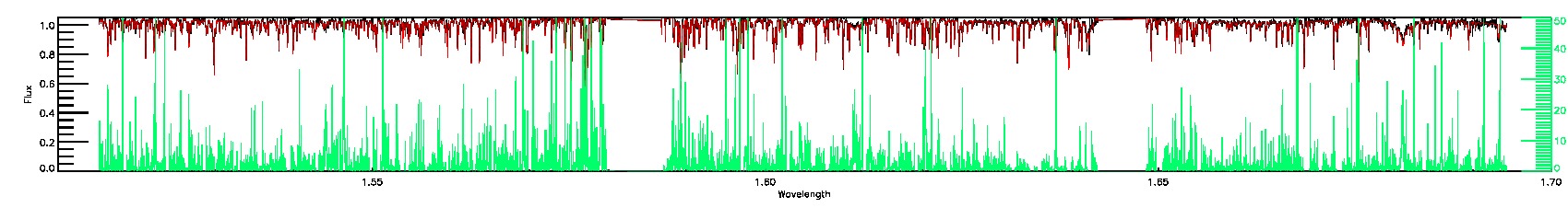
160-45
| 5346. | +/- | 12. |
| 5334. | +/- | 128. |
| 4.47 | +/- | 0. |
| 4.43 | +/- | 0. |
| 1.17 | +/- | 0. |
| 1.17 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 1.57 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.22 |
| -9999.00 |

