PERSIST_LOW,SUSPECT_BROAD_LINES
ANDR3
| 7845. | +/- | 12. |
| 7656. | +/- | 220. |
| 4.61 | +/- | 0. |
| 4.32 | +/- | 0. |
| 4.09 | +/- | 0. |
| 4.09 | +/- | -NaN |
| -0.32 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 19.37 | +/- | 0. |
| 1.29 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.34 |
| -9999.00 |
SUSPECT_RV_COMBINATION
STAR_BAD
STAR_WARN,COLORTE_WARN
ANDR3
| 8143. | +/- | 24. |
| -10000. | +/- | -NaN |
| 4.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -0.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.72 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.26 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.23 |
| -9999.00 |
SUSPECT_RV_COMBINATION
STAR_BAD
STAR_WARN,COLORTE_WARN
ANDR3
| 8229. | +/- | 25. |
| -10000. | +/- | -NaN |
| 4.31 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -0.61 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.98 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.84 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.55 |
| -9999.00 |
SUSPECT_BROAD_LINES
ANDR3
| 7009. | +/- | 11. |
| 6805. | +/- | 164. |
| 4.15 | +/- | 0. |
| 3.79 | +/- | 0. |
| 4.02 | +/- | 0. |
| 4.02 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 28.56 | +/- | 0. |
| 1.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.26 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.21 |
| -9999.00 |
SUSPECT_RV_COMBINATION
STAR_BAD
ANDR3
| 6620. | +/- | 14. |
| -10000. | +/- | -NaN |
| 3.73 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.14 | +/- | 0. |
| 1.14 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.31 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 94.93 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.28 |
| -9999.00 |
PERSIST_LOW,SUSPECT_BROAD_LINES
STAR_BAD
ANDR3
| 7341. | +/- | 10. |
| -10000. | +/- | -NaN |
| 4.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.43 | +/- | 0. |
| 4.43 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 16.74 | +/- | 0. |
| 1.22 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.12 |
| -9999.00 |
SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES
STAR_WARN,COLORTE_WARN
ANDR3
| 8817. | +/- | 60. |
| 8682. | +/- | 340. |
| 3.90 | +/- | 0. |
| 4.12 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -1.57 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 85.86 | +/- | 0. |
| 1.93 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.57 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.57 |
| -9999.00 |
SUSPECT_RV_COMBINATION
STAR_BAD
ANDR3
| 6685. | +/- | 16. |
| -10000. | +/- | -NaN |
| 3.92 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.78 | +/- | 0. |
| 0.78 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.90 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -0.49 |
| -9999.00 |
SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES
ANDR3
| 7098. | +/- | 13. |
| 6914. | +/- | 178. |
| 4.27 | +/- | 0. |
| 4.10 | +/- | 0. |
| 2.35 | +/- | 0. |
| 2.35 | +/- | -NaN |
| -0.41 | +/- | 0. |
| -0.41 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.10 | +/- | 0. |
| 32.75 | +/- | 0. |
| 1.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| 0.31 |
| -9999.00 |
SUSPECT_RV_COMBINATION
STAR_BAD
STAR_WARN,COLORTE_WARN
ANDR3
| 6269. | +/- | 17. |
| -10000. | +/- | -NaN |
| 3.67 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.37 | +/- | 0. |
| 1.37 | +/- | -NaN |
| -0.55 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.24 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.48 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.31 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.84 |
| -9999.00 |
SUSPECT_RV_COMBINATION
STAR_WARN,COLORTE_WARN
ANDR3
| 7767. | +/- | 14. |
| 7580. | +/- | 216. |
| 4.46 | +/- | 0. |
| 4.18 | +/- | 0. |
| 1.32 | +/- | 0. |
| 1.32 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 80.48 | +/- | 0. |
| 1.91 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -1.22 |
| -9999.00 |
SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,COLORTE_WARN
ANDR3
| 8912. | +/- | 61. |
| -10000. | +/- | -NaN |
| 3.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -1.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.21 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.60 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.89 |
| -9999.00 |
SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES
ANDR3
| 6861. | +/- | 14. |
| 6679. | +/- | 160. |
| 3.96 | +/- | 0. |
| 3.70 | +/- | 0. |
| 3.04 | +/- | 0. |
| 3.04 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.16 | +/- | 0. |
| 73.25 | +/- | 0. |
| 1.86 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.21 |
| -9999.00 |
SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_WARN,COLORTE_WARN
ANDR3
| 8370. | +/- | 25. |
| 8267. | +/- | 325. |
| 4.45 | +/- | 0. |
| 4.98 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.32 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.34 | +/- | 0. |
| 1.01 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.46 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -2.30 |
| -9999.00 |
SUSPECT_RV_COMBINATION
STAR_BAD
ANDR3
| 8791. | +/- | 43. |
| -10000. | +/- | -NaN |
| 4.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.72 | +/- | 0. |
| 0.76 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.44 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -2.44 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,ROTATION_BAD
STAR_WARN,ROTATION_WARN
ANDR3
| 3809. | +/- | 3. |
| -10000. | +/- | -NaN |
| 1.71 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.52 | +/- | 0. |
| 0.52 | +/- | -NaN |
| -0.98 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.24 | +/- | 0. |
| 0.72 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.43 |
| -9999.00 |
LOW_SNR,PERSIST_MED,PERSIST_LOW,PERSIST_JUMP_NEG,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
ANDR3
| 4496. | +/- | 636. |
| -10000. | +/- | -NaN |
| 4.49 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 1.60 | +/- | 4. |
| 1.60 | +/- | -NaN |
| -2.46 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.28 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| -0.17 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.48 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 12.50 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.27 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.47 |
| -9999.00 |

LOW_SNR,PERSIST_LOW,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 5682. | +/- | 1227. |
| -10000. | +/- | -NaN |
| 5.36 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 0.86 | +/- | 57. |
| 0.86 | +/- | -NaN |
| -2.46 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| -0.18 | +/- | 18. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 87.42 | +/- | 3. |
| 1.94 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -2.45 |
| -9999.00 |
LOW_SNR,PERSIST_MED,PERSIST_LOW,PERSIST_JUMP_NEG,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 4492. | +/- | 116. |
| -10000. | +/- | -NaN |
| 0.63 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.67 | +/- | 1. |
| 0.67 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.58 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.24 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| -0.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.76 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.60 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.50 |
| -9999.00 |

LOW_SNR,PERSIST_LOW,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 6932. | +/- | 285. |
| -10000. | +/- | -NaN |
| 4.79 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.79 | +/- | 2. |
| 0.79 | +/- | -NaN |
| -2.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| -0.34 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 2.35 | +/- | 2. |
| 0.37 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
LOW_SNR,PERSIST_LOW,PERSIST_JUMP_NEG,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 6278. | +/- | 315. |
| -10000. | +/- | -NaN |
| 4.39 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.27 | +/- | 1. |
| 1.27 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.30 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.50 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.28 |
| -9999.00 |
LOW_SNR,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 5762. | +/- | 1647. |
| -10000. | +/- | -NaN |
| 5.42 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.95 | +/- | 10. |
| 0.95 | +/- | -NaN |
| -2.21 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 6. |
| -9999.99 | +/- | -NaN |
| -0.12 | +/- | 10. |
| -9999.99 | +/- | -NaN |
| 0.28 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 91.94 | +/- | 2. |
| 1.96 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| -1.69 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -2.48 |
| -9999.00 |

LOW_SNR,PERSIST_MED,PERSIST_LOW,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 4771. | +/- | 139. |
| -10000. | +/- | -NaN |
| 4.82 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.61 | +/- | 1. |
| 0.61 | +/- | -NaN |
| -2.32 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 1.00 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 44.33 | +/- | 0. |
| 1.65 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -2.45 |
| -9999.00 |

BRIGHT_NEIGHBOR,LOW_SNR,PERSIST_LOW,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 4731. | +/- | 243. |
| -10000. | +/- | -NaN |
| 3.84 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.87 | +/- | 5. |
| 0.87 | +/- | -NaN |
| -2.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.64 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 11.32 | +/- | 0. |
| 1.05 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.52 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.14 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.47 |
| -9999.00 |

LOW_SNR,PERSIST_LOW,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 3800. | +/- | 107. |
| -10000. | +/- | -NaN |
| 5.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.67 | +/- | 1. |
| 0.67 | +/- | -NaN |
| -1.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 64.92 | +/- | 0. |
| 1.81 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| 0.99 |
| -9999.00 |
LOW_SNR,PERSIST_LOW,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 3637. | +/- | 67. |
| -10000. | +/- | -NaN |
| 2.85 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.93 | +/- | 1. |
| 0.93 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.15 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.73 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.23 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.73 |
| -9999.00 |
VERY_BRIGHT_NEIGHBOR,LOW_SNR,PERSIST_MED,PERSIST_LOW,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
ANDR3
| 3906. | +/- | 81. |
| -10000. | +/- | -NaN |
| 1.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.72 | +/- | 1. |
| 0.72 | +/- | -NaN |
| -2.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.26 | +/- | 0. |
| 1.09 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -2.42 |
| -9999.00 |
LOW_SNR,PERSIST_LOW,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 4426. | +/- | 180. |
| -10000. | +/- | -NaN |
| 5.09 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.97 | +/- | 2. |
| 0.97 | +/- | -NaN |
| -2.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.32 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 9.18 | +/- | 0. |
| 0.96 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.46 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.44 |
| -9999.00 |

LOW_SNR,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 5941. | +/- | 139. |
| -10000. | +/- | -NaN |
| 3.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.82 | +/- | 6. |
| 0.82 | +/- | -NaN |
| -2.31 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.26 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.10 | +/- | 8. |
| -9999.99 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.65 | +/- | 1. |
| 0.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.26 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| 1.00 |
| -9999.00 |
PERSIST_LOW,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 6443. | +/- | 427. |
| -10000. | +/- | -NaN |
| 4.99 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 2.83 | +/- | 3. |
| 2.83 | +/- | -NaN |
| -1.86 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 10. |
| -9999.99 | +/- | -NaN |
| -0.74 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.04 | +/- | 2. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -2.30 |
| -9999.00 |
LOW_SNR,PERSIST_LOW,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 9515. | +/- | 947. |
| -10000. | +/- | -NaN |
| 4.31 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -1.24 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.67 | +/- | 2. |
| 0.82 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.15 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -1.38 |
| -9999.00 |
PERSIST_LOW,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 5551. | +/- | 110. |
| -10000. | +/- | -NaN |
| 3.93 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.83 | +/- | 8. |
| 0.83 | +/- | -NaN |
| -2.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.88 | +/- | 1. |
| 0.27 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
LOW_SNR,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
ANDR3
| 3000. | +/- | 0. |
| -10000. | +/- | -NaN |
| 1.30 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.19 | +/- | 0. |
| 3.19 | +/- | -NaN |
| -1.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.39 | +/- | 0. |
| 0.73 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 1.60 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -1.04 |
| -9999.00 |
PERSIST_LOW,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
ANDR3
| 3719. | +/- | 20. |
| -10000. | +/- | -NaN |
| 1.45 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.59 | +/- | 0. |
| 1.59 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.76 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.27 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.50 |
| -9999.00 |
BRIGHT_NEIGHBOR,LOW_SNR,PERSIST_LOW,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
ANDR3
| 3783. | +/- | 35. |
| -10000. | +/- | -NaN |
| -0.41 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.95 | +/- | 0. |
| 3.95 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.76 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -2.50 |
| -9999.00 |
LOW_SNR,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 4282. | +/- | 103. |
| -10000. | +/- | -NaN |
| 2.21 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.44 | +/- | 2. |
| 1.44 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.69 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| -0.41 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.76 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.69 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.49 |
| -9999.00 |

PERSIST_LOW,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD
STAR_WARN,SN_WARN
ANDR3
| 5719. | +/- | 197. |
| -10000. | +/- | -NaN |
| 4.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.92 | +/- | 1. |
| 1.92 | +/- | -NaN |
| -2.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| 0.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 93.80 | +/- | 0. |
| 1.97 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.49 |
| -9999.00 |

PERSIST_LOW,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,ROTATION_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
ANDR3
| 3810. | +/- | 37. |
| -10000. | +/- | -NaN |
| 1.41 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.80 | +/- | 0. |
| 0.80 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.76 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -0.41 |
| -9999.00 |
PERSIST_LOW,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 6299. | +/- | 200. |
| -10000. | +/- | -NaN |
| 4.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.78 | +/- | 1. |
| 1.78 | +/- | -NaN |
| -2.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| -0.61 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.57 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -2.20 |
| -9999.00 |
SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,ROTATION_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
ANDR3
| 3930. | +/- | 27. |
| -10000. | +/- | -NaN |
| 1.59 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.80 | +/- | 0. |
| 0.80 | +/- | -NaN |
| -2.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.60 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -1.96 |
| -9999.00 |
PERSIST_LOW,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
ANDR3
| 3945. | +/- | 18. |
| -10000. | +/- | -NaN |
| 0.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.69 | +/- | 0. |
| 0.69 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.42 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.76 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
LOW_SNR,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
ANDR3
| 3902. | +/- | 85. |
| -10000. | +/- | -NaN |
| 1.57 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.84 | +/- | 1. |
| 2.84 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.16 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.75 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.04 |
| -9999.00 |
BRIGHT_NEIGHBOR,PERSIST_MED,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
ANDR3
| 6194. | +/- | 128. |
| 6190. | +/- | 219. |
| 4.56 | +/- | 0. |
| 5.38 | +/- | 0. |
| 1.84 | +/- | 2. |
| 1.84 | +/- | -NaN |
| -2.42 | +/- | 0. |
| -2.42 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.10 | +/- | 6. |
| 0.10 | +/- | -NaN |
| -0.64 | +/- | 0. |
| -0.65 | +/- | 0. |
| 3.24 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.34 |
| -9999.00 |
PERSIST_LOW,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,ROTATION_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
ANDR3
| 4768. | +/- | 214. |
| -10000. | +/- | -NaN |
| 4.30 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.67 | +/- | 5. |
| 0.67 | +/- | -NaN |
| -2.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.18 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.34 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.64 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.55 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.41 |
| -9999.00 |

SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD
ANDR3
| 6259. | +/- | 48. |
| -10000. | +/- | -NaN |
| 4.22 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.47 | +/- | 1. |
| 1.47 | +/- | -NaN |
| -2.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| -0.52 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.92 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| 0.86 |
| -9999.00 |
LOW_SNR,PERSIST_MED,PERSIST_LOW,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 3961. | +/- | 75. |
| -10000. | +/- | -NaN |
| 4.92 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 3.58 | +/- | 1. |
| 3.58 | +/- | -NaN |
| -2.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.91 | +/- | 0. |
| 0.77 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| -2.49 |
| -9999.00 |
SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
ANDR3
| 8601. | +/- | 119. |
| 8489. | +/- | 447. |
| 4.19 | +/- | 0. |
| 4.64 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.11 | +/- | 1. |
| 1.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -1.98 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.10 |
| -9999.00 |
PERSIST_LOW,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD
STAR_WARN,SN_WARN
ANDR3
| 3723. | +/- | 33. |
| -10000. | +/- | -NaN |
| 0.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.76 | +/- | 0. |
| 0.76 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.41 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.77 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.50 |
| -9999.00 |
PERSIST_LOW,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD
ANDR3
| 7280. | +/- | 47. |
| -10000. | +/- | -NaN |
| 5.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.47 | +/- | 3. |
| 1.47 | +/- | -NaN |
| -2.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.74 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -2.42 |
| -9999.00 |
SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES
STAR_BAD,ROTATION_BAD
STAR_WARN,ROTATION_WARN
ANDR3
| 3758. | +/- | 3. |
| -10000. | +/- | -NaN |
| 0.65 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.47 | +/- | 0. |
| 0.47 | +/- | -NaN |
| -2.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.04 | +/- | 0. |
| 1.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -1.93 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -1.62 |
| -9999.00 |
SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,ROTATION_BAD
STAR_WARN,ROTATION_WARN
ANDR3
| 3952. | +/- | 21. |
| -10000. | +/- | -NaN |
| 2.76 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.76 | +/- | 0. |
| 0.76 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.74 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -2.46 |
| -9999.00 |
LOW_SNR,PERSIST_MED,PERSIST_JUMP_NEG,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 3746. | +/- | 52. |
| -10000. | +/- | -NaN |
| 2.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.77 | +/- | 0. |
| 0.77 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.37 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.76 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.23 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.71 |
| -9999.00 |
PERSIST_MED,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 3895. | +/- | 31. |
| -10000. | +/- | -NaN |
| 1.67 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.79 | +/- | 1. |
| 0.79 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.74 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 1.00 |
| -9999.00 |
PERSIST_MED,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD
ANDR3
| 7782. | +/- | 28. |
| -10000. | +/- | -NaN |
| 5.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.21 | +/- | 1. |
| 3.21 | +/- | -NaN |
| -2.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| -0.72 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.65 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -2.21 |
| -9999.00 |
SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 5401. | +/- | 342. |
| -10000. | +/- | -NaN |
| 4.38 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 1.54 | +/- | 17. |
| 1.54 | +/- | -NaN |
| -2.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.24 | +/- | 8. |
| -9999.99 | +/- | -NaN |
| -0.08 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| -0.74 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.51 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.57 |
| -9999.00 |

LOW_SNR,PERSIST_LOW,PERSIST_JUMP_NEG,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
ANDR3
| 3794. | +/- | 130. |
| -10000. | +/- | -NaN |
| 2.99 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 2.88 | +/- | 1. |
| 2.88 | +/- | -NaN |
| -2.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.69 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.36 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| 1.68 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -2.49 |
| -9999.00 |
SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD
ANDR3
| 6041. | +/- | 98. |
| -10000. | +/- | -NaN |
| 4.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.77 | +/- | 1. |
| 1.77 | +/- | -NaN |
| -2.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| -0.41 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.53 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.40 |
| -9999.00 |
SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,ROTATION_BAD
STAR_WARN,ROTATION_WARN
ANDR3
| 3870. | +/- | 11. |
| -10000. | +/- | -NaN |
| 1.22 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.70 | +/- | 0. |
| 0.70 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.74 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.29 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
LOW_SNR,PERSIST_LOW,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
ANDR3
| 3742. | +/- | 29. |
| -10000. | +/- | -NaN |
| 0.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.78 | +/- | 0. |
| 0.78 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.74 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -2.14 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.49 |
| -9999.00 |
LOW_SNR,PERSIST_LOW,PERSIST_JUMP_NEG,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
ANDR3
| 5204. | +/- | 662. |
| -10000. | +/- | -NaN |
| 4.43 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 2.24 | +/- | 5. |
| 2.24 | +/- | -NaN |
| -2.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 12.55 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.18 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.49 |
| -9999.00 |

PERSIST_MED,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD
ANDR3
| 7414. | +/- | 66. |
| -10000. | +/- | -NaN |
| 5.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.33 | +/- | 1. |
| 1.33 | +/- | -NaN |
| -2.42 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| -0.59 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.54 | +/- | 1. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -2.03 |
| -9999.00 |
SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,ROTATION_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
ANDR3
| 3808. | +/- | 45. |
| -10000. | +/- | -NaN |
| 2.87 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.89 | +/- | 1. |
| 0.89 | +/- | -NaN |
| -2.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.26 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.42 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.29 | +/- | 0. |
| 1.09 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.26 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,ROTATION_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
ANDR3
| 3713. | +/- | 19. |
| -10000. | +/- | -NaN |
| 0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.68 | +/- | 0. |
| 1.68 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.77 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.03 |
| -9999.00 |
LOW_SNR,PERSIST_LOW,PERSIST_JUMP_NEG,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 6605. | +/- | 214. |
| -10000. | +/- | -NaN |
| 3.57 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.54 | +/- | 1. |
| 0.54 | +/- | -NaN |
| -1.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 1.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 79.12 | +/- | 0. |
| 1.90 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -2.42 |
| -9999.00 |
LOW_SNR,PERSIST_LOW,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
ANDR3
| 4478. | +/- | 258. |
| -10000. | +/- | -NaN |
| 2.94 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 3.86 | +/- | 1. |
| 3.86 | +/- | -NaN |
| -2.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.11 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.28 | +/- | 0. |
| 1.09 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| 0.40 |
| -9999.00 |

LOW_SNR,PERSIST_LOW,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 6675. | +/- | 173. |
| -10000. | +/- | -NaN |
| 3.76 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.68 | +/- | 12. |
| 0.68 | +/- | -NaN |
| -2.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 7. |
| -9999.99 | +/- | -NaN |
| -0.29 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 5.46 | +/- | 3. |
| 0.74 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| 0.59 |
| -9999.00 |
PERSIST_MED,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,ROTATION_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
ANDR3
| 3599. | +/- | 16. |
| -10000. | +/- | -NaN |
| 0.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.69 | +/- | 0. |
| 0.69 | +/- | -NaN |
| -2.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.45 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.25 | +/- | 0. |
| 1.09 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -2.43 |
| -9999.00 |
SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,ROTATION_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
ANDR3
| 3893. | +/- | 28. |
| -10000. | +/- | -NaN |
| 1.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.79 | +/- | 0. |
| 0.79 | +/- | -NaN |
| -2.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.72 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.23 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.78 |
| -9999.00 |
LOW_SNR,PERSIST_LOW,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 5564. | +/- | 474. |
| -10000. | +/- | -NaN |
| 4.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.56 | +/- | 14. |
| 0.56 | +/- | -NaN |
| -2.41 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 77.21 | +/- | 1. |
| 1.89 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -1.93 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD
STAR_WARN,SN_WARN
ANDR3
| 3815. | +/- | 20. |
| -10000. | +/- | -NaN |
| 0.31 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.63 | +/- | 0. |
| 1.63 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.26 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.46 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.77 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.24 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.46 |
| -9999.00 |
LOW_SNR,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 3692. | +/- | 185. |
| -10000. | +/- | -NaN |
| 4.29 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 1.55 | +/- | 1. |
| 1.55 | +/- | -NaN |
| -2.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.15 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 11.74 | +/- | 0. |
| 1.07 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.48 |
| -9999.00 |

SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,ROTATION_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
ANDR3
| 3846. | +/- | 27. |
| -10000. | +/- | -NaN |
| 0.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.62 | +/- | 0. |
| 2.62 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.75 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.25 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -2.28 |
| -9999.00 |
LOW_SNR,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 5956. | +/- | 750. |
| -10000. | +/- | -NaN |
| 4.52 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.12 | +/- | 8. |
| 1.12 | +/- | -NaN |
| -2.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 16. |
| -9999.99 | +/- | -NaN |
| -0.44 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 50.54 | +/- | 1. |
| 1.70 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| 0.04 |
| -9999.00 |

LOW_SNR,PERSIST_LOW,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 3770. | +/- | 32. |
| -10000. | +/- | -NaN |
| 0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.77 | +/- | 0. |
| 2.77 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.75 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.27 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -2.48 |
| -9999.00 |
LOW_SNR,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 5030. | +/- | 441. |
| -10000. | +/- | -NaN |
| 5.20 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 1.06 | +/- | 4. |
| 1.06 | +/- | -NaN |
| -2.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 0.69 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 45.82 | +/- | 1. |
| 1.66 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.32 |
| -9999.00 |
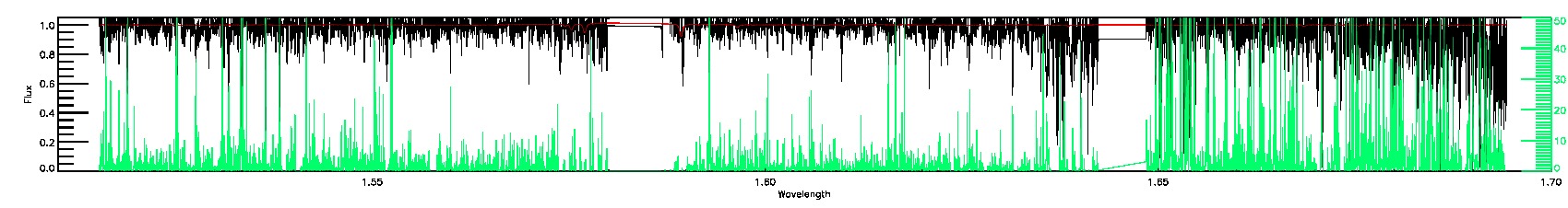
LOW_SNR,PERSIST_LOW,PERSIST_JUMP_NEG,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 3631. | +/- | 84. |
| -10000. | +/- | -NaN |
| 2.89 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.59 | +/- | 0. |
| 0.59 | +/- | -NaN |
| -2.41 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.08 | +/- | 0. |
| 1.08 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.76 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 1.99 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -2.25 |
| -9999.00 |
LOW_SNR,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 4015. | +/- | 166. |
| -10000. | +/- | -NaN |
| 4.60 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 1.53 | +/- | 1. |
| 1.53 | +/- | -NaN |
| -2.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.65 | +/- | 1. |
| 0.75 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -0.62 |
| -9999.00 |

LOW_SNR,PERSIST_LOW,PERSIST_JUMP_NEG,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 3818. | +/- | 112. |
| -10000. | +/- | -NaN |
| 2.84 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.78 | +/- | 2. |
| 0.78 | +/- | -NaN |
| -2.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.24 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.61 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.57 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| 2.00 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -1.69 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
LOW_SNR,PERSIST_LOW,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
ANDR3
| 4843. | +/- | 379. |
| -10000. | +/- | -NaN |
| 3.92 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.60 | +/- | 3. |
| 0.60 | +/- | -NaN |
| -2.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| -0.24 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.53 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.42 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.45 |
| -9999.00 |

LOW_SNR,PERSIST_JUMP_NEG,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 4109. | +/- | 72. |
| -10000. | +/- | -NaN |
| 3.73 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.10 | +/- | 0. |
| 1.10 | +/- | -NaN |
| -0.54 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.45 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 94.17 | +/- | 0. |
| 1.97 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 1.45 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.88 |
| -9999.00 |

LOW_SNR,PERSIST_MED,PERSIST_JUMP_NEG,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 5325. | +/- | 714. |
| -10000. | +/- | -NaN |
| 4.44 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 3.31 | +/- | 5. |
| 3.31 | +/- | -NaN |
| -2.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.41 | +/- | 8. |
| -9999.99 | +/- | -NaN |
| -0.20 | +/- | 6. |
| -9999.99 | +/- | -NaN |
| -0.41 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.22 | +/- | 0. |
| 1.09 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.58 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.42 |
| -9999.00 |

LOW_SNR,PERSIST_LOW,PERSIST_JUMP_NEG,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 5329. | +/- | 1501. |
| -10000. | +/- | -NaN |
| 4.84 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 0.50 | +/- | 56. |
| 0.50 | +/- | -NaN |
| -2.41 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.20 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| -0.34 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 1.50 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.41 |
| -9999.00 |

LOW_SNR,PERSIST_LOW,PERSIST_JUMP_POS,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 6208. | +/- | 757. |
| -10000. | +/- | -NaN |
| 4.94 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 3.57 | +/- | 1. |
| 3.57 | +/- | -NaN |
| -1.35 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| -0.11 | +/- | 8. |
| -9999.99 | +/- | -NaN |
| 0.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.76 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.47 |
| -9999.00 |
VERY_BRIGHT_NEIGHBOR,LOW_SNR,PERSIST_MED,PERSIST_JUMP_NEG,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 5613. | +/- | 912. |
| -10000. | +/- | -NaN |
| 4.73 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.65 | +/- | 19. |
| 0.65 | +/- | -NaN |
| -2.46 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 6. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.17 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 1.98 | +/- | 7. |
| 0.30 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -2.14 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.46 |
| -9999.00 |
LOW_SNR,PERSIST_LOW,PERSIST_JUMP_NEG,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
ANDR3
| 3357. | +/- | 65. |
| -10000. | +/- | -NaN |
| 1.96 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.17 | +/- | 1. |
| 1.17 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.24 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.29 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.76 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.41 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 1.95 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.50 |
| -9999.00 |
VERY_BRIGHT_NEIGHBOR,LOW_SNR,PERSIST_JUMP_NEG,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
ANDR3
| 5293. | +/- | 167. |
| -10000. | +/- | -NaN |
| 2.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.93 | +/- | 0. |
| 0.93 | +/- | -NaN |
| -1.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.34 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.62 | +/- | 0. |
| 0.75 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
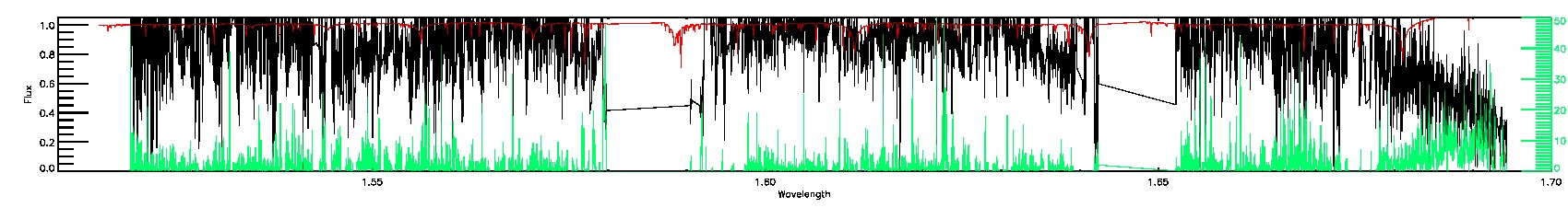
BRIGHT_NEIGHBOR,LOW_SNR,PERSIST_JUMP_NEG,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
ANDR3
| 4167. | +/- | 79. |
| -10000. | +/- | -NaN |
| 5.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.64 | +/- | 1. |
| 0.64 | +/- | -NaN |
| -0.58 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.47 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 82.58 | +/- | 0. |
| 1.92 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.50 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 1.47 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -1.65 |
| -9999.00 |

BRIGHT_NEIGHBOR,VERY_BRIGHT_NEIGHBOR,LOW_SNR,PERSIST_JUMP_NEG,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
ANDR3
| 3118. | +/- | 50. |
| -10000. | +/- | -NaN |
| 2.90 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.74 | +/- | 1. |
| 1.74 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.75 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.27 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.50 |
| -9999.00 |
SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES
ANDR3
| 6433. | +/- | 32. |
| 6296. | +/- | 175. |
| 4.25 | +/- | 0. |
| 4.02 | +/- | 0. |
| 1.08 | +/- | 0. |
| 1.08 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.29 | +/- | 0. |
| 0.29 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.08 | +/- | 0. |
| 48.51 | +/- | 0. |
| 1.69 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -2.28 |
| -9999.00 |
