STAR_WARN,COLORTE_WARN
010-07-C
| 12117. | +/- | 128. |
| -10000. | +/- | -NaN |
| 4.41 | +/- | 0. |
| 4.03 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 27.28 | +/- | 0. |
| 1.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.08 |
| -9999.00 |
STAR_WARN,SN_WARN
010-07-C
| 4647. | +/- | 26. |
| 4752. | +/- | 128. |
| 2.44 | +/- | 0. |
| 2.32 | +/- | 0. |
| 1.51 | +/- | 0. |
| 1.51 | +/- | -NaN |
| -0.81 | +/- | 0. |
| -0.81 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.26 | +/- | 0. |
| 4.74 | +/- | 0. |
| 0.68 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.48 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4241. | +/- | 4. |
| 4315. | +/- | 94. |
| 2.47 | +/- | 0. |
| 2.20 | +/- | 0. |
| 1.22 | +/- | 0. |
| 1.22 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.37 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.45 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| 0.63 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
010-07-C
| 3789. | +/- | 2. |
| 3863. | +/- | 80. |
| 1.69 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.26 | +/- | 0. |
| 1.26 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.44 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.37 |
| -9999.00 |
010-07-C
| 8560. | +/- | 84. |
| 8419. | +/- | 399. |
| 3.87 | +/- | 0. |
| 4.02 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -1.41 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.13 | +/- | 0. |
| 1.01 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.41 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -1.40 |
| -9999.00 |
010-07-C
| 3874. | +/- | 2. |
| 3969. | +/- | 86. |
| 1.49 | +/- | 0. |
| 1.32 | +/- | 0. |
| 1.67 | +/- | 0. |
| 1.67 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -0.16 | +/- | 0. |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.22 | +/- | 0. |
| 3.24 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.50 |
| -9999.00 |
010-07-C
| 3767. | +/- | 2. |
| 3860. | +/- | 83. |
| 1.28 | +/- | 0. |
| 1.14 | +/- | 0. |
| 1.76 | +/- | 0. |
| 1.76 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | 0. |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.21 | +/- | 0. |
| 3.16 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.14 |
| -9999.00 |
010-07-C
| 3841. | +/- | 2. |
| 3953. | +/- | 88. |
| 1.16 | +/- | 0. |
| 1.08 | +/- | 0. |
| 2.08 | +/- | 0. |
| 2.08 | +/- | -NaN |
| -0.55 | +/- | 0. |
| -0.55 | +/- | 0. |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.26 | +/- | 0. |
| 4.07 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.05 |
| -9999.00 |
STAR_WARN,SN_WARN
010-07-C
| 4316. | +/- | 9. |
| 4391. | +/- | 102. |
| 2.45 | +/- | 0. |
| 2.26 | +/- | 0. |
| 1.22 | +/- | 0. |
| 1.22 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.46 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -0.79 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,ROTATION_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
010-07-C
| 4250. | +/- | 12. |
| -10000. | +/- | -NaN |
| 2.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.47 | +/- | 0. |
| 0.47 | +/- | -NaN |
| -0.55 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.07 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.27 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.11 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 3794. | +/- | 3. |
| 3870. | +/- | 83. |
| 1.45 | +/- | 0. |
| 1.23 | +/- | 0. |
| 1.51 | +/- | 0. |
| 1.51 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.29 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.50 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| 0.60 |
| -9999.00 |
010-07-C
| 4106. | +/- | 2. |
| 4192. | +/- | 88. |
| 1.99 | +/- | 0. |
| 1.75 | +/- | 0. |
| 1.37 | +/- | 0. |
| 1.37 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.07 | +/- | 0. |
| 2.82 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| -0.46 |
| -9999.00 |

010-07-C
| 4720. | +/- | 10. |
| 4780. | +/- | 108. |
| 3.08 | +/- | 0. |
| 2.88 | +/- | 0. |
| 1.19 | +/- | 0. |
| 1.19 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.85 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -0.04 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4388. | +/- | 13. |
| 4503. | +/- | 116. |
| 2.29 | +/- | 0. |
| 2.14 | +/- | 0. |
| 1.49 | +/- | 0. |
| 1.49 | +/- | -NaN |
| -0.63 | +/- | 0. |
| -0.63 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.20 | +/- | 0. |
| 4.26 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.32 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4437. | +/- | 10. |
| 4513. | +/- | 104. |
| 2.89 | +/- | 0. |
| 2.64 | +/- | 0. |
| 0.64 | +/- | 0. |
| 0.64 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.04 | +/- | 0. |
| 2.48 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.21 |
| -9999.00 |

010-07-C
| 3852. | +/- | 2. |
| 3938. | +/- | 85. |
| 1.40 | +/- | 0. |
| 1.21 | +/- | 0. |
| 1.62 | +/- | 0. |
| 1.62 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.07 | +/- | 0. |
| 2.83 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.23 |
| -9999.00 |
STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 4410. | +/- | 9. |
| 4480. | +/- | 102. |
| 2.70 | +/- | 0. |
| 2.42 | +/- | 0. |
| 1.10 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.44 | +/- | 0. |
| 0.44 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.41 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.28 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 1.00 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
010-07-C
| 4463. | +/- | 15. |
| -10000. | +/- | -NaN |
| 2.66 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.13 | +/- | 0. |
| 1.13 | +/- | -NaN |
| 0.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.09 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4319. | +/- | 8. |
| 4430. | +/- | 112. |
| 1.92 | +/- | 0. |
| 1.76 | +/- | 0. |
| 1.53 | +/- | 0. |
| 1.53 | +/- | -NaN |
| -0.53 | +/- | 0. |
| -0.52 | +/- | 0. |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.29 | +/- | 0. |
| 4.02 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.21 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -0.55 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4562. | +/- | 12. |
| 4643. | +/- | 112. |
| 2.63 | +/- | 0. |
| 2.34 | +/- | 0. |
| 1.44 | +/- | 0. |
| 1.44 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.08 | +/- | 0. |
| 2.94 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.48 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4718. | +/- | 14. |
| 4800. | +/- | 118. |
| 2.56 | +/- | 0. |
| 2.31 | +/- | 0. |
| 1.70 | +/- | 0. |
| 1.70 | +/- | -NaN |
| -0.43 | +/- | 0. |
| -0.43 | +/- | 0. |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.23 | +/- | 0. |
| 3.81 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.77 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4532. | +/- | 17. |
| 4617. | +/- | 114. |
| 2.53 | +/- | 0. |
| 2.30 | +/- | 0. |
| 1.38 | +/- | 0. |
| 1.38 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.06 | +/- | 0. |
| 2.95 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.40 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4410. | +/- | 15. |
| 4529. | +/- | 120. |
| 1.96 | +/- | 0. |
| 1.82 | +/- | 0. |
| 1.69 | +/- | 0. |
| 1.69 | +/- | -NaN |
| -0.72 | +/- | 0. |
| -0.72 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.27 | +/- | 0. |
| 4.50 | +/- | 0. |
| 0.65 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.25 |
| -9999.00 |

010-07-C
| 4452. | +/- | 6. |
| 4533. | +/- | 100. |
| 2.80 | +/- | 0. |
| 2.57 | +/- | 0. |
| 0.59 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.69 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 0.16 |
| -9999.00 |

STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 4515. | +/- | 12. |
| 4599. | +/- | 111. |
| 2.84 | +/- | 0. |
| 2.62 | +/- | 0. |
| 1.05 | +/- | 0. |
| 1.05 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.04 | +/- | 0. |
| 2.85 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| 0.85 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4376. | +/- | 10. |
| 4449. | +/- | 104. |
| 2.80 | +/- | 0. |
| 2.54 | +/- | 0. |
| 1.21 | +/- | 0. |
| 1.21 | +/- | -NaN |
| 0.36 | +/- | 0. |
| 0.36 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.39 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.96 |
| -9999.00 |

010-07-C
| 4669. | +/- | 10. |
| 4747. | +/- | 111. |
| 3.02 | +/- | 0. |
| 2.85 | +/- | 0. |
| 1.28 | +/- | 0. |
| 1.28 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -0.22 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.17 | +/- | 0. |
| 3.37 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.01 |
| -9999.00 |

010-07-C
| 4037. | +/- | 2. |
| 4171. | +/- | 89. |
| 1.10 | +/- | 0. |
| 1.09 | +/- | 0. |
| 2.10 | +/- | 0. |
| 2.10 | +/- | -NaN |
| -1.05 | +/- | 0. |
| -1.05 | +/- | 0. |
| -0.24 | +/- | 0. |
| -0.24 | +/- | -NaN |
| 0.92 | +/- | 0. |
| 0.92 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.24 | +/- | 0. |
| 5.47 | +/- | 0. |
| 0.74 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.26 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.93 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4442. | +/- | 7. |
| 4514. | +/- | 101. |
| 2.70 | +/- | 0. |
| 2.39 | +/- | 0. |
| 1.30 | +/- | 0. |
| 1.30 | +/- | -NaN |
| 0.38 | +/- | 0. |
| 0.39 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.36 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.42 |
| -9999.00 |

STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 3923. | +/- | 4. |
| 4027. | +/- | 93. |
| 1.30 | +/- | 0. |
| 1.19 | +/- | 0. |
| 1.49 | +/- | 0. |
| 1.49 | +/- | -NaN |
| -0.37 | +/- | 0. |
| -0.37 | +/- | 0. |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.19 | +/- | 0. |
| 3.67 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.25 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| -0.75 |
| -9999.00 |
010-07-C
| 4216. | +/- | 4. |
| 4350. | +/- | 105. |
| 1.58 | +/- | 0. |
| 1.52 | +/- | 0. |
| 1.82 | +/- | 0. |
| 1.82 | +/- | -NaN |
| -1.07 | +/- | 0. |
| -1.07 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.28 | +/- | 0. |
| 5.53 | +/- | 0. |
| 0.74 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -1.76 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4193. | +/- | 5. |
| 4266. | +/- | 96. |
| 2.34 | +/- | 0. |
| 2.06 | +/- | 0. |
| 1.20 | +/- | 0. |
| 1.20 | +/- | -NaN |
| 0.37 | +/- | 0. |
| 0.37 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.43 |
| -9999.00 |
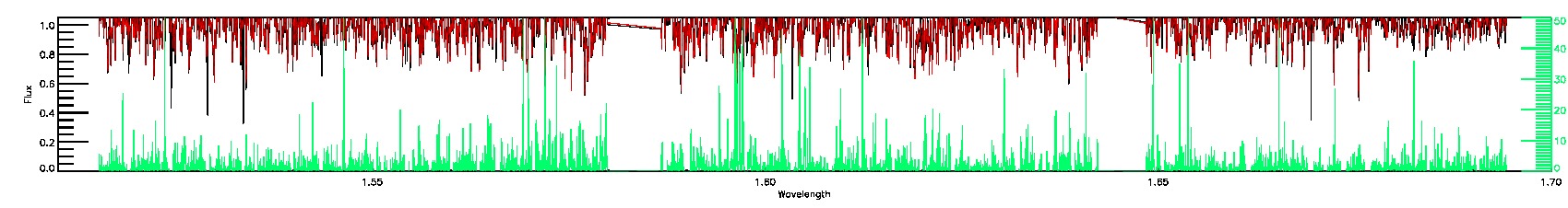
010-07-C
| 4295. | +/- | 3. |
| 4405. | +/- | 96. |
| 1.95 | +/- | 0. |
| 1.78 | +/- | 0. |
| 1.70 | +/- | 0. |
| 1.70 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -0.50 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.19 | +/- | 0. |
| 3.97 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.49 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4484. | +/- | 23. |
| 4620. | +/- | 128. |
| 1.92 | +/- | 0. |
| 1.84 | +/- | 0. |
| 1.33 | +/- | 0. |
| 1.33 | +/- | -NaN |
| -1.12 | +/- | 0. |
| -1.11 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.27 | +/- | 0. |
| 5.68 | +/- | 0. |
| 0.75 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| -1.95 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4573. | +/- | 15. |
| 4647. | +/- | 112. |
| 2.70 | +/- | 0. |
| 2.38 | +/- | 0. |
| 1.34 | +/- | 0. |
| 1.34 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.74 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.94 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4500. | +/- | 14. |
| 4589. | +/- | 113. |
| 2.89 | +/- | 0. |
| 2.68 | +/- | 0. |
| 0.94 | +/- | 0. |
| 0.94 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.12 | +/- | 0. |
| 2.96 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| -0.85 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4326. | +/- | 7. |
| 4410. | +/- | 102. |
| 2.39 | +/- | 0. |
| 2.14 | +/- | 0. |
| 1.29 | +/- | 0. |
| 1.29 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.80 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.89 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4480. | +/- | 12. |
| 4611. | +/- | 120. |
| 2.12 | +/- | 0. |
| 2.02 | +/- | 0. |
| 1.83 | +/- | 0. |
| 1.83 | +/- | -NaN |
| -1.00 | +/- | 0. |
| -1.00 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| 0.87 | +/- | 0. |
| 0.87 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.22 | +/- | 0. |
| 5.32 | +/- | 0. |
| 0.73 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -1.67 |
| -9999.00 |

010-07-C
| 4238. | +/- | 4. |
| 4346. | +/- | 102. |
| 2.12 | +/- | 0. |
| 1.94 | +/- | 0. |
| 1.37 | +/- | 0. |
| 1.37 | +/- | -NaN |
| -0.46 | +/- | 0. |
| -0.46 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.17 | +/- | 0. |
| 3.87 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.83 |
| -9999.00 |

BRIGHT_NEIGHBOR
STAR_WARN,SN_WARN
010-07-C
| 4201. | +/- | 8. |
| 4296. | +/- | 106. |
| 2.05 | +/- | 0. |
| 1.84 | +/- | 0. |
| 1.03 | +/- | 0. |
| 1.03 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -0.16 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.14 | +/- | 0. |
| 3.24 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| -0.30 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4552. | +/- | 19. |
| 4623. | +/- | 112. |
| 2.89 | +/- | 0. |
| 2.53 | +/- | 0. |
| 0.74 | +/- | 0. |
| 0.74 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.05 | +/- | 0. |
| 2.52 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 1.00 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4455. | +/- | 11. |
| 4542. | +/- | 108. |
| 2.45 | +/- | 0. |
| 2.22 | +/- | 0. |
| 1.34 | +/- | 0. |
| 1.34 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.29 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.88 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| 0.35 |
| -9999.00 |

010-07-C
| 4758. | +/- | 10. |
| 4810. | +/- | 107. |
| 2.89 | +/- | 0. |
| 2.53 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.43 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.72 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.46 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4205. | +/- | 4. |
| 4288. | +/- | 95. |
| 2.19 | +/- | 0. |
| 1.93 | +/- | 0. |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.11 | +/- | 0. |
| 2.74 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.12 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4399. | +/- | 11. |
| 4521. | +/- | 118. |
| 2.12 | +/- | 0. |
| 1.99 | +/- | 0. |
| 1.50 | +/- | 0. |
| 1.50 | +/- | -NaN |
| -0.78 | +/- | 0. |
| -0.78 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.24 | +/- | 0. |
| 4.68 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.88 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4324. | +/- | 6. |
| 4430. | +/- | 105. |
| 2.25 | +/- | 0. |
| 2.06 | +/- | 0. |
| 1.20 | +/- | 0. |
| 1.20 | +/- | -NaN |
| -0.41 | +/- | 0. |
| -0.41 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.18 | +/- | 0. |
| 3.77 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.86 |
| -9999.00 |

010-07-C
| 4181. | +/- | 4. |
| 4275. | +/- | 96. |
| 2.01 | +/- | 0. |
| 1.79 | +/- | 0. |
| 1.57 | +/- | 0. |
| 1.57 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.13 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 3.20 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.55 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 3962. | +/- | 3. |
| 4034. | +/- | 87. |
| 1.93 | +/- | 0. |
| 1.64 | +/- | 0. |
| 1.33 | +/- | 0. |
| 1.33 | +/- | -NaN |
| 0.38 | +/- | 0. |
| 0.38 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.37 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.61 |
| -9999.00 |
STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 4338. | +/- | 11. |
| 4448. | +/- | 113. |
| 1.99 | +/- | 0. |
| 1.82 | +/- | 0. |
| 1.52 | +/- | 0. |
| 1.52 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -0.49 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.22 | +/- | 0. |
| 3.94 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -1.05 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4514. | +/- | 15. |
| 4582. | +/- | 108. |
| 2.83 | +/- | 0. |
| 2.48 | +/- | 0. |
| 0.95 | +/- | 0. |
| 0.95 | +/- | -NaN |
| 0.45 | +/- | 0. |
| 0.45 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.04 | +/- | 0. |
| 2.27 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| 1.00 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
010-07-C
| 11894. | +/- | 196. |
| -10000. | +/- | -NaN |
| 4.44 | +/- | 0. |
| 4.25 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -0.56 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 14.15 | +/- | 0. |
| 1.15 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.70 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.71 |
| -9999.00 |
STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 4128. | +/- | 5. |
| 4215. | +/- | 97. |
| 1.97 | +/- | 0. |
| 1.73 | +/- | 0. |
| 1.38 | +/- | 0. |
| 1.38 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.88 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.06 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4904. | +/- | 21. |
| 4963. | +/- | 126. |
| 2.85 | +/- | 0. |
| 2.49 | +/- | 0. |
| 1.75 | +/- | 0. |
| 1.75 | +/- | -NaN |
| -0.42 | +/- | 0. |
| -0.41 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.13 | +/- | 0. |
| 3.77 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.44 |
| -9999.00 |

010-07-C
| 3938. | +/- | 2. |
| 4039. | +/- | 90. |
| 1.54 | +/- | 0. |
| 1.39 | +/- | 0. |
| 1.55 | +/- | 0. |
| 1.55 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -0.31 | +/- | 0. |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.21 | +/- | 0. |
| 3.54 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -1.01 |
| -9999.00 |
010-07-C
| 3884. | +/- | 2. |
| 3960. | +/- | 82. |
| 1.75 | +/- | 0. |
| 1.48 | +/- | 0. |
| 1.52 | +/- | 0. |
| 1.52 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.48 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.58 |
| -9999.00 |
STAR_WARN,SN_WARN
010-07-C
| 3844. | +/- | 2. |
| 3936. | +/- | 87. |
| 1.31 | +/- | 0. |
| 1.17 | +/- | 0. |
| 1.70 | +/- | 0. |
| 1.70 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.10 | +/- | 0. |
| 3.12 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.38 |
| -9999.00 |
STAR_WARN,SN_WARN
010-07-C
| 4230. | +/- | 6. |
| 4315. | +/- | 101. |
| 2.34 | +/- | 0. |
| 2.10 | +/- | 0. |
| 1.21 | +/- | 0. |
| 1.21 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.84 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.21 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4477. | +/- | 11. |
| 4550. | +/- | 106. |
| 2.73 | +/- | 0. |
| 2.41 | +/- | 0. |
| 1.33 | +/- | 0. |
| 1.33 | +/- | -NaN |
| 0.37 | +/- | 0. |
| 0.37 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.45 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_WARN,COLORTE_WARN
010-07-C
| 9120. | +/- | 107. |
| 8987. | +/- | 445. |
| 4.17 | +/- | 0. |
| 4.41 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -1.61 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.07 | +/- | 0. |
| 1.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.75 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.29 |
| -9999.00 |
010-07-C
| 4648. | +/- | 11. |
| 4739. | +/- | 114. |
| 2.77 | +/- | 0. |
| 2.62 | +/- | 0. |
| 1.23 | +/- | 0. |
| 1.23 | +/- | -NaN |
| -0.46 | +/- | 0. |
| -0.46 | +/- | 0. |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.19 | +/- | 0. |
| 3.86 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -1.25 |
| -9999.00 |

010-07-C
| 4908. | +/- | 6. |
| 4945. | +/- | 94. |
| 3.04 | +/- | 0. |
| 2.66 | +/- | 0. |
| 1.50 | +/- | 0. |
| 1.50 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.82 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.85 |
| -9999.00 |
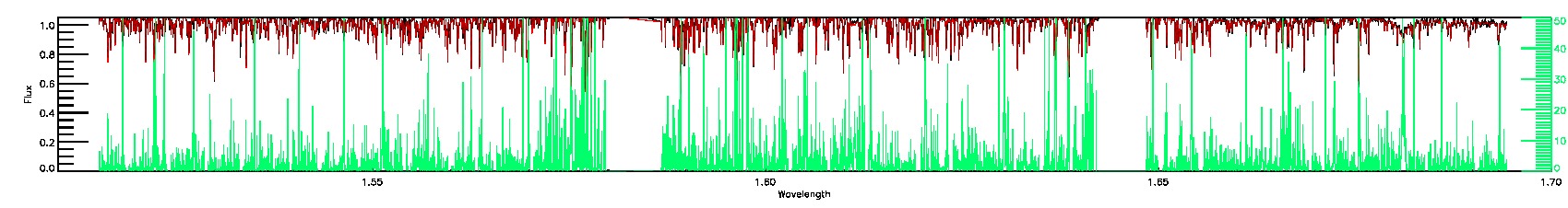
010-07-C
| 8313. | +/- | 36. |
| 8189. | +/- | 316. |
| 4.38 | +/- | 0. |
| 4.71 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -1.83 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 55.72 | +/- | 0. |
| 1.75 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.30 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -1.48 |
| -9999.00 |
STAR_WARN,SN_WARN
010-07-C
| 4390. | +/- | 13. |
| 4551. | +/- | 124. |
| 1.46 | +/- | 0. |
| 1.50 | +/- | 0. |
| 1.01 | +/- | 0. |
| 1.01 | +/- | -NaN |
| -1.70 | +/- | 0. |
| -1.70 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -0.29 | +/- | -NaN |
| 0.42 | +/- | 0. |
| 0.39 | +/- | 0. |
| 7.98 | +/- | 0. |
| 0.90 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.15 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -1.69 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.48 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4127. | +/- | 7. |
| 4203. | +/- | 98. |
| 2.19 | +/- | 0. |
| 1.92 | +/- | 0. |
| 1.29 | +/- | 0. |
| 1.29 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.49 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.05 | +/- | 0. |
| 2.51 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.10 |
| -9999.00 |
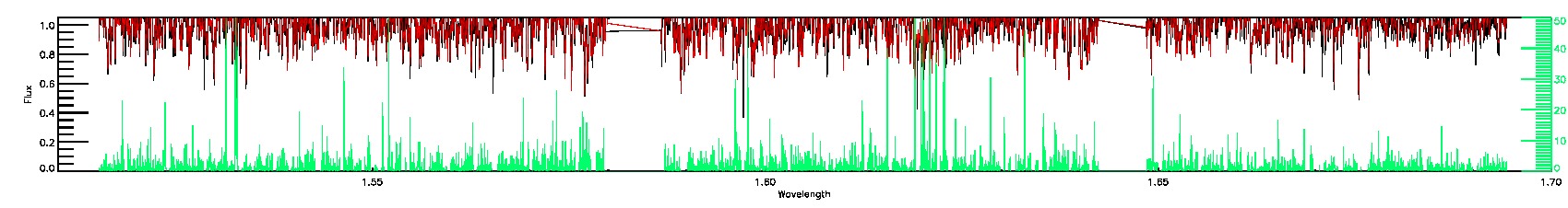
BRIGHT_NEIGHBOR
010-07-C
| 4157. | +/- | 3. |
| 4272. | +/- | 99. |
| 1.50 | +/- | 0. |
| 1.40 | +/- | 0. |
| 1.78 | +/- | 0. |
| 1.78 | +/- | -NaN |
| -0.62 | +/- | 0. |
| -0.62 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.24 | +/- | 0. |
| 4.26 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -1.07 |
| -9999.00 |

010-07-C
| 4452. | +/- | 7. |
| 4572. | +/- | 111. |
| 2.19 | +/- | 0. |
| 2.05 | +/- | 0. |
| 1.55 | +/- | 0. |
| 1.55 | +/- | -NaN |
| -0.75 | +/- | 0. |
| -0.75 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.27 | +/- | 0. |
| 4.58 | +/- | 0. |
| 0.66 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -0.99 |
| -9999.00 |

010-07-C
| 3736. | +/- | 1. |
| 3819. | +/- | 77. |
| 1.13 | +/- | 0. |
| 0.96 | +/- | 0. |
| 1.70 | +/- | 0. |
| 1.70 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.06 | +/- | 0. |
| 2.75 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.22 |
| -9999.00 |
010-07-C
| 4144. | +/- | 3. |
| 4273. | +/- | 98. |
| 1.46 | +/- | 0. |
| 1.40 | +/- | 0. |
| 1.93 | +/- | 0. |
| 1.93 | +/- | -NaN |
| -0.96 | +/- | 0. |
| -0.96 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.28 | +/- | 0. |
| 5.18 | +/- | 0. |
| 0.71 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -1.30 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4420. | +/- | 9. |
| 4509. | +/- | 107. |
| 2.61 | +/- | 0. |
| 2.38 | +/- | 0. |
| 1.12 | +/- | 0. |
| 1.12 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 2.97 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.51 |
| -9999.00 |

010-07-C
| 8603. | +/- | 89. |
| 8483. | +/- | 428. |
| 4.22 | +/- | 0. |
| 4.59 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -1.92 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 17.11 | +/- | 0. |
| 1.23 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.76 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| -1.98 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.94 |
| -9999.00 |
STAR_WARN,SN_WARN
010-07-C
| 4624. | +/- | 18. |
| 4719. | +/- | 121. |
| 2.69 | +/- | 0. |
| 2.53 | +/- | 0. |
| 1.27 | +/- | 0. |
| 1.27 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -0.49 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.20 | +/- | 0. |
| 3.93 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.91 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4980. | +/- | 26. |
| 5042. | +/- | 132. |
| 2.57 | +/- | 0. |
| 2.31 | +/- | 0. |
| 1.90 | +/- | 0. |
| 1.90 | +/- | -NaN |
| -0.69 | +/- | 0. |
| -0.69 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.17 | +/- | 0. |
| 4.44 | +/- | 0. |
| 0.65 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| -0.55 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4601. | +/- | 11. |
| 4681. | +/- | 111. |
| 2.79 | +/- | 0. |
| 2.59 | +/- | 0. |
| 1.25 | +/- | 0. |
| 1.25 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | 0. |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.16 | +/- | 0. |
| 3.11 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.21 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| -0.61 |
| -9999.00 |

010-07-C
| 4126. | +/- | 4. |
| 4259. | +/- | 103. |
| 1.34 | +/- | 0. |
| 1.31 | +/- | 0. |
| 2.13 | +/- | 0. |
| 2.13 | +/- | -NaN |
| -1.04 | +/- | 0. |
| -1.04 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.20 | +/- | 0. |
| 5.44 | +/- | 0. |
| 0.74 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -1.62 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4144. | +/- | 6. |
| 4214. | +/- | 96. |
| 2.35 | +/- | 0. |
| 2.05 | +/- | 0. |
| 1.33 | +/- | 0. |
| 1.33 | +/- | -NaN |
| 0.45 | +/- | 0. |
| 0.45 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.28 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.18 |
| -9999.00 |
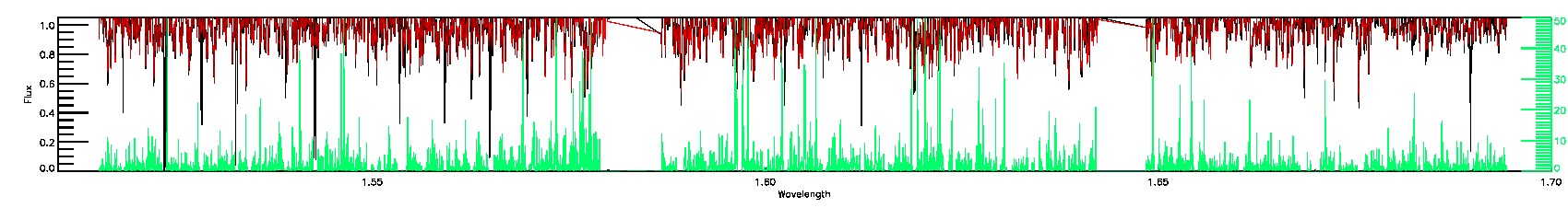
010-07-C
| 3843. | +/- | 2. |
| 3912. | +/- | 81. |
| 1.82 | +/- | 0. |
| 1.54 | +/- | 0. |
| 1.31 | +/- | 0. |
| 1.31 | +/- | -NaN |
| 0.44 | +/- | 0. |
| 0.44 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.29 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.24 |
| -9999.00 |
STAR_WARN,SN_WARN
010-07-C
| 4467. | +/- | 12. |
| 4534. | +/- | 106. |
| 2.69 | +/- | 0. |
| 2.38 | +/- | 0. |
| 1.29 | +/- | 0. |
| 1.29 | +/- | -NaN |
| 0.51 | +/- | 0. |
| 0.51 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.42 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.20 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.77 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4918. | +/- | 23. |
| 4953. | +/- | 121. |
| 3.86 | +/- | 0. |
| 3.84 | +/- | 0. |
| 0.58 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.04 | +/- | 0. |
| 2.77 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.24 |
| -9999.00 |

STAR_BAD
STAR_WARN,COLORTE_WARN
010-07-C
| 7997. | +/- | 13. |
| -10000. | +/- | -NaN |
| 4.94 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.24 | +/- | 1. |
| 1.24 | +/- | -NaN |
| -2.46 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.16 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.29 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.17 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.31 |
| -9999.00 |
010-07-C
| 3933. | +/- | 3. |
| 4042. | +/- | 92. |
| 1.47 | +/- | 0. |
| 1.35 | +/- | 0. |
| 1.66 | +/- | 0. |
| 1.66 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -0.49 | +/- | 0. |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.29 | +/- | 0. |
| 3.95 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.48 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_WARN,COLORTE_WARN
010-07-C
| 14910. | +/- | 132. |
| -10000. | +/- | -NaN |
| 4.74 | +/- | 0. |
| 4.34 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 33.18 | +/- | 0. |
| 1.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.23 |
| -9999.00 |
STAR_WARN,SN_WARN
010-07-C
| 4121. | +/- | 3. |
| 4231. | +/- | 99. |
| 1.72 | +/- | 0. |
| 1.57 | +/- | 0. |
| 1.70 | +/- | 0. |
| 1.70 | +/- | -NaN |
| -0.51 | +/- | 0. |
| -0.51 | +/- | 0. |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.21 | +/- | 0. |
| 3.99 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.33 |
| -9999.00 |

BRIGHT_NEIGHBOR
010-07-C
| 4153. | +/- | 3. |
| 4224. | +/- | 89. |
| 2.28 | +/- | 0. |
| 1.99 | +/- | 0. |
| 1.36 | +/- | 0. |
| 1.36 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.40 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.44 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.35 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.67 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4565. | +/- | 13. |
| 4660. | +/- | 116. |
| 2.74 | +/- | 0. |
| 2.57 | +/- | 0. |
| 1.36 | +/- | 0. |
| 1.36 | +/- | -NaN |
| -0.34 | +/- | 0. |
| -0.34 | +/- | 0. |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.23 | +/- | 0. |
| 3.60 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| 0.99 |
| -9999.00 |

010-07-C
| 4674. | +/- | 7. |
| 4753. | +/- | 104. |
| 2.69 | +/- | 0. |
| 2.50 | +/- | 0. |
| 1.42 | +/- | 0. |
| 1.42 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.24 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.17 | +/- | 0. |
| 3.41 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.19 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 3924. | +/- | 4. |
| 4039. | +/- | 99. |
| 1.45 | +/- | 0. |
| 1.35 | +/- | 0. |
| 1.77 | +/- | 0. |
| 1.77 | +/- | -NaN |
| -0.64 | +/- | 0. |
| -0.64 | +/- | 0. |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.22 | +/- | 0. |
| 4.29 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.04 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
010-07-C
| 4560. | +/- | 21. |
| -10000. | +/- | -NaN |
| 2.76 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.72 | +/- | 0. |
| 0.72 | +/- | -NaN |
| 0.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.54 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.39 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.08 |
| -9999.00 |

010-07-C
| 4704. | +/- | 6. |
| 4771. | +/- | 97. |
| 2.76 | +/- | 0. |
| 2.43 | +/- | 0. |
| 1.39 | +/- | 0. |
| 1.39 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.05 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| -0.13 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4574. | +/- | 20. |
| 4715. | +/- | 131. |
| 1.99 | +/- | 0. |
| 1.95 | +/- | 0. |
| 1.87 | +/- | 0. |
| 1.87 | +/- | -NaN |
| -1.43 | +/- | 0. |
| -1.43 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.15 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.24 | +/- | 0. |
| 6.81 | +/- | 0. |
| 0.83 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.16 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -0.98 |
| -9999.00 |

BRIGHT_NEIGHBOR
STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 4609. | +/- | 18. |
| 4721. | +/- | 124. |
| 2.50 | +/- | 0. |
| 2.39 | +/- | 0. |
| 0.64 | +/- | 0. |
| 0.64 | +/- | -NaN |
| -0.86 | +/- | 0. |
| -0.86 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.16 | +/- | 0. |
| 4.90 | +/- | 0. |
| 0.69 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.41 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
010-07-C
| 4501. | +/- | 18. |
| -10000. | +/- | -NaN |
| 2.66 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.99 | +/- | 0. |
| 0.99 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.25 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.76 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4351. | +/- | 12. |
| 4458. | +/- | 114. |
| 2.26 | +/- | 0. |
| 2.08 | +/- | 0. |
| 1.27 | +/- | 0. |
| 1.27 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -0.44 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.19 | +/- | 0. |
| 3.83 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -1.27 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
010-07-C
| 4871. | +/- | 10. |
| 4905. | +/- | 107. |
| 3.40 | +/- | 0. |
| 3.20 | +/- | 0. |
| 1.12 | +/- | 0. |
| 1.12 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.40 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.52 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.23 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4117. | +/- | 4. |
| 4241. | +/- | 104. |
| 1.61 | +/- | 0. |
| 1.51 | +/- | 0. |
| 1.71 | +/- | 0. |
| 1.71 | +/- | -NaN |
| -0.83 | +/- | 0. |
| -0.83 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.21 | +/- | 0. |
| 4.81 | +/- | 0. |
| 0.68 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -0.88 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
010-07-C
| 8265. | +/- | 45. |
| 8156. | +/- | 358. |
| 4.32 | +/- | 0. |
| 4.79 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 40.35 | +/- | 0. |
| 1.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.85 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_WARN,COLORTE_WARN
010-07-C
| 7700. | +/- | 14. |
| 7590. | +/- | 261. |
| 3.58 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.23 | +/- | 1. |
| 1.23 | +/- | -NaN |
| -2.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.12 | +/- | 2. |
| -0.12 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 85.98 | +/- | 0. |
| 1.93 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.38 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| -1.98 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -1.99 |
| -9999.00 |
STAR_WARN,SN_WARN
010-07-C
| 4443. | +/- | 14. |
| 4560. | +/- | 121. |
| 2.12 | +/- | 0. |
| 1.97 | +/- | 0. |
| 1.52 | +/- | 0. |
| 1.52 | +/- | -NaN |
| -0.68 | +/- | 0. |
| -0.68 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.37 | +/- | 0. |
| 0.34 | +/- | 0. |
| 4.40 | +/- | 0. |
| 0.64 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.52 |
| -9999.00 |

010-07-C
| 4670. | +/- | 9. |
| 4739. | +/- | 107. |
| 3.34 | +/- | 0. |
| 3.17 | +/- | 0. |
| 0.71 | +/- | 0. |
| 0.71 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.11 | +/- | 0. |
| 2.97 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.61 |
| -9999.00 |

010-07-C
| 4799. | +/- | 5. |
| 4844. | +/- | 87. |
| 2.83 | +/- | 0. |
| 2.59 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.43 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | 0. |
| -0.38 | +/- | 0. |
| -0.38 | +/- | -NaN |
| 0.60 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.61 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.39 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.18 |
| -9999.00 |

010-07-C
| 4131. | +/- | 3. |
| 4225. | +/- | 94. |
| 1.90 | +/- | 0. |
| 1.69 | +/- | 0. |
| 1.48 | +/- | 0. |
| 1.48 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.13 | +/- | 0. |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.17 | +/- | 0. |
| 3.19 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.35 |
| -9999.00 |
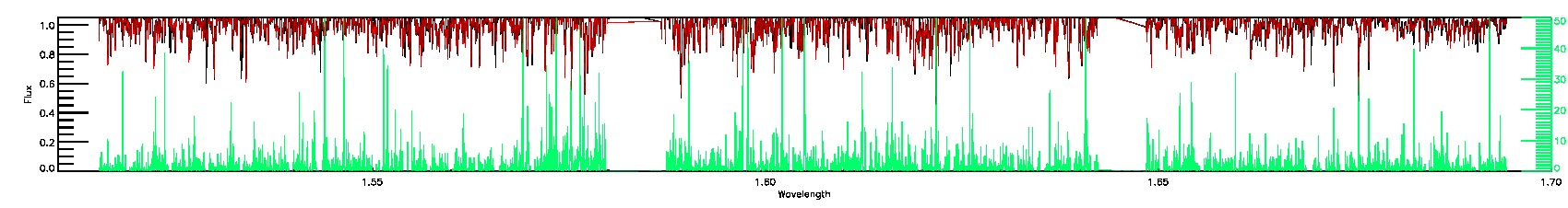
STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 4688. | +/- | 17. |
| 4762. | +/- | 118. |
| 2.71 | +/- | 0. |
| 2.39 | +/- | 0. |
| 1.27 | +/- | 0. |
| 1.27 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.17 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.27 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.00 |
| -9999.00 |

STAR_BAD,COLORTE_BAD,SN_BAD
STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 5113. | +/- | 61. |
| -10000. | +/- | -NaN |
| 2.69 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.92 | +/- | 0. |
| 1.92 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.26 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.31 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.95 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.29 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -1.01 |
| -9999.00 |
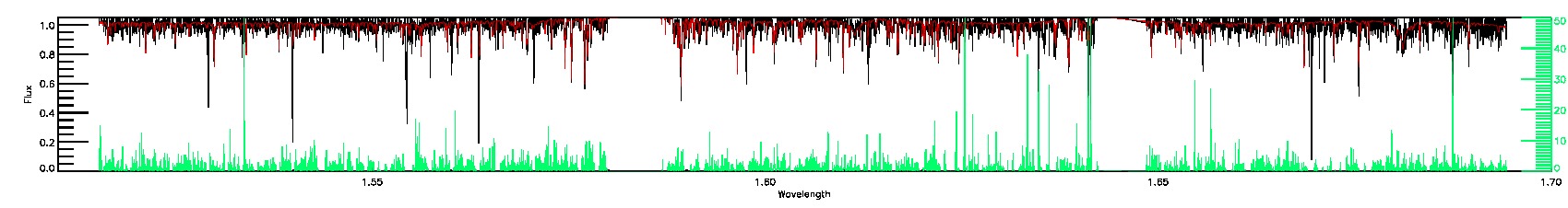
010-07-C
| 3757. | +/- | 2. |
| 3841. | +/- | 81. |
| 1.49 | +/- | 0. |
| 1.28 | +/- | 0. |
| 1.57 | +/- | 0. |
| 1.57 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.20 | +/- | 0. |
| 2.78 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 1.00 |
| -9999.00 |
STAR_WARN,SN_WARN
010-07-C
| 4657. | +/- | 16. |
| 4738. | +/- | 119. |
| 2.74 | +/- | 0. |
| 2.41 | +/- | 0. |
| 1.22 | +/- | 0. |
| 1.22 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -0.26 | +/- | 0. |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.18 | +/- | 0. |
| 3.45 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -0.57 |
| -9999.00 |

010-07-C
| 4470. | +/- | 5. |
| 4542. | +/- | 96. |
| 2.76 | +/- | 0. |
| 2.43 | +/- | 0. |
| 1.04 | +/- | 0. |
| 1.04 | +/- | -NaN |
| 0.38 | +/- | 0. |
| 0.38 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.36 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.80 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4796. | +/- | 24. |
| 4901. | +/- | 135. |
| 2.00 | +/- | 0. |
| 1.93 | +/- | 0. |
| 1.88 | +/- | 0. |
| 1.88 | +/- | -NaN |
| -1.21 | +/- | 0. |
| -1.20 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.37 | +/- | 0. |
| 0.33 | +/- | 0. |
| 5.98 | +/- | 0. |
| 0.78 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -2.47 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
010-07-C
| 6069. | +/- | 126. |
| -10000. | +/- | -NaN |
| 4.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.79 | +/- | 0. |
| 0.79 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 53.93 | +/- | 0. |
| 1.73 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.21 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -0.06 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
010-07-C
| 5962. | +/- | 118. |
| -10000. | +/- | -NaN |
| 3.90 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.18 | +/- | 0. |
| 3.18 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.82 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.96 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| -0.22 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 4193. | +/- | 10. |
| -10000. | +/- | -NaN |
| 1.69 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.64 | +/- | 0. |
| 1.64 | +/- | -NaN |
| -0.72 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.50 | +/- | 0. |
| 0.65 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.15 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
010-07-C
| 4258. | +/- | 20. |
| -10000. | +/- | -NaN |
| 2.52 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.67 | +/- | 0. |
| 0.67 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.04 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.77 |
| -9999.00 |

010-07-C
| 3879. | +/- | 2. |
| 3991. | +/- | 91. |
| 1.33 | +/- | 0. |
| 1.25 | +/- | 0. |
| 1.85 | +/- | 0. |
| 1.85 | +/- | -NaN |
| -0.57 | +/- | 0. |
| -0.57 | +/- | 0. |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.23 | +/- | 0. |
| 4.12 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.33 |
| -9999.00 |
STAR_WARN,SN_WARN
010-07-C
| 3868. | +/- | 4. |
| 3947. | +/- | 90. |
| 1.59 | +/- | 0. |
| 1.36 | +/- | 0. |
| 1.42 | +/- | 0. |
| 1.42 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.05 | +/- | 0. |
| 2.60 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.49 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_WARN,COLORTE_WARN
010-07-C
| 4392. | +/- | 14. |
| 4578. | +/- | 127. |
| 0.86 | +/- | 0. |
| 1.01 | +/- | 0. |
| 2.65 | +/- | 0. |
| 2.65 | +/- | -NaN |
| -2.28 | +/- | 0. |
| -2.28 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.15 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.44 | +/- | 0. |
| 0.41 | +/- | 0. |
| 11.20 | +/- | 0. |
| 1.05 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.31 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -1.14 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4258. | +/- | 9. |
| 4358. | +/- | 109. |
| 2.30 | +/- | 0. |
| 2.10 | +/- | 0. |
| 1.29 | +/- | 0. |
| 1.29 | +/- | -NaN |
| -0.27 | +/- | 0. |
| -0.27 | +/- | 0. |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.21 | +/- | 0. |
| 3.47 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -0.99 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 3771. | +/- | 5. |
| 3847. | +/- | 87. |
| 1.57 | +/- | 0. |
| 1.33 | +/- | 0. |
| 1.40 | +/- | 0. |
| 1.40 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.04 | +/- | 0. |
| 2.51 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.31 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
010-07-C
| 4471. | +/- | 17. |
| -10000. | +/- | -NaN |
| 2.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.24 | +/- | 0. |
| 1.24 | +/- | -NaN |
| -0.62 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.24 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -1.37 |
| -9999.00 |

010-07-C
| 3848. | +/- | 2. |
| 3918. | +/- | 79. |
| 1.80 | +/- | 0. |
| 1.52 | +/- | 0. |
| 1.39 | +/- | 0. |
| 1.39 | +/- | -NaN |
| 0.43 | +/- | 0. |
| 0.43 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.30 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.46 |
| -9999.00 |
010-07-C
| 3736. | +/- | 2. |
| 3808. | +/- | 79. |
| 1.55 | +/- | 0. |
| 1.30 | +/- | 0. |
| 1.42 | +/- | 0. |
| 1.42 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.35 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.43 |
| -9999.00 |
STAR_WARN,SN_WARN
010-07-C
| 4393. | +/- | 14. |
| 4508. | +/- | 118. |
| 2.19 | +/- | 0. |
| 2.04 | +/- | 0. |
| 1.50 | +/- | 0. |
| 1.50 | +/- | -NaN |
| -0.63 | +/- | 0. |
| -0.63 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.22 | +/- | 0. |
| 4.28 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -0.32 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4951. | +/- | 70. |
| 5061. | +/- | 157. |
| 2.47 | +/- | 0. |
| 2.47 | +/- | 0. |
| 0.53 | +/- | 0. |
| 0.53 | +/- | -NaN |
| -1.73 | +/- | 0. |
| -1.73 | +/- | 0. |
| 0.29 | +/- | 0. |
| 0.29 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.26 | +/- | 0. |
| 8.15 | +/- | 0. |
| 0.91 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.25 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.69 |
| -9999.00 |
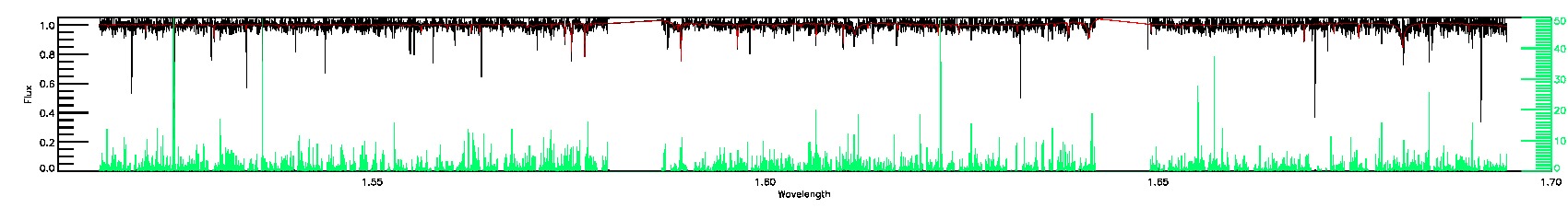
STAR_WARN,SN_WARN
010-07-C
| 4182. | +/- | 7. |
| 4255. | +/- | 99. |
| 2.22 | +/- | 0. |
| 1.93 | +/- | 0. |
| 1.52 | +/- | 0. |
| 1.52 | +/- | -NaN |
| 0.36 | +/- | 0. |
| 0.36 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.40 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.64 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4869. | +/- | 27. |
| 4935. | +/- | 129. |
| 3.09 | +/- | 0. |
| 2.96 | +/- | 0. |
| 1.24 | +/- | 0. |
| 1.24 | +/- | -NaN |
| -0.47 | +/- | 0. |
| -0.47 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.17 | +/- | 0. |
| 3.88 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| 0.48 |
| -9999.00 |

010-07-C
| 4688. | +/- | 11. |
| 4761. | +/- | 108. |
| 2.80 | +/- | 0. |
| 2.61 | +/- | 0. |
| 1.16 | +/- | 0. |
| 1.16 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.23 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.70 |
| -9999.00 |

010-07-C
| 3904. | +/- | 2. |
| 3988. | +/- | 85. |
| 1.58 | +/- | 0. |
| 1.36 | +/- | 0. |
| 1.68 | +/- | 0. |
| 1.68 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| 0.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.17 | +/- | 0. |
| 2.77 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.36 |
| -9999.00 |
STAR_WARN,SN_WARN
010-07-C
| 4600. | +/- | 11. |
| 4674. | +/- | 109. |
| 2.70 | +/- | 0. |
| 2.39 | +/- | 0. |
| 1.23 | +/- | 0. |
| 1.23 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.85 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.24 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4803. | +/- | 14. |
| 4843. | +/- | 111. |
| 3.43 | +/- | 0. |
| 3.22 | +/- | 0. |
| 0.89 | +/- | 0. |
| 0.89 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.47 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| 0.41 |
| -9999.00 |

010-07-C
| 4888. | +/- | 11. |
| 4946. | +/- | 113. |
| 3.02 | +/- | 0. |
| 2.87 | +/- | 0. |
| 1.28 | +/- | 0. |
| 1.28 | +/- | -NaN |
| -0.34 | +/- | 0. |
| -0.34 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.06 | +/- | 0. |
| 3.60 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.45 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4816. | +/- | 47. |
| 4934. | +/- | 147. |
| 2.26 | +/- | 0. |
| 2.23 | +/- | 0. |
| 0.64 | +/- | 0. |
| 0.64 | +/- | -NaN |
| -1.56 | +/- | 0. |
| -1.56 | +/- | 0. |
| 0.41 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.62 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.20 | +/- | 0. |
| 7.35 | +/- | 0. |
| 0.87 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.41 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -1.40 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 3943. | +/- | 5. |
| 4048. | +/- | 97. |
| 1.58 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.57 | +/- | 0. |
| 1.57 | +/- | -NaN |
| -0.38 | +/- | 0. |
| -0.38 | +/- | 0. |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.24 | +/- | 0. |
| 3.70 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -1.10 |
| -9999.00 |
STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 4851. | +/- | 66. |
| 4966. | +/- | 150. |
| 2.33 | +/- | 0. |
| 2.31 | +/- | 0. |
| 0.69 | +/- | 0. |
| 0.69 | +/- | -NaN |
| -1.57 | +/- | 0. |
| -1.57 | +/- | 0. |
| -0.20 | +/- | 0. |
| -0.20 | +/- | -NaN |
| 0.89 | +/- | 0. |
| 0.89 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.10 | +/- | 0. |
| 7.39 | +/- | 0. |
| 0.87 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.31 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -1.93 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -2.41 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4071. | +/- | 5. |
| 4148. | +/- | 94. |
| 2.03 | +/- | 0. |
| 1.76 | +/- | 0. |
| 1.31 | +/- | 0. |
| 1.31 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.51 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.99 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4864. | +/- | 47. |
| 4972. | +/- | 145. |
| 2.37 | +/- | 0. |
| 2.33 | +/- | 0. |
| 1.67 | +/- | 0. |
| 1.67 | +/- | -NaN |
| -1.45 | +/- | 0. |
| -1.45 | +/- | 0. |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.72 | +/- | 0. |
| 0.72 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.10 | +/- | 0. |
| 6.92 | +/- | 0. |
| 0.84 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.29 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -1.98 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -2.43 |
| -9999.00 |

010-07-C
| 4054. | +/- | 2. |
| 4121. | +/- | 83. |
| 2.16 | +/- | 0. |
| 1.86 | +/- | 0. |
| 1.40 | +/- | 0. |
| 1.40 | +/- | -NaN |
| 0.51 | +/- | 0. |
| 0.51 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.43 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.19 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.95 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
010-07-C
| 3964. | +/- | 2. |
| 4076. | +/- | 92. |
| 1.39 | +/- | 0. |
| 1.29 | +/- | 0. |
| 1.85 | +/- | 0. |
| 1.85 | +/- | -NaN |
| -0.56 | +/- | 0. |
| -0.56 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.22 | +/- | 0. |
| 4.09 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.49 |
| -9999.00 |
STAR_WARN,SN_WARN
010-07-C
| 4081. | +/- | 3. |
| 4199. | +/- | 101. |
| 1.42 | +/- | 0. |
| 1.34 | +/- | 0. |
| 1.75 | +/- | 0. |
| 1.75 | +/- | -NaN |
| -0.69 | +/- | 0. |
| -0.69 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.28 | +/- | 0. |
| 4.44 | +/- | 0. |
| 0.65 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -1.54 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4153. | +/- | 4. |
| 4268. | +/- | 104. |
| 1.78 | +/- | 0. |
| 1.64 | +/- | 0. |
| 1.47 | +/- | 0. |
| 1.47 | +/- | -NaN |
| -0.62 | +/- | 0. |
| -0.62 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.22 | +/- | 0. |
| 4.25 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -1.22 |
| -9999.00 |

010-07-C
| 3946. | +/- | 2. |
| 4071. | +/- | 96. |
| 1.24 | +/- | 0. |
| 1.20 | +/- | 0. |
| 2.03 | +/- | 0. |
| 2.03 | +/- | -NaN |
| -0.86 | +/- | 0. |
| -0.86 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.24 | +/- | 0. |
| 4.88 | +/- | 0. |
| 0.69 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.94 |
| -9999.00 |
STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 4430. | +/- | 20. |
| 4584. | +/- | 128. |
| 1.52 | +/- | 0. |
| 1.53 | +/- | 0. |
| 1.95 | +/- | 0. |
| 1.95 | +/- | -NaN |
| -1.54 | +/- | 0. |
| -1.54 | +/- | 0. |
| -0.18 | +/- | 0. |
| -0.18 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.20 | +/- | 0. |
| 7.27 | +/- | 0. |
| 0.86 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.19 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.83 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4768. | +/- | 53. |
| 4896. | +/- | 147. |
| 2.37 | +/- | 0. |
| 2.36 | +/- | 0. |
| 0.49 | +/- | 0. |
| 0.49 | +/- | -NaN |
| -1.66 | +/- | 0. |
| -1.66 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.85 | +/- | 0. |
| 0.85 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.21 | +/- | 0. |
| 7.80 | +/- | 0. |
| 0.89 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.33 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.54 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 6035. | +/- | 139. |
| -10000. | +/- | -NaN |
| 3.62 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.29 | +/- | 0. |
| 2.29 | +/- | -NaN |
| -0.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.24 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 26.83 | +/- | 0. |
| 1.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.26 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| 1.00 |
| -9999.00 |
STAR_WARN,SN_WARN
010-07-C
| 4025. | +/- | 6. |
| 4096. | +/- | 93. |
| 2.11 | +/- | 0. |
| 1.82 | +/- | 0. |
| 1.37 | +/- | 0. |
| 1.37 | +/- | -NaN |
| 0.40 | +/- | 0. |
| 0.40 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.34 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.05 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 3917. | +/- | 4. |
| 3994. | +/- | 88. |
| 1.71 | +/- | 0. |
| 1.46 | +/- | 0. |
| 1.49 | +/- | 0. |
| 1.49 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.52 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.27 |
| -9999.00 |
010-07-C
| 8293. | +/- | 43. |
| 8167. | +/- | 353. |
| 4.37 | +/- | 0. |
| 4.68 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -1.77 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 30.05 | +/- | 0. |
| 1.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.00 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -1.98 |
| -9999.00 |
| -1.93 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -1.98 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -1.75 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
010-07-C
| 5064. | +/- | 24. |
| 5168. | +/- | 147. |
| 2.09 | +/- | 0. |
| 2.10 | +/- | 0. |
| 2.08 | +/- | 0. |
| 2.08 | +/- | -NaN |
| -1.89 | +/- | 0. |
| -1.89 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.36 | +/- | 0. |
| 0.33 | +/- | 0. |
| 8.94 | +/- | 0. |
| 0.95 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.27 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| -0.93 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4612. | +/- | 12. |
| 4681. | +/- | 110. |
| 2.67 | +/- | 0. |
| 2.37 | +/- | 0. |
| 1.46 | +/- | 0. |
| 1.46 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.71 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.96 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
010-07-C
| 4515. | +/- | 21. |
| -10000. | +/- | -NaN |
| 4.66 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.43 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.70 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -2.40 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4465. | +/- | 6. |
| 4541. | +/- | 101. |
| 2.68 | +/- | 0. |
| 2.37 | +/- | 0. |
| 1.23 | +/- | 0. |
| 1.23 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.28 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.06 | +/- | 0. |
| 2.52 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.94 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4152. | +/- | 4. |
| 4260. | +/- | 103. |
| 1.90 | +/- | 0. |
| 1.73 | +/- | 0. |
| 1.71 | +/- | 0. |
| 1.71 | +/- | -NaN |
| -0.45 | +/- | 0. |
| -0.45 | +/- | 0. |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.23 | +/- | 0. |
| 3.84 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.08 |
| -9999.00 |
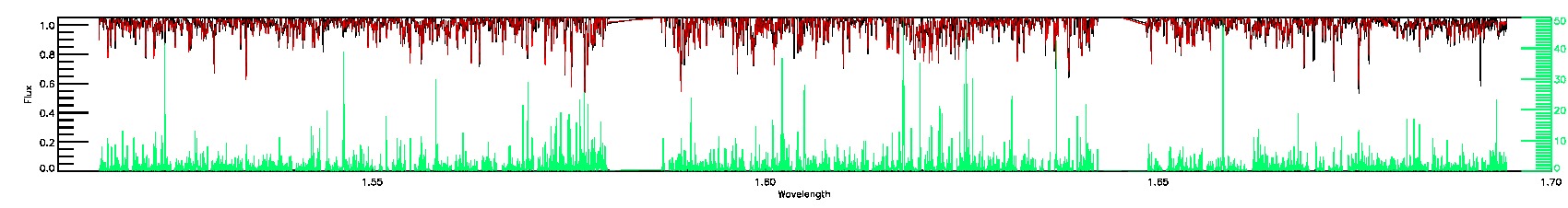
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
010-07-C
| 5086. | +/- | 64. |
| -10000. | +/- | -NaN |
| 3.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.60 | +/- | 0. |
| 1.60 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.11 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 1.00 |
| -9999.00 |

010-07-C
| 4829. | +/- | 27. |
| 4951. | +/- | 138. |
| 2.23 | +/- | 0. |
| 2.22 | +/- | 0. |
| 0.61 | +/- | 0. |
| 0.61 | +/- | -NaN |
| -1.69 | +/- | 0. |
| -1.68 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.28 | +/- | 0. |
| 7.93 | +/- | 0. |
| 0.90 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.14 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| -2.36 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4120. | +/- | 5. |
| 4216. | +/- | 102. |
| 2.01 | +/- | 0. |
| 1.80 | +/- | 0. |
| 1.40 | +/- | 0. |
| 1.40 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.17 | +/- | 0. |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.20 | +/- | 0. |
| 3.27 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.54 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4916. | +/- | 51. |
| 5021. | +/- | 150. |
| 2.39 | +/- | 0. |
| 2.37 | +/- | 0. |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| -1.52 | +/- | 0. |
| -1.52 | +/- | 0. |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.90 | +/- | 0. |
| 0.90 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.20 | +/- | 0. |
| 7.19 | +/- | 0. |
| 0.86 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.40 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.05 |
| -9999.00 |

STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 5245. | +/- | 40. |
| 5288. | +/- | 151. |
| 2.78 | +/- | 0. |
| 2.44 | +/- | 0. |
| 2.53 | +/- | 0. |
| 2.53 | +/- | -NaN |
| -0.95 | +/- | 0. |
| -0.95 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.26 | +/- | 0. |
| 5.17 | +/- | 0. |
| 0.71 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.30 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4343. | +/- | 11. |
| 4454. | +/- | 115. |
| 2.18 | +/- | 0. |
| 2.01 | +/- | 0. |
| 1.35 | +/- | 0. |
| 1.35 | +/- | -NaN |
| -0.51 | +/- | 0. |
| -0.51 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.24 | +/- | 0. |
| 4.00 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.51 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 4064. | +/- | 8. |
| -10000. | +/- | -NaN |
| 2.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.46 | +/- | 0. |
| 1.46 | +/- | -NaN |
| 0.52 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.35 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.19 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.93 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4852. | +/- | 40. |
| 4970. | +/- | 146. |
| 2.25 | +/- | 0. |
| 2.24 | +/- | 0. |
| 0.75 | +/- | 0. |
| 0.75 | +/- | -NaN |
| -1.66 | +/- | 0. |
| -1.66 | +/- | 0. |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.16 | +/- | 0. |
| 7.81 | +/- | 0. |
| 0.89 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.21 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -1.99 |
| -9999.00 |

STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 4779. | +/- | 37. |
| 4904. | +/- | 142. |
| 2.29 | +/- | 0. |
| 2.27 | +/- | 0. |
| 1.32 | +/- | 0. |
| 1.32 | +/- | -NaN |
| -1.62 | +/- | 0. |
| -1.62 | +/- | 0. |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.74 | +/- | 0. |
| 0.74 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.18 | +/- | 0. |
| 7.63 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.35 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.99 |
| -9999.00 |

STAR_BAD
STAR_WARN,COLORTE_WARN
010-07-C
| 8879. | +/- | 88. |
| -10000. | +/- | -NaN |
| 4.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.57 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.39 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.95 |
| -9999.00 |
010-07-C
| 4591. | +/- | 7. |
| 4733. | +/- | 105. |
| 1.86 | +/- | 0. |
| 1.83 | +/- | 0. |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| -1.49 | +/- | 0. |
| -1.49 | +/- | 0. |
| -0.16 | +/- | 0. |
| -0.16 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.24 | +/- | 0. |
| 7.06 | +/- | 0. |
| 0.85 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.36 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.59 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4881. | +/- | 36. |
| 4991. | +/- | 143. |
| 2.47 | +/- | 0. |
| 2.45 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.43 | +/- | -NaN |
| -1.54 | +/- | 0. |
| -1.54 | +/- | 0. |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 1.04 | +/- | 0. |
| 1.04 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.26 | +/- | 0. |
| 7.26 | +/- | 0. |
| 0.86 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.34 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 1.03 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.07 |
| -9999.00 |

010-07-C
| 4770. | +/- | 26. |
| 4898. | +/- | 137. |
| 2.18 | +/- | 0. |
| 2.16 | +/- | 0. |
| 0.68 | +/- | 0. |
| 0.68 | +/- | -NaN |
| -1.67 | +/- | 0. |
| -1.67 | +/- | 0. |
| 0.37 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.27 | +/- | 0. |
| 7.84 | +/- | 0. |
| 0.89 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.28 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.69 |
| -9999.00 |
| -1.98 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.82 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4711. | +/- | 37. |
| 4811. | +/- | 134. |
| 2.87 | +/- | 0. |
| 2.78 | +/- | 0. |
| 0.48 | +/- | 0. |
| 0.48 | +/- | -NaN |
| -0.85 | +/- | 0. |
| -0.85 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.26 | +/- | 0. |
| 4.87 | +/- | 0. |
| 0.69 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.22 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4966. | +/- | 54. |
| 5067. | +/- | 151. |
| 2.47 | +/- | 0. |
| 2.45 | +/- | 0. |
| 1.06 | +/- | 0. |
| 1.06 | +/- | -NaN |
| -1.54 | +/- | 0. |
| -1.54 | +/- | 0. |
| -0.60 | +/- | 0. |
| -0.60 | +/- | -NaN |
| 1.16 | +/- | 0. |
| 1.16 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.20 | +/- | 0. |
| 7.30 | +/- | 0. |
| 0.86 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.83 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| 1.05 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.50 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 6380. | +/- | 31. |
| -10000. | +/- | -NaN |
| 2.53 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.09 | +/- | 0. |
| 4.09 | +/- | -NaN |
| -1.87 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 7. |
| -9999.99 | +/- | -NaN |
| 0.46 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.45 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -1.93 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.39 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
010-07-C
| 12240. | +/- | 149. |
| -10000. | +/- | -NaN |
| 4.37 | +/- | 0. |
| 4.34 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -0.96 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 15.42 | +/- | 0. |
| 1.19 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.89 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -0.96 |
| -9999.00 |
STAR_WARN,SN_WARN
010-07-C
| 4783. | +/- | 45. |
| 4916. | +/- | 147. |
| 2.28 | +/- | 0. |
| 2.29 | +/- | 0. |
| 1.21 | +/- | 0. |
| 1.21 | +/- | -NaN |
| -1.82 | +/- | 0. |
| -1.82 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.25 | +/- | 0. |
| 8.59 | +/- | 0. |
| 0.93 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.35 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -1.93 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.85 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4808. | +/- | 54. |
| 4941. | +/- | 152. |
| 2.19 | +/- | 0. |
| 2.21 | +/- | 0. |
| 0.82 | +/- | 0. |
| 0.82 | +/- | -NaN |
| -1.87 | +/- | 0. |
| -1.87 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.27 | +/- | 0. |
| 8.82 | +/- | 0. |
| 0.95 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.41 |
| -9999.00 |
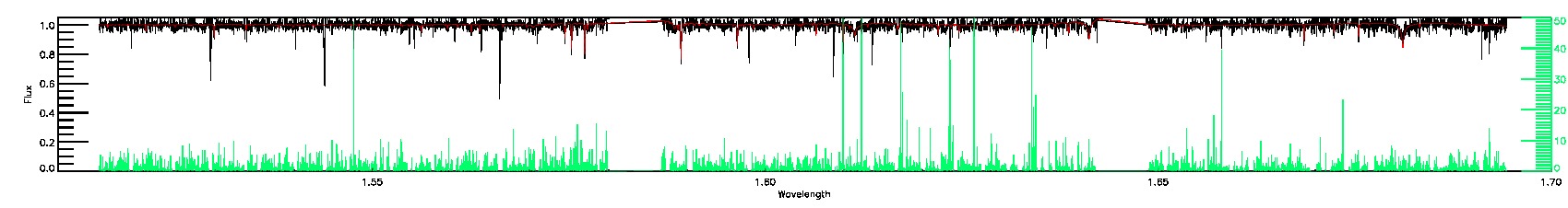
010-07-C
| 4773. | +/- | 26. |
| 4898. | +/- | 135. |
| 2.06 | +/- | 0. |
| 2.04 | +/- | 0. |
| 1.86 | +/- | 0. |
| 1.86 | +/- | -NaN |
| -1.60 | +/- | 0. |
| -1.60 | +/- | 0. |
| -0.18 | +/- | 0. |
| -0.18 | +/- | -NaN |
| 0.77 | +/- | 0. |
| 0.77 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.13 | +/- | 0. |
| 7.54 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.23 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -1.69 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -2.01 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4799. | +/- | 28. |
| 4916. | +/- | 136. |
| 2.27 | +/- | 0. |
| 2.23 | +/- | 0. |
| 1.64 | +/- | 0. |
| 1.64 | +/- | -NaN |
| -1.47 | +/- | 0. |
| -1.47 | +/- | 0. |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.80 | +/- | 0. |
| 0.80 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.13 | +/- | 0. |
| 6.98 | +/- | 0. |
| 0.84 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.20 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -2.14 |
| -9999.00 |
| -2.24 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4878. | +/- | 39. |
| 4993. | +/- | 146. |
| 2.25 | +/- | 0. |
| 2.23 | +/- | 0. |
| 0.61 | +/- | 0. |
| 0.61 | +/- | -NaN |
| -1.66 | +/- | 0. |
| -1.65 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.53 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.16 | +/- | 0. |
| 7.79 | +/- | 0. |
| 0.89 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.44 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4903. | +/- | 64. |
| 5023. | +/- | 156. |
| 2.33 | +/- | 0. |
| 2.34 | +/- | 0. |
| 2.22 | +/- | 0. |
| 2.22 | +/- | -NaN |
| -1.84 | +/- | 0. |
| -1.84 | +/- | 0. |
| -0.72 | +/- | 1. |
| -0.72 | +/- | -NaN |
| 0.16 | +/- | 3. |
| 0.16 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.27 | +/- | 0. |
| 8.69 | +/- | 0. |
| 0.94 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.79 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -1.69 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.84 |
| -9999.00 |

010-07-C
| 3757. | +/- | 2. |
| 3864. | +/- | 88. |
| 1.22 | +/- | 0. |
| 1.13 | +/- | 0. |
| 1.87 | +/- | 0. |
| 1.87 | +/- | -NaN |
| -0.43 | +/- | 0. |
| -0.43 | +/- | 0. |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.27 | +/- | 0. |
| 3.79 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.56 |
| -9999.00 |
010-07-C
| 4203. | +/- | 4. |
| 4303. | +/- | 96. |
| 1.68 | +/- | 0. |
| 1.50 | +/- | 0. |
| 1.47 | +/- | 0. |
| 1.47 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -0.28 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.54 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 3.49 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.47 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4836. | +/- | 47. |
| 4953. | +/- | 148. |
| 2.36 | +/- | 0. |
| 2.34 | +/- | 0. |
| 0.83 | +/- | 0. |
| 0.83 | +/- | -NaN |
| -1.58 | +/- | 0. |
| -1.58 | +/- | 0. |
| 0.64 | +/- | 0. |
| 0.64 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.41 | +/- | 0. |
| 0.37 | +/- | 0. |
| 7.47 | +/- | 0. |
| 0.87 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.77 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.02 |
| -9999.00 |

010-07-C
| 4832. | +/- | 25. |
| 4949. | +/- | 135. |
| 2.17 | +/- | 0. |
| 2.15 | +/- | 0. |
| 1.48 | +/- | 0. |
| 1.48 | +/- | -NaN |
| -1.58 | +/- | 0. |
| -1.58 | +/- | 0. |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.20 | +/- | 0. |
| 7.45 | +/- | 0. |
| 0.87 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.20 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.34 |
| -9999.00 |

010-07-C
| 4212. | +/- | 3. |
| 4370. | +/- | 87. |
| 1.22 | +/- | 0. |
| 1.29 | +/- | 0. |
| 1.93 | +/- | 0. |
| 1.93 | +/- | -NaN |
| -1.62 | +/- | 0. |
| -1.62 | +/- | 0. |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.96 | +/- | 0. |
| 0.96 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.22 | +/- | 0. |
| 7.65 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.25 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -1.09 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4890. | +/- | 37. |
| 5001. | +/- | 144. |
| 2.33 | +/- | 0. |
| 2.31 | +/- | 0. |
| 1.14 | +/- | 0. |
| 1.14 | +/- | -NaN |
| -1.60 | +/- | 0. |
| -1.60 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.73 | +/- | 0. |
| 0.73 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.23 | +/- | 0. |
| 7.53 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.17 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.25 |
| -9999.00 |

010-07-C
| 4805. | +/- | 22. |
| 4942. | +/- | 138. |
| 1.71 | +/- | 0. |
| 1.75 | +/- | 0. |
| 1.90 | +/- | 0. |
| 1.90 | +/- | -NaN |
| -1.97 | +/- | 0. |
| -1.97 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.24 | +/- | 0. |
| 9.35 | +/- | 0. |
| 0.97 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.53 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4771. | +/- | 30. |
| 4903. | +/- | 141. |
| 2.18 | +/- | 0. |
| 2.18 | +/- | 0. |
| 0.96 | +/- | 0. |
| 0.96 | +/- | -NaN |
| -1.77 | +/- | 0. |
| -1.77 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -0.29 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.30 | +/- | 0. |
| 8.31 | +/- | 0. |
| 0.92 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -1.93 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.24 |
| -9999.00 |
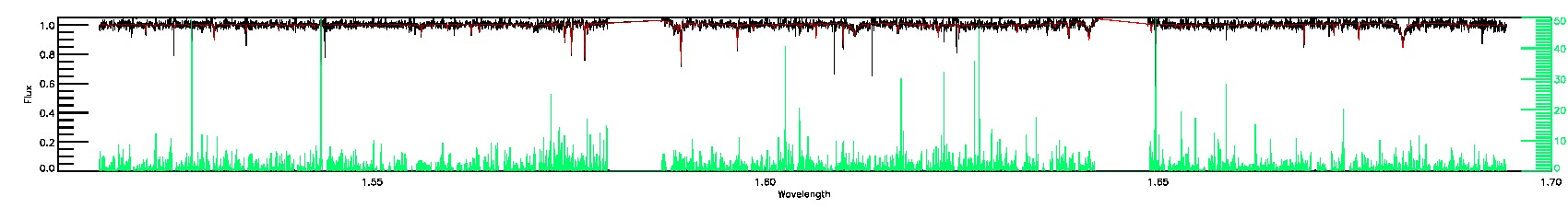
STAR_WARN,SN_WARN
010-07-C
| 4075. | +/- | 3. |
| 4150. | +/- | 91. |
| 2.00 | +/- | 0. |
| 1.72 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.43 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.48 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.84 |
| -9999.00 |

010-07-C
| 4523. | +/- | 7. |
| 4677. | +/- | 105. |
| 1.70 | +/- | 0. |
| 1.70 | +/- | 0. |
| 1.55 | +/- | 0. |
| 1.55 | +/- | -NaN |
| -1.59 | +/- | 0. |
| -1.59 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.87 | +/- | 0. |
| 0.87 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.26 | +/- | 0. |
| 7.51 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -1.47 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_WARN,COLORTE_WARN
010-07-C
| 8910. | +/- | 53. |
| 8794. | +/- | 380. |
| 4.01 | +/- | 0. |
| 4.41 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.57 | +/- | 0. |
| 0.75 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.02 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.17 |
| -9999.00 |
010-07-C
| 4328. | +/- | 3. |
| 4486. | +/- | 90. |
| 1.32 | +/- | 0. |
| 1.38 | +/- | 0. |
| 0.74 | +/- | 0. |
| 0.74 | +/- | -NaN |
| -1.64 | +/- | 0. |
| -1.63 | +/- | 0. |
| -0.45 | +/- | 0. |
| -0.45 | +/- | -NaN |
| -0.45 | +/- | 0. |
| -0.45 | +/- | -NaN |
| 0.43 | +/- | 0. |
| 0.39 | +/- | 0. |
| 7.70 | +/- | 0. |
| 0.89 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.57 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.97 |
| -9999.00 |

010-07-C
| 4690. | +/- | 21. |
| 4828. | +/- | 131. |
| 2.00 | +/- | 0. |
| 1.99 | +/- | 0. |
| 1.36 | +/- | 0. |
| 1.36 | +/- | -NaN |
| -1.68 | +/- | 0. |
| -1.68 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.25 | +/- | 0. |
| 7.88 | +/- | 0. |
| 0.90 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.49 |
| -9999.00 |

STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN
010-07-C
| 4798. | +/- | 16. |
| -10000. | +/- | -NaN |
| 1.67 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.17 | +/- | 0. |
| 2.17 | +/- | -NaN |
| -1.74 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.18 | +/- | 0. |
| 0.91 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.44 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.04 |
| -9999.00 |

STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 4952. | +/- | 55. |
| 5057. | +/- | 152. |
| 2.42 | +/- | 0. |
| 2.40 | +/- | 0. |
| 1.07 | +/- | 0. |
| 1.07 | +/- | -NaN |
| -1.62 | +/- | 0. |
| -1.62 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.17 | +/- | 0. |
| 7.64 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.23 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.19 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4769. | +/- | 40. |
| 4894. | +/- | 143. |
| 2.24 | +/- | 0. |
| 2.21 | +/- | 0. |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -1.58 | +/- | 0. |
| -1.58 | +/- | 0. |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.86 | +/- | 0. |
| 0.86 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.18 | +/- | 0. |
| 7.45 | +/- | 0. |
| 0.87 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -2.36 |
| -9999.00 |
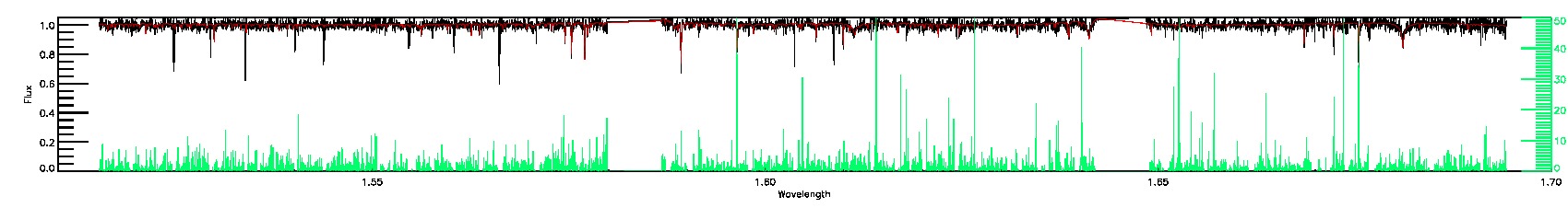
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
010-07-C
| 4647. | +/- | 64. |
| -10000. | +/- | -NaN |
| 1.94 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.52 | +/- | 0. |
| 1.52 | +/- | -NaN |
| -1.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.37 | +/- | 0. |
| 0.92 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.43 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -2.44 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4831. | +/- | 32. |
| 4942. | +/- | 141. |
| 2.48 | +/- | 0. |
| 2.44 | +/- | 0. |
| 0.37 | +/- | 0. |
| 0.37 | +/- | -NaN |
| -1.42 | +/- | 0. |
| -1.42 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.41 | +/- | 0. |
| 0.38 | +/- | 0. |
| 6.78 | +/- | 0. |
| 0.83 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.44 |
| -9999.00 |

010-07-C
| 4856. | +/- | 28. |
| 4980. | +/- | 141. |
| 2.16 | +/- | 0. |
| 2.17 | +/- | 0. |
| 1.75 | +/- | 0. |
| 1.75 | +/- | -NaN |
| -1.79 | +/- | 0. |
| -1.79 | +/- | 0. |
| -0.20 | +/- | 0. |
| -0.20 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -0.44 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.29 | +/- | 0. |
| 8.44 | +/- | 0. |
| 0.93 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.20 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.10 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
010-07-C
| 7316. | +/- | 110. |
| -10000. | +/- | -NaN |
| 3.64 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.59 | +/- | 2. |
| 0.59 | +/- | -NaN |
| -0.84 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 6. |
| -9999.99 | +/- | -NaN |
| -0.34 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 15.19 | +/- | 0. |
| 1.18 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -1.00 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
010-07-C
| 7307. | +/- | 104. |
| -10000. | +/- | -NaN |
| 3.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.02 | +/- | 1. |
| 1.02 | +/- | -NaN |
| -0.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.34 | +/- | 0. |
| 1.01 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -1.29 |
| -9999.00 |
STAR_WARN,SN_WARN
010-07-C
| 4743. | +/- | 23. |
| 4868. | +/- | 134. |
| 2.21 | +/- | 0. |
| 2.17 | +/- | 0. |
| 0.61 | +/- | 0. |
| 0.61 | +/- | -NaN |
| -1.51 | +/- | 0. |
| -1.51 | +/- | 0. |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| 0.45 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.30 | +/- | 0. |
| 7.14 | +/- | 0. |
| 0.85 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.28 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.14 |
| -9999.00 |

SUSPECT_BROAD_LINES
010-07-C
| 8381. | +/- | 47. |
| 8256. | +/- | 366. |
| 4.40 | +/- | 0. |
| 4.72 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -1.80 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 15.11 | +/- | 0. |
| 1.18 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.01 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -1.79 |
| -9999.00 |
STAR_WARN,SN_WARN
010-07-C
| 3977. | +/- | 3. |
| 4056. | +/- | 89. |
| 1.84 | +/- | 0. |
| 1.58 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.43 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.61 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.23 |
| -9999.00 |
STAR_WARN,SN_WARN
010-07-C
| 4904. | +/- | 81. |
| 5033. | +/- | 161. |
| 2.37 | +/- | 0. |
| 2.41 | +/- | 0. |
| 0.60 | +/- | 0. |
| 0.60 | +/- | -NaN |
| -2.04 | +/- | 0. |
| -2.04 | +/- | 0. |
| -0.48 | +/- | 1. |
| -0.48 | +/- | -NaN |
| -0.31 | +/- | 1. |
| -0.31 | +/- | -NaN |
| 0.40 | +/- | 0. |
| 0.37 | +/- | 0. |
| 9.75 | +/- | 0. |
| 0.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.35 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.26 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4845. | +/- | 41. |
| 4965. | +/- | 145. |
| 2.33 | +/- | 0. |
| 2.32 | +/- | 0. |
| 1.00 | +/- | 0. |
| 1.00 | +/- | -NaN |
| -1.67 | +/- | 0. |
| -1.67 | +/- | 0. |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.71 | +/- | 0. |
| 0.71 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.18 | +/- | 0. |
| 7.87 | +/- | 0. |
| 0.90 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.28 |
| -9999.00 |

010-07-C
| 4323. | +/- | 4. |
| 4481. | +/- | 90. |
| 1.27 | +/- | 0. |
| 1.34 | +/- | 0. |
| 1.80 | +/- | 0. |
| 1.80 | +/- | -NaN |
| -1.64 | +/- | 0. |
| -1.64 | +/- | 0. |
| -0.31 | +/- | 0. |
| -0.31 | +/- | -NaN |
| 0.55 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.25 | +/- | 0. |
| 7.70 | +/- | 0. |
| 0.89 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.35 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -2.38 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 5025. | +/- | 43. |
| 5117. | +/- | 150. |
| 2.40 | +/- | 0. |
| 2.37 | +/- | 0. |
| 2.17 | +/- | 0. |
| 2.17 | +/- | -NaN |
| -1.50 | +/- | 0. |
| -1.50 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.97 | +/- | 0. |
| 0.97 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.15 | +/- | 0. |
| 7.13 | +/- | 0. |
| 0.85 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 1.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.78 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4875. | +/- | 51. |
| 4988. | +/- | 149. |
| 2.23 | +/- | 0. |
| 2.21 | +/- | 0. |
| 0.45 | +/- | 0. |
| 0.45 | +/- | -NaN |
| -1.60 | +/- | 0. |
| -1.60 | +/- | 0. |
| -0.42 | +/- | 0. |
| -0.42 | +/- | -NaN |
| 0.52 | +/- | 1. |
| 0.52 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.23 | +/- | 0. |
| 7.53 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.14 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4828. | +/- | 77. |
| 4958. | +/- | 155. |
| 2.35 | +/- | 0. |
| 2.36 | +/- | 0. |
| 1.85 | +/- | 0. |
| 1.85 | +/- | -NaN |
| -1.86 | +/- | 0. |
| -1.86 | +/- | 0. |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.26 | +/- | 0. |
| 8.76 | +/- | 0. |
| 0.94 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.45 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.96 |
| -9999.00 |

STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN
010-07-C
| 4664. | +/- | 11. |
| -10000. | +/- | -NaN |
| 1.57 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.21 | +/- | 0. |
| 2.21 | +/- | -NaN |
| -1.67 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.75 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.85 | +/- | 0. |
| 0.89 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.30 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -1.98 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.19 |
| -9999.00 |

BRIGHT_NEIGHBOR
010-07-C
| 4825. | +/- | 30. |
| 4945. | +/- | 139. |
| 2.20 | +/- | 0. |
| 2.18 | +/- | 0. |
| 1.25 | +/- | 0. |
| 1.25 | +/- | -NaN |
| -1.62 | +/- | 0. |
| -1.62 | +/- | 0. |
| -1.17 | +/- | 0. |
| -1.17 | +/- | -NaN |
| 1.56 | +/- | 0. |
| 1.56 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.09 | +/- | 0. |
| 7.62 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.06 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| 1.63 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.14 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
010-07-C
| 6448. | +/- | 81. |
| -10000. | +/- | -NaN |
| 2.59 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.29 | +/- | 0. |
| 1.29 | +/- | -NaN |
| -0.62 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 70.91 | +/- | 0. |
| 1.85 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.18 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
010-07-C
| 7666. | +/- | 154. |
| -10000. | +/- | -NaN |
| 2.76 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.85 | +/- | 1. |
| 0.85 | +/- | -NaN |
| -0.86 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.31 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 15.14 | +/- | 0. |
| 1.18 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -1.58 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
010-07-C
| 4848. | +/- | 81. |
| -10000. | +/- | -NaN |
| 2.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.07 | +/- | 0. |
| 1.07 | +/- | -NaN |
| -1.91 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 9.06 | +/- | 0. |
| 0.96 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.58 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -2.34 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
010-07-C
| 6990. | +/- | 80. |
| -10000. | +/- | -NaN |
| 3.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.01 | +/- | 1. |
| 1.01 | +/- | -NaN |
| -1.96 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 8. |
| -9999.99 | +/- | -NaN |
| 0.22 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.78 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -2.12 |
| -9999.00 |
STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 4248. | +/- | 7. |
| 4354. | +/- | 109. |
| 2.12 | +/- | 0. |
| 1.94 | +/- | 0. |
| 1.82 | +/- | 0. |
| 1.82 | +/- | -NaN |
| -0.43 | +/- | 0. |
| -0.43 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.40 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.20 | +/- | 0. |
| 3.80 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.19 |
| -9999.00 |

010-07-C
| 4656. | +/- | 9. |
| 4793. | +/- | 101. |
| 1.71 | +/- | 0. |
| 1.70 | +/- | 0. |
| 2.36 | +/- | 0. |
| 2.36 | +/- | -NaN |
| -1.54 | +/- | 0. |
| -1.54 | +/- | 0. |
| -0.26 | +/- | 0. |
| -0.26 | +/- | -NaN |
| 0.41 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.04 | +/- | 0. |
| 7.28 | +/- | 0. |
| 0.86 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.29 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.80 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 3913. | +/- | 3. |
| 4011. | +/- | 93. |
| 1.45 | +/- | 0. |
| 1.30 | +/- | 0. |
| 1.57 | +/- | 0. |
| 1.57 | +/- | -NaN |
| -0.21 | +/- | 0. |
| -0.21 | +/- | 0. |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.23 | +/- | 0. |
| 3.34 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.88 |
| -9999.00 |
010-07-C
| 4669. | +/- | 8. |
| 4808. | +/- | 100. |
| 1.50 | +/- | 0. |
| 1.52 | +/- | 0. |
| 2.42 | +/- | 0. |
| 2.42 | +/- | -NaN |
| -1.63 | +/- | 0. |
| -1.63 | +/- | 0. |
| -0.35 | +/- | 0. |
| -0.35 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.29 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.11 | +/- | 0. |
| 7.67 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.60 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.90 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
010-07-C
| 4556. | +/- | 9. |
| 4713. | +/- | 110. |
| 1.64 | +/- | 0. |
| 1.67 | +/- | 0. |
| 1.65 | +/- | 0. |
| 1.65 | +/- | -NaN |
| -1.76 | +/- | 0. |
| -1.76 | +/- | 0. |
| -0.37 | +/- | 0. |
| -0.37 | +/- | -NaN |
| -0.41 | +/- | 0. |
| -0.41 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.27 | +/- | 0. |
| 8.27 | +/- | 0. |
| 0.92 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.50 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.33 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 5357. | +/- | 53. |
| 5412. | +/- | 168. |
| 2.80 | +/- | 0. |
| 2.79 | +/- | 0. |
| 2.49 | +/- | 0. |
| 2.49 | +/- | -NaN |
| -1.55 | +/- | 0. |
| -1.55 | +/- | 0. |
| -0.36 | +/- | 0. |
| -0.36 | +/- | -NaN |
| 0.04 | +/- | 1. |
| 0.04 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.18 | +/- | 0. |
| 7.30 | +/- | 0. |
| 0.86 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.37 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.94 |
| -9999.00 |

010-07-C
| 4232. | +/- | 3. |
| 4397. | +/- | 90. |
| 0.93 | +/- | 0. |
| 1.02 | +/- | 0. |
| 2.16 | +/- | 0. |
| 2.16 | +/- | -NaN |
| -1.80 | +/- | 0. |
| -1.80 | +/- | 0. |
| -0.55 | +/- | 0. |
| -0.55 | +/- | -NaN |
| 0.49 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.25 | +/- | 0. |
| 8.45 | +/- | 0. |
| 0.93 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.62 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -2.14 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -2.36 |
| -9999.00 |
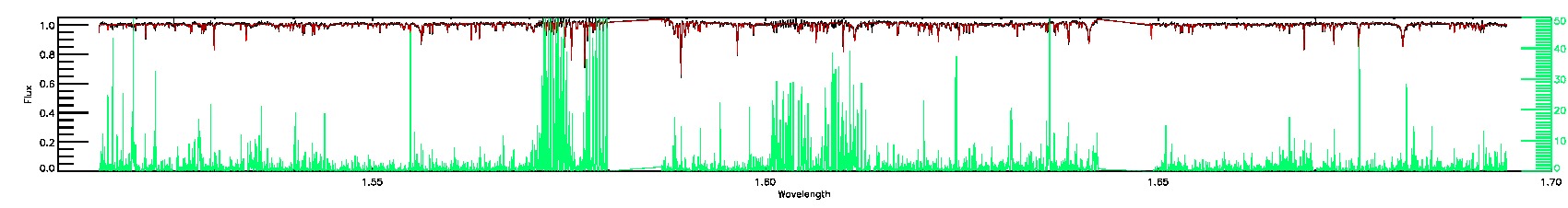
STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 5543. | +/- | 77. |
| 5573. | +/- | 182. |
| 3.05 | +/- | 0. |
| 2.68 | +/- | 0. |
| 2.82 | +/- | 0. |
| 2.82 | +/- | -NaN |
| -1.47 | +/- | 0. |
| -1.46 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.25 | +/- | 0. |
| 2.37 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.31 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
010-07-C
| 5585. | +/- | 98. |
| -10000. | +/- | -NaN |
| 1.95 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.49 | +/- | 2. |
| 4.49 | +/- | -NaN |
| -2.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.12 | +/- | 14. |
| -9999.99 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.60 | +/- | 0. |
| 1.03 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.36 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.20 |
| -9999.00 |
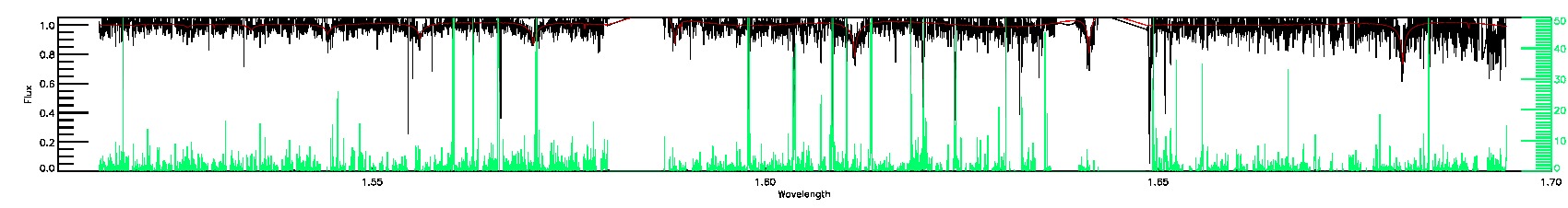
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 4996. | +/- | 42. |
| -10000. | +/- | -NaN |
| 2.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.80 | +/- | 0. |
| 1.80 | +/- | -NaN |
| -1.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.30 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.05 | +/- | 0. |
| 0.85 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.39 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 1.07 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.36 |
| -9999.00 |

010-07-C
| 4729. | +/- | 26. |
| 4863. | +/- | 134. |
| 1.95 | +/- | 0. |
| 1.94 | +/- | 0. |
| 1.86 | +/- | 0. |
| 1.86 | +/- | -NaN |
| -1.68 | +/- | 0. |
| -1.68 | +/- | 0. |
| -0.21 | +/- | 0. |
| -0.21 | +/- | -NaN |
| 0.67 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.07 | +/- | 0. |
| 7.88 | +/- | 0. |
| 0.90 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.17 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -1.69 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -1.64 |
| -9999.00 |

BRIGHT_NEIGHBOR
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
010-07-C
| 4881. | +/- | 88. |
| -10000. | +/- | -NaN |
| 2.46 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.26 | +/- | 0. |
| 2.26 | +/- | -NaN |
| -1.70 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.77 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.01 | +/- | 0. |
| 0.90 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.60 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.59 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4829. | +/- | 41. |
| 4949. | +/- | 145. |
| 2.12 | +/- | 0. |
| 2.10 | +/- | 0. |
| 1.01 | +/- | 0. |
| 1.01 | +/- | -NaN |
| -1.63 | +/- | 0. |
| -1.63 | +/- | 0. |
| -0.13 | +/- | 0. |
| -0.13 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.10 | +/- | 0. |
| 7.67 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.36 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.47 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4650. | +/- | 19. |
| 4788. | +/- | 134. |
| 2.12 | +/- | 0. |
| 2.09 | +/- | 0. |
| 0.80 | +/- | 0. |
| 0.80 | +/- | -NaN |
| -1.59 | +/- | 0. |
| -1.59 | +/- | 0. |
| 0.77 | +/- | 0. |
| 0.77 | +/- | -NaN |
| 0.56 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.29 | +/- | 0. |
| 7.48 | +/- | 0. |
| 0.87 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.76 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -2.27 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 3900. | +/- | 3. |
| 3970. | +/- | 86. |
| 1.84 | +/- | 0. |
| 1.55 | +/- | 0. |
| 1.22 | +/- | 0. |
| 1.22 | +/- | -NaN |
| 0.42 | +/- | 0. |
| 0.42 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.31 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.26 |
| -9999.00 |
STAR_WARN,SN_WARN
010-07-C
| 4114. | +/- | 4. |
| 4202. | +/- | 97. |
| 1.83 | +/- | 0. |
| 1.60 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.43 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.08 | +/- | 0. |
| 2.94 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.16 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4878. | +/- | 70. |
| 4988. | +/- | 152. |
| 2.37 | +/- | 0. |
| 2.34 | +/- | 0. |
| 2.78 | +/- | 0. |
| 2.78 | +/- | -NaN |
| -1.54 | +/- | 0. |
| -1.54 | +/- | 0. |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.76 | +/- | 0. |
| 0.76 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.11 | +/- | 0. |
| 7.28 | +/- | 0. |
| 0.86 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.35 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.06 |
| -9999.00 |

STAR_BAD,COLORTE_BAD,SN_BAD
STAR_WARN,CHI2_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 6612. | +/- | 116. |
| -10000. | +/- | -NaN |
| 3.89 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.39 | +/- | 0. |
| 4.39 | +/- | -NaN |
| -1.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.47 | +/- | 0. |
| 0.74 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -2.44 |
| -9999.00 |
STAR_WARN,SN_WARN
010-07-C
| 4946. | +/- | 17. |
| 4973. | +/- | 119. |
| 3.15 | +/- | 0. |
| 2.78 | +/- | 0. |
| 1.39 | +/- | 0. |
| 1.39 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.44 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.60 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.09 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4739. | +/- | 40. |
| 4867. | +/- | 141. |
| 2.06 | +/- | 0. |
| 2.04 | +/- | 0. |
| 1.45 | +/- | 0. |
| 1.45 | +/- | -NaN |
| -1.59 | +/- | 0. |
| -1.59 | +/- | 0. |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.89 | +/- | 0. |
| 0.89 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.12 | +/- | 0. |
| 7.48 | +/- | 0. |
| 0.87 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -2.07 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
010-07-C
| 5118. | +/- | 118. |
| -10000. | +/- | -NaN |
| 2.68 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.92 | +/- | 0. |
| 0.92 | +/- | -NaN |
| -1.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.35 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.00 | +/- | 0. |
| 0.84 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.23 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 1.04 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -2.46 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 5032. | +/- | 81. |
| 5128. | +/- | 161. |
| 2.35 | +/- | 0. |
| 2.33 | +/- | 0. |
| 2.13 | +/- | 0. |
| 2.13 | +/- | -NaN |
| -1.64 | +/- | 0. |
| -1.64 | +/- | 0. |
| -0.17 | +/- | 0. |
| -0.17 | +/- | -NaN |
| -0.08 | +/- | 1. |
| -0.08 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.10 | +/- | 0. |
| 7.73 | +/- | 0. |
| 0.89 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -2.39 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 6395. | +/- | 65. |
| -10000. | +/- | -NaN |
| 2.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.05 | +/- | 0. |
| 2.05 | +/- | -NaN |
| -2.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.19 | +/- | 10. |
| -9999.99 | +/- | -NaN |
| 0.53 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 14.26 | +/- | 0. |
| 1.15 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -1.97 |
| -9999.00 |
STAR_WARN,SN_WARN
010-07-C
| 4954. | +/- | 42. |
| 5058. | +/- | 147. |
| 2.38 | +/- | 0. |
| 2.36 | +/- | 0. |
| 1.68 | +/- | 0. |
| 1.68 | +/- | -NaN |
| -1.60 | +/- | 0. |
| -1.60 | +/- | 0. |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.56 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.09 | +/- | 0. |
| 7.53 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -1.93 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.91 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 5153. | +/- | 32. |
| 5242. | +/- | 156. |
| 2.21 | +/- | 0. |
| 2.21 | +/- | 0. |
| 1.48 | +/- | 0. |
| 1.48 | +/- | -NaN |
| -1.78 | +/- | 0. |
| -1.78 | +/- | 0. |
| -0.17 | +/- | 0. |
| -0.17 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.44 | +/- | 0. |
| 0.41 | +/- | 0. |
| 8.38 | +/- | 0. |
| 0.92 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -1.93 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.31 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 4734. | +/- | 80. |
| -10000. | +/- | -NaN |
| 1.97 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.80 | +/- | 0. |
| 1.80 | +/- | -NaN |
| -1.81 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.24 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.18 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.54 | +/- | 0. |
| 0.93 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.24 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -0.59 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4716. | +/- | 37. |
| 4848. | +/- | 142. |
| 2.12 | +/- | 0. |
| 2.10 | +/- | 0. |
| 1.49 | +/- | 0. |
| 1.49 | +/- | -NaN |
| -1.61 | +/- | 0. |
| -1.61 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 1.15 | +/- | 0. |
| 1.15 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.25 | +/- | 0. |
| 7.59 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 1.15 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -2.31 |
| -9999.00 |

010-07-C
| 4562. | +/- | 7. |
| 4628. | +/- | 100. |
| 2.81 | +/- | 0. |
| 2.46 | +/- | 0. |
| 1.28 | +/- | 0. |
| 1.28 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.41 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.42 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.34 |
| -9999.00 |

STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 4844. | +/- | 50. |
| -10000. | +/- | -NaN |
| 2.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| -1.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.98 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.11 | +/- | 0. |
| 0.85 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.33 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.20 |
| -9999.00 |
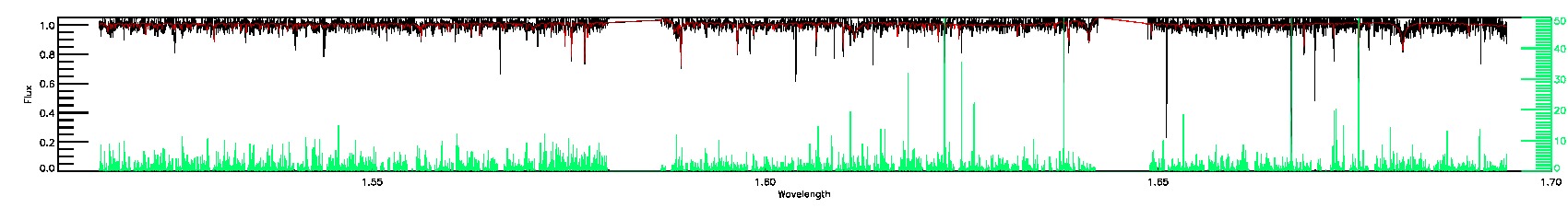
STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 4830. | +/- | 59. |
| 4964. | +/- | 155. |
| 2.16 | +/- | 0. |
| 2.19 | +/- | 0. |
| 1.22 | +/- | 0. |
| 1.22 | +/- | -NaN |
| -1.96 | +/- | 0. |
| -1.96 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.41 | +/- | 0. |
| 0.38 | +/- | 0. |
| 9.33 | +/- | 0. |
| 0.97 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -1.93 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.24 |
| -9999.00 |
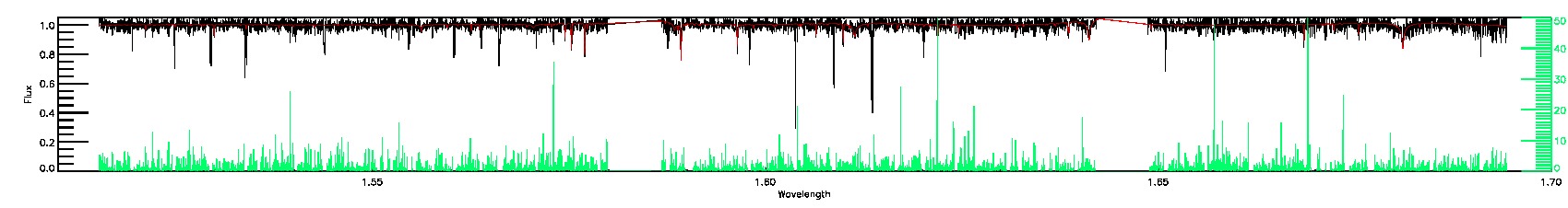
STAR_WARN,SN_WARN
010-07-C
| 4822. | +/- | 29. |
| 4936. | +/- | 139. |
| 2.20 | +/- | 0. |
| 2.16 | +/- | 0. |
| 1.16 | +/- | 0. |
| 1.16 | +/- | -NaN |
| -1.48 | +/- | 0. |
| -1.48 | +/- | 0. |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.83 | +/- | 0. |
| 0.83 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.15 | +/- | 0. |
| 7.03 | +/- | 0. |
| 0.85 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.31 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.33 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4911. | +/- | 82. |
| 5022. | +/- | 156. |
| 2.39 | +/- | 0. |
| 2.38 | +/- | 0. |
| 1.34 | +/- | 0. |
| 1.34 | +/- | -NaN |
| -1.64 | +/- | 0. |
| -1.64 | +/- | 0. |
| -0.36 | +/- | 0. |
| -0.36 | +/- | -NaN |
| 0.88 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.09 | +/- | 0. |
| 7.72 | +/- | 0. |
| 0.89 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.31 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| 1.10 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.62 |
| -9999.00 |
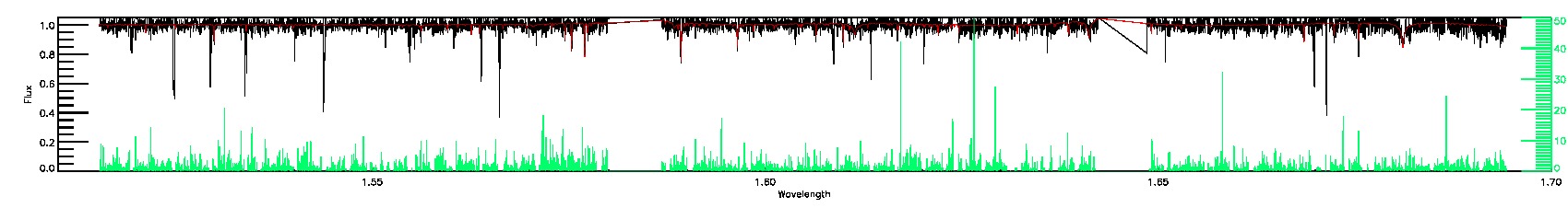
010-07-C
| 3972. | +/- | 2. |
| 4087. | +/- | 95. |
| 1.40 | +/- | 0. |
| 1.31 | +/- | 0. |
| 1.78 | +/- | 0. |
| 1.78 | +/- | -NaN |
| -0.62 | +/- | 0. |
| -0.62 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.27 | +/- | 0. |
| 4.26 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.30 |
| -9999.00 |
STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 4903. | +/- | 47. |
| 5013. | +/- | 150. |
| 2.32 | +/- | 0. |
| 2.30 | +/- | 0. |
| 1.49 | +/- | 0. |
| 1.49 | +/- | -NaN |
| -1.60 | +/- | 0. |
| -1.60 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.38 | +/- | 0. |
| 0.34 | +/- | 0. |
| 7.54 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.78 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4426. | +/- | 13. |
| 4524. | +/- | 114. |
| 2.56 | +/- | 0. |
| 2.37 | +/- | 0. |
| 1.23 | +/- | 0. |
| 1.23 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.24 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.19 | +/- | 0. |
| 3.40 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.03 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
010-07-C
| 4956. | +/- | 110. |
| -10000. | +/- | -NaN |
| 2.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.17 | +/- | 0. |
| 2.17 | +/- | -NaN |
| -1.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.22 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.36 | +/- | 0. |
| 0.92 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.10 |
| -9999.00 |

010-07-C
| 4033. | +/- | 3. |
| 4151. | +/- | 97. |
| 1.47 | +/- | 0. |
| 1.38 | +/- | 0. |
| 1.86 | +/- | 0. |
| 1.86 | +/- | -NaN |
| -0.71 | +/- | 0. |
| -0.71 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.25 | +/- | 0. |
| 4.47 | +/- | 0. |
| 0.65 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -1.49 |
| -9999.00 |

STAR_BAD,COLORTE_BAD,SN_BAD
STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 5820. | +/- | 136. |
| -10000. | +/- | -NaN |
| 3.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.37 | +/- | 0. |
| 4.37 | +/- | -NaN |
| -1.46 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 7. |
| -9999.99 | +/- | -NaN |
| 0.21 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.69 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| -1.92 |
| -9999.00 |
STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 4787. | +/- | 47. |
| 4914. | +/- | 147. |
| 2.20 | +/- | 0. |
| 2.19 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| -1.68 | +/- | 0. |
| -1.68 | +/- | 0. |
| 0.40 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.51 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.23 | +/- | 0. |
| 7.89 | +/- | 0. |
| 0.90 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.40 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.12 |
| -9999.00 |

010-07-C
| 4780. | +/- | 24. |
| 4903. | +/- | 133. |
| 2.11 | +/- | 0. |
| 2.08 | +/- | 0. |
| 1.48 | +/- | 0. |
| 1.48 | +/- | -NaN |
| -1.58 | +/- | 0. |
| -1.58 | +/- | 0. |
| -0.20 | +/- | 0. |
| -0.20 | +/- | -NaN |
| 0.63 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.09 | +/- | 0. |
| 7.46 | +/- | 0. |
| 0.87 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.25 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.28 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 5092. | +/- | 95. |
| -10000. | +/- | -NaN |
| 2.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.46 | +/- | 0. |
| 2.46 | +/- | -NaN |
| -1.61 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.89 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.37 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.59 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.10 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -1.69 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -1.69 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.24 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 5023. | +/- | 62. |
| 5125. | +/- | 159. |
| 2.34 | +/- | 0. |
| 2.34 | +/- | 0. |
| 2.72 | +/- | 0. |
| 2.72 | +/- | -NaN |
| -1.75 | +/- | 0. |
| -1.75 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.12 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.12 | +/- | 0. |
| 8.22 | +/- | 0. |
| 0.92 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 1.26 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.23 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
010-07-C
| 5677. | +/- | 113. |
| -10000. | +/- | -NaN |
| 3.21 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.24 | +/- | 0. |
| 3.24 | +/- | -NaN |
| -1.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.83 | +/- | 0. |
| 0.83 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.21 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.34 |
| -9999.00 |
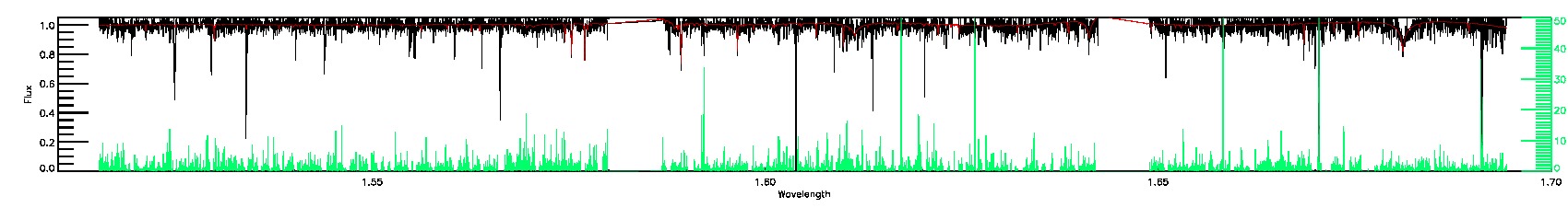
STAR_WARN,SN_WARN
010-07-C
| 4714. | +/- | 22. |
| 4788. | +/- | 123. |
| 2.65 | +/- | 0. |
| 2.36 | +/- | 0. |
| 1.52 | +/- | 0. |
| 1.52 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -0.22 | +/- | 0. |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.24 | +/- | 0. |
| 3.37 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.23 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.98 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 3975. | +/- | 3. |
| 4090. | +/- | 99. |
| 1.41 | +/- | 0. |
| 1.31 | +/- | 0. |
| 2.06 | +/- | 0. |
| 2.06 | +/- | -NaN |
| -0.62 | +/- | 0. |
| -0.62 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.22 | +/- | 0. |
| 4.25 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| -0.67 |
| -9999.00 |
STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 4902. | +/- | 80. |
| 5019. | +/- | 157. |
| 2.28 | +/- | 0. |
| 2.28 | +/- | 0. |
| 1.66 | +/- | 0. |
| 1.66 | +/- | -NaN |
| -1.77 | +/- | 0. |
| -1.77 | +/- | 0. |
| -0.61 | +/- | 0. |
| -0.61 | +/- | -NaN |
| -0.38 | +/- | 1. |
| -0.38 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.15 | +/- | 0. |
| 8.31 | +/- | 0. |
| 0.92 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.38 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -2.40 |
| -9999.00 |

STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 4793. | +/- | 32. |
| 4918. | +/- | 141. |
| 2.07 | +/- | 0. |
| 2.05 | +/- | 0. |
| 1.50 | +/- | 0. |
| 1.50 | +/- | -NaN |
| -1.66 | +/- | 0. |
| -1.66 | +/- | 0. |
| -0.30 | +/- | 0. |
| -0.30 | +/- | -NaN |
| 0.85 | +/- | 0. |
| 0.85 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.20 | +/- | 0. |
| 7.81 | +/- | 0. |
| 0.89 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.30 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| -1.69 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -1.69 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.50 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 6026. | +/- | 67. |
| 5929. | +/- | 174. |
| 4.52 | +/- | 0. |
| 4.30 | +/- | 0. |
| 1.05 | +/- | 0. |
| 1.05 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.19 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.00 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.52 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
010-07-C
| 4291. | +/- | 12. |
| -10000. | +/- | -NaN |
| 2.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.44 | +/- | 0. |
| 1.44 | +/- | -NaN |
| -0.53 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.22 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.04 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -1.31 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4974. | +/- | 62. |
| 5083. | +/- | 159. |
| 2.28 | +/- | 0. |
| 2.28 | +/- | 0. |
| 2.21 | +/- | 0. |
| 2.21 | +/- | -NaN |
| -1.78 | +/- | 0. |
| -1.78 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.47 | +/- | 0. |
| 0.44 | +/- | 0. |
| 8.37 | +/- | 0. |
| 0.92 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.38 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,COLORTE_WARN,SN_WARN
010-07-C
| 6620. | +/- | 127. |
| -10000. | +/- | -NaN |
| 4.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.16 | +/- | 0. |
| 3.16 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 9.60 | +/- | 0. |
| 0.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.98 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
010-07-C
| 4582. | +/- | 33. |
| -10000. | +/- | -NaN |
| 1.71 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.69 | +/- | 0. |
| 2.69 | +/- | -NaN |
| -1.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.30 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.90 | +/- | 0. |
| 0.77 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.17 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| -2.29 |
| -9999.00 |
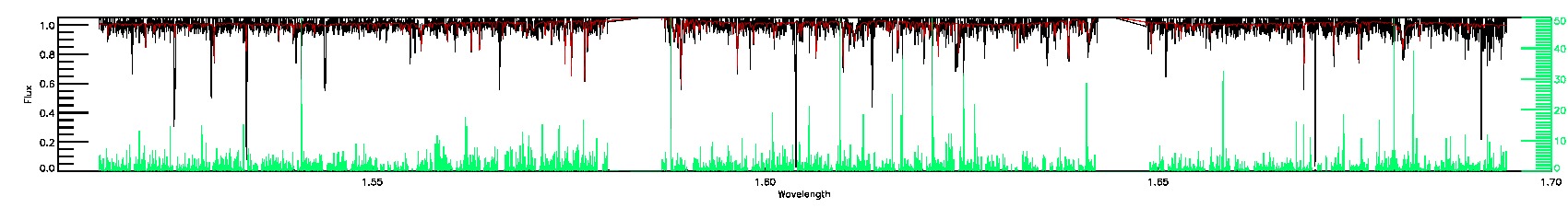
STAR_WARN,SN_WARN
010-07-C
| 4334. | +/- | 7. |
| 4429. | +/- | 106. |
| 2.33 | +/- | 0. |
| 2.12 | +/- | 0. |
| 1.33 | +/- | 0. |
| 1.33 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.07 | +/- | 0. |
| 3.22 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.11 |
| -9999.00 |

BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
010-07-C
| 4530. | +/- | 10. |
| 4618. | +/- | 102. |
| 4.07 | +/- | 0. |
| 4.23 | +/- | 0. |
| 1.09 | +/- | 0. |
| 1.09 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 39.04 | +/- | 0. |
| 1.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.23 |
| -9999.00 |

STAR_WARN,SN_WARN
010-07-C
| 4084. | +/- | 5. |
| 4191. | +/- | 102. |
| 1.89 | +/- | 0. |
| 1.71 | +/- | 0. |
| 1.42 | +/- | 0. |
| 1.42 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -0.44 | +/- | 0. |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.19 | +/- | 0. |
| 3.83 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| 0.11 |
| -9999.00 |

