055-53-O
| 4369. | +/- | 2. |
| 4464. | +/- | 77. |
| 4.39 | +/- | 0. |
| 4.63 | +/- | 0. |
| 0.57 | +/- | 0. |
| 0.57 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.16 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.11 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.34 |
| -9999.00 |

055-53-O
| 4884. | +/- | 5. |
| 4916. | +/- | 86. |
| 4.33 | +/- | 0. |
| 4.28 | +/- | 0. |
| 1.17 | +/- | 0. |
| 1.17 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.12 | +/- | 0. |
| 1.72 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.32 |
| -9999.00 |
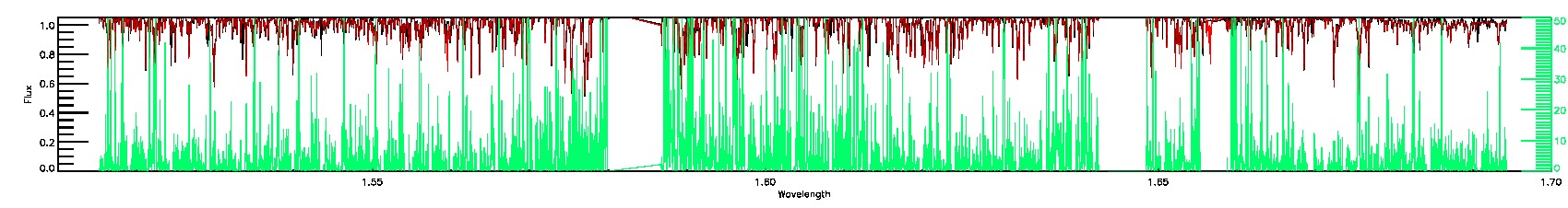
055-53-O
| 4901. | +/- | 10. |
| 4951. | +/- | 107. |
| 3.40 | +/- | 0. |
| 3.26 | +/- | 0. |
| 1.14 | +/- | 0. |
| 1.14 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -0.19 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.30 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.77 |
| -9999.00 |
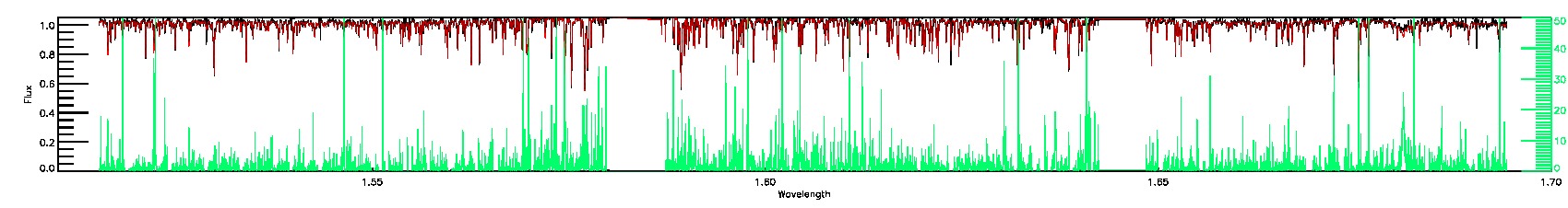
055-53-O
| 4816. | +/- | 10. |
| 4878. | +/- | 109. |
| 4.45 | +/- | 0. |
| 4.63 | +/- | 0. |
| 1.52 | +/- | 0. |
| 1.52 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.23 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.02 | +/- | 0. |
| 1.83 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.67 |
| -9999.00 |
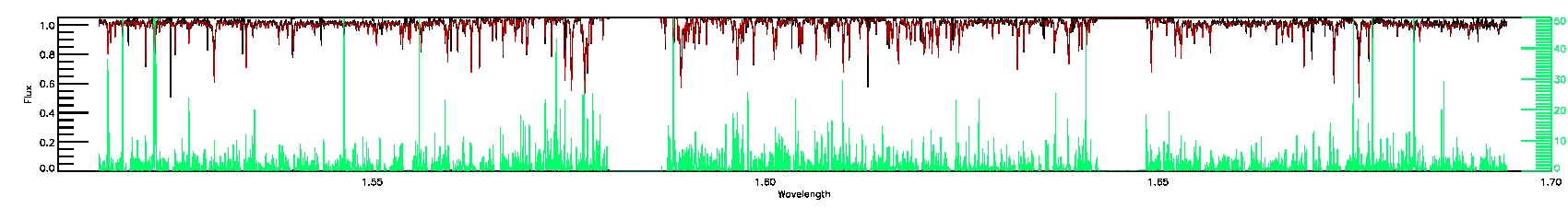
SUSPECT_BROAD_LINES
STAR_BAD
055-53-O
| 4799. | +/- | 7. |
| -10000. | +/- | -NaN |
| 3.93 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.77 | +/- | 0. |
| 4.77 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 62.57 | +/- | 0. |
| 1.80 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.04 |
| -9999.00 |
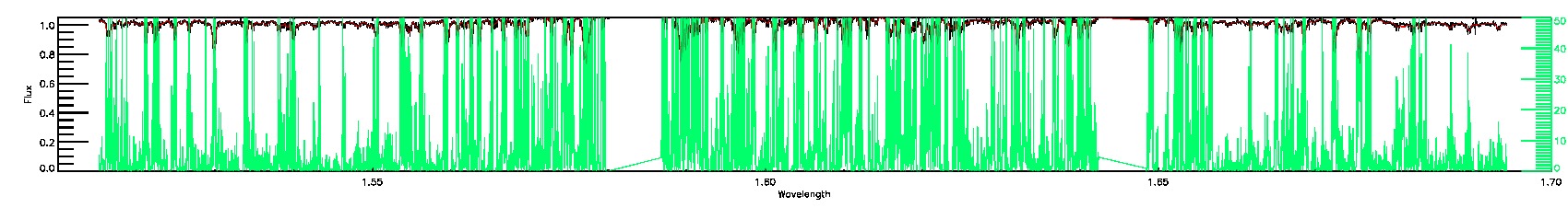
055-53-O
| 4830. | +/- | 10. |
| 4885. | +/- | 108. |
| 3.62 | +/- | 0. |
| 3.56 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.12 | +/- | 0. |
| 3.16 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -1.02 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD
055-53-O
| 4874. | +/- | 22. |
| -10000. | +/- | -NaN |
| 3.59 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.50 | +/- | 0. |
| 1.50 | +/- | -NaN |
| -0.73 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 96.01 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.68 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
055-53-O
| 3527. | +/- | 2. |
| 3622. | +/- | 60. |
| 4.41 | +/- | 0. |
| 4.81 | +/- | 0. |
| 1.20 | +/- | 0. |
| 1.20 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.16 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -0.44 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 5.05 | +/- | 0. |
| 0.70 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.20 |
| -9999.00 |
055-53-O
| 4670. | +/- | 4. |
| 4748. | +/- | 85. |
| 2.67 | +/- | 0. |
| 2.37 | +/- | 0. |
| 1.38 | +/- | 0. |
| 1.38 | +/- | -NaN |
| -0.21 | +/- | 0. |
| -0.21 | +/- | 0. |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.18 | +/- | 0. |
| 3.35 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |

055-53-O
| 4621. | +/- | 8. |
| 4733. | +/- | 108. |
| 2.26 | +/- | 0. |
| 2.14 | +/- | 0. |
| 1.74 | +/- | 0. |
| 1.74 | +/- | -NaN |
| -0.88 | +/- | 0. |
| -0.88 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.13 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.13 | +/- | 0. |
| 4.96 | +/- | 0. |
| 0.70 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.55 |
| -9999.00 |

055-53-O
| 4929. | +/- | 8. |
| 4993. | +/- | 99. |
| 3.41 | +/- | 0. |
| 3.32 | +/- | 0. |
| 1.17 | +/- | 0. |
| 1.17 | +/- | -NaN |
| -0.58 | +/- | 0. |
| -0.58 | +/- | 0. |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| -0.27 | +/- | 0. |
| -0.27 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.22 | +/- | 0. |
| 4.14 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.68 |
| -9999.00 |

055-53-O
| 4751. | +/- | 9. |
| 4809. | +/- | 106. |
| 4.51 | +/- | 0. |
| 4.59 | +/- | 0. |
| 1.41 | +/- | 0. |
| 1.41 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 1.66 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.16 |
| -9999.00 |
055-53-O
| 5630. | +/- | 14. |
| 5594. | +/- | 122. |
| 4.46 | +/- | 0. |
| 4.45 | +/- | 0. |
| 1.48 | +/- | 0. |
| 1.48 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -0.14 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 1.66 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.09 |
| -9999.00 |
055-53-O
| 5416. | +/- | 14. |
| 5397. | +/- | 119. |
| 4.28 | +/- | 0. |
| 4.25 | +/- | 0. |
| 0.70 | +/- | 0. |
| 0.70 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 5.51 | +/- | 0. |
| 0.74 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 1.00 |
| -9999.00 |
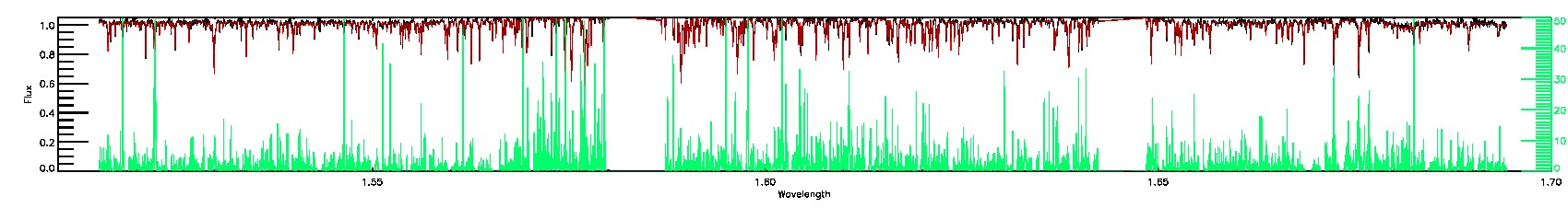
055-53-O
| 5500. | +/- | 10. |
| 5473. | +/- | 109. |
| 4.50 | +/- | 0. |
| 4.47 | +/- | 0. |
| 0.51 | +/- | 0. |
| 0.51 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.15 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.97 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.05 |
| -9999.00 |
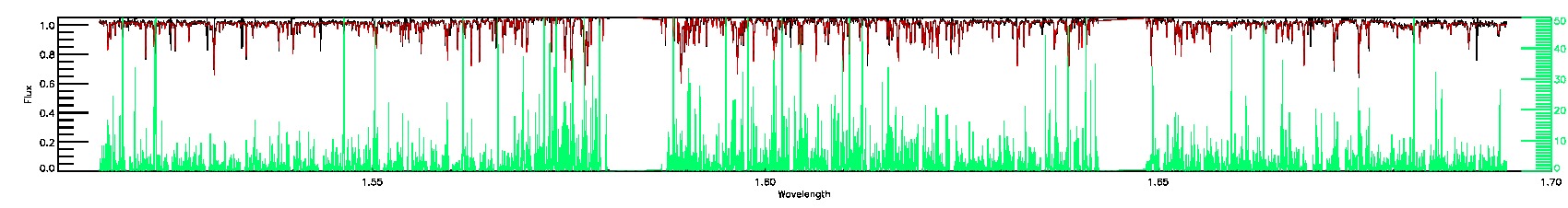
055-53-O
| 4872. | +/- | 5. |
| 4916. | +/- | 88. |
| 3.48 | +/- | 0. |
| 3.32 | +/- | 0. |
| 0.80 | +/- | 0. |
| 0.80 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.93 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.00 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
055-53-O
| 3608. | +/- | 2. |
| 3700. | +/- | 62. |
| 4.49 | +/- | 0. |
| 4.84 | +/- | 0. |
| 1.08 | +/- | 0. |
| 1.08 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -0.40 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 3.47 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.06 |
| -9999.00 |
055-53-O
| 5286. | +/- | 25. |
| 5312. | +/- | 138. |
| 3.83 | +/- | 0. |
| 4.00 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| -0.67 | +/- | 0. |
| -0.67 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.25 | +/- | 0. |
| 4.38 | +/- | 0. |
| 0.64 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.66 |
| -9999.00 |

055-53-O
| 4076. | +/- | 3. |
| 4167. | +/- | 77. |
| 4.34 | +/- | 0. |
| 4.59 | +/- | 0. |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.06 | +/- | 0. |
| 5.71 | +/- | 0. |
| 0.76 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.09 |
| -9999.00 |

055-53-O
| 5697. | +/- | 15. |
| 5646. | +/- | 123. |
| 4.46 | +/- | 0. |
| 4.37 | +/- | 0. |
| 0.83 | +/- | 0. |
| 0.83 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.03 | +/- | 0. |
| -0.16 | +/- | 0. |
| -0.16 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 6.74 | +/- | 0. |
| 0.83 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.04 |
| -9999.00 |

055-53-O
| 5617. | +/- | 10. |
| 5572. | +/- | 108. |
| 4.63 | +/- | 0. |
| 4.53 | +/- | 0. |
| 1.44 | +/- | 0. |
| 1.44 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | 0. |
| -0.31 | +/- | 0. |
| -0.31 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 5.36 | +/- | 0. |
| 0.73 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.22 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.40 |
| -9999.00 |

055-53-O
| 4317. | +/- | 2. |
| 4432. | +/- | 80. |
| 4.35 | +/- | 0. |
| 4.78 | +/- | 0. |
| 0.81 | +/- | 0. |
| 0.81 | +/- | -NaN |
| -0.63 | +/- | 0. |
| -0.62 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.17 | +/- | 0. |
| 2.07 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.67 |
| -9999.00 |

055-53-O
| 5779. | +/- | 16. |
| 5723. | +/- | 132. |
| 4.45 | +/- | 0. |
| 4.40 | +/- | 0. |
| 1.17 | +/- | 0. |
| 1.17 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.16 | +/- | 0. |
| -0.16 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.02 | +/- | 0. |
| 3.40 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.11 |
| -9999.00 |
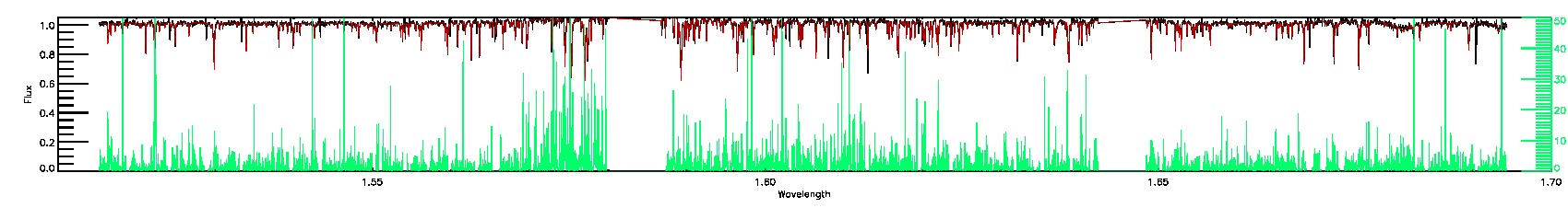
STAR_WARN,COLORTE_WARN
055-53-O
| 3677. | +/- | 6. |
| 3789. | +/- | 85. |
| 4.54 | +/- | 0. |
| 5.07 | +/- | 0. |
| 1.04 | +/- | 0. |
| 1.04 | +/- | -NaN |
| -0.55 | +/- | 0. |
| -0.55 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -0.40 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.00 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.02 |
| -9999.00 |
055-53-O
| 5420. | +/- | 10. |
| 5398. | +/- | 114. |
| 4.47 | +/- | 0. |
| 4.41 | +/- | 0. |
| 1.08 | +/- | 0. |
| 1.08 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.98 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.01 |
| -9999.00 |

055-53-O
| 3621. | +/- | 1. |
| 3729. | +/- | 64. |
| 0.78 | +/- | 0. |
| 0.69 | +/- | 0. |
| 1.97 | +/- | 0. |
| 1.97 | +/- | -NaN |
| -0.48 | +/- | 0. |
| -0.48 | +/- | 0. |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.26 | +/- | 0. |
| 3.91 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.85 |
| -9999.00 |
055-53-O
| 5856. | +/- | 15. |
| 5786. | +/- | 135. |
| 4.51 | +/- | 0. |
| 4.38 | +/- | 0. |
| 0.96 | +/- | 0. |
| 0.96 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.54 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.01 |
| -9999.00 |
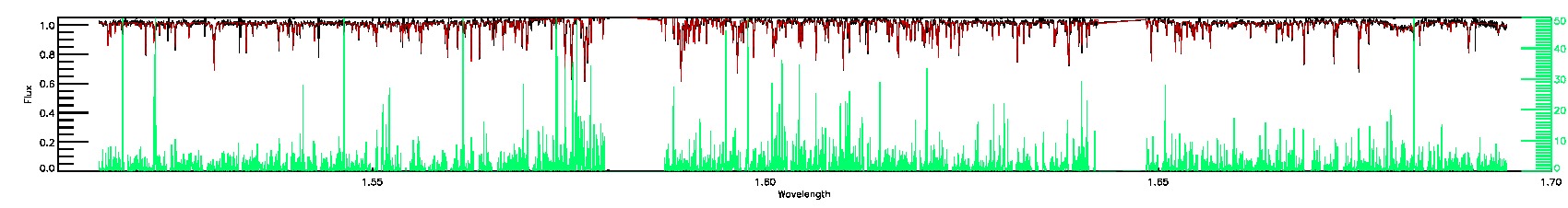
055-53-O
| 5299. | +/- | 9. |
| 5294. | +/- | 108. |
| 4.41 | +/- | 0. |
| 4.39 | +/- | 0. |
| 1.03 | +/- | 0. |
| 1.03 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.03 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 1.78 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.91 |
| -9999.00 |

055-53-O
| 5230. | +/- | 8. |
| 5235. | +/- | 102. |
| 4.60 | +/- | 0. |
| 4.63 | +/- | 0. |
| 1.64 | +/- | 0. |
| 1.64 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.31 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.11 |
| -9999.00 |

055-53-O
| 6154. | +/- | 15. |
| 6051. | +/- | 129. |
| 4.38 | +/- | 0. |
| 4.22 | +/- | 0. |
| 1.60 | +/- | 0. |
| 1.60 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| 0.46 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 5.85 | +/- | 0. |
| 0.77 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.46 |
| -9999.00 |
055-53-O
| 5815. | +/- | 11. |
| 5750. | +/- | 124. |
| 4.35 | +/- | 0. |
| 4.23 | +/- | 0. |
| 1.07 | +/- | 0. |
| 1.07 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.04 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 3.25 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.89 |
| -9999.00 |
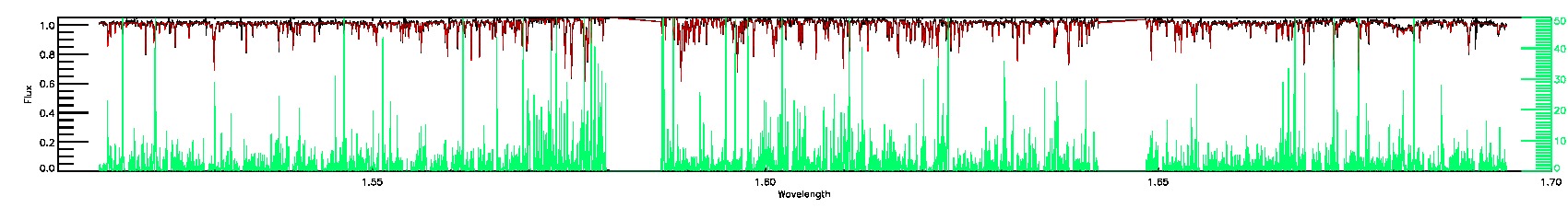
055-53-O
| 5278. | +/- | 19. |
| 5288. | +/- | 127. |
| 4.43 | +/- | 0. |
| 4.54 | +/- | 0. |
| 1.60 | +/- | 0. |
| 1.60 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -0.28 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.15 | +/- | 0. |
| 1.77 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.20 |
| -9999.00 |
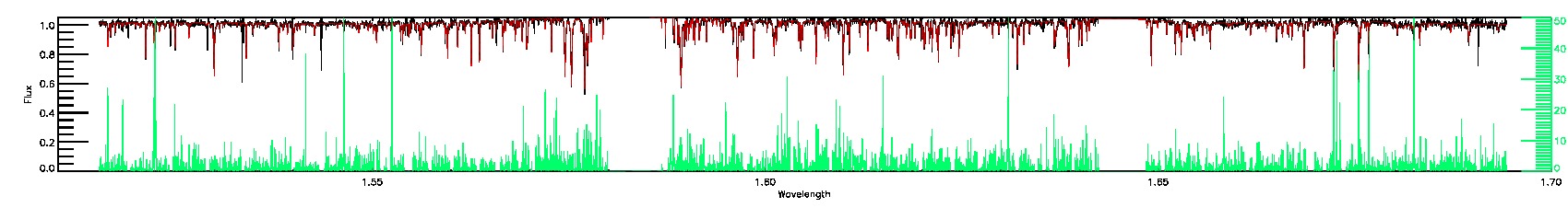
BRIGHT_NEIGHBOR
055-53-O
| 4461. | +/- | 5. |
| 4562. | +/- | 100. |
| 4.19 | +/- | 0. |
| 4.46 | +/- | 0. |
| 1.80 | +/- | 0. |
| 1.80 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -0.29 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 13.87 | +/- | 0. |
| 1.14 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.14 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.63 |
| -9999.00 |

055-53-O
| 5709. | +/- | 17. |
| 5668. | +/- | 130. |
| 4.08 | +/- | 0. |
| 4.11 | +/- | 0. |
| 1.18 | +/- | 0. |
| 1.18 | +/- | -NaN |
| -0.27 | +/- | 0. |
| -0.26 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.06 | +/- | 0. |
| 3.46 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 1.00 |
| -9999.00 |

055-53-O
| 5658. | +/- | 18. |
| 5611. | +/- | 134. |
| 4.43 | +/- | 0. |
| 4.34 | +/- | 0. |
| 1.49 | +/- | 0. |
| 1.49 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.03 | +/- | 0. |
| -0.14 | +/- | 0. |
| -0.14 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 1.74 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.99 |
| -9999.00 |

055-53-O
| 4740. | +/- | 5. |
| 4822. | +/- | 90. |
| 2.86 | +/- | 0. |
| 2.72 | +/- | 0. |
| 1.04 | +/- | 0. |
| 1.04 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -0.50 | +/- | 0. |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.21 | +/- | 0. |
| 3.95 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.56 |
| -9999.00 |

055-53-O
| 6137. | +/- | 36. |
| 6038. | +/- | 164. |
| 4.48 | +/- | 0. |
| 4.34 | +/- | 0. |
| 1.23 | +/- | 0. |
| 1.23 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.87 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -0.19 |
| -9999.00 |
055-53-O
| 4493. | +/- | 3. |
| 4605. | +/- | 84. |
| 2.51 | +/- | 0. |
| 2.36 | +/- | 0. |
| 1.36 | +/- | 0. |
| 1.36 | +/- | -NaN |
| -0.55 | +/- | 0. |
| -0.55 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.21 | +/- | 0. |
| 4.09 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.83 |
| -9999.00 |

BRIGHT_NEIGHBOR
STAR_WARN,COLORTE_WARN
055-53-O
| 3476. | +/- | 4. |
| 3571. | +/- | 74. |
| 4.58 | +/- | 0. |
| 4.98 | +/- | 0. |
| 1.36 | +/- | 0. |
| 1.36 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -0.16 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.06 | +/- | 0. |
| 2.24 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.18 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
055-53-O
| 6395. | +/- | 24. |
| 6253. | +/- | 166. |
| 5.10 | +/- | 0. |
| 4.79 | +/- | 0. |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| 0.95 | +/- | 0. |
| 0.95 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 7.32 | +/- | 0. |
| 0.86 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.22 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.99 |
| -9999.00 |
055-53-O
| 5806. | +/- | 26. |
| 5741. | +/- | 152. |
| 4.18 | +/- | 0. |
| 4.05 | +/- | 0. |
| 1.39 | +/- | 0. |
| 1.39 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 3.39 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.47 |
| -9999.00 |

055-53-O
| 5669. | +/- | 15. |
| 5615. | +/- | 126. |
| 4.39 | +/- | 0. |
| 4.25 | +/- | 0. |
| 0.78 | +/- | 0. |
| 0.78 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.61 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.12 |
| -9999.00 |
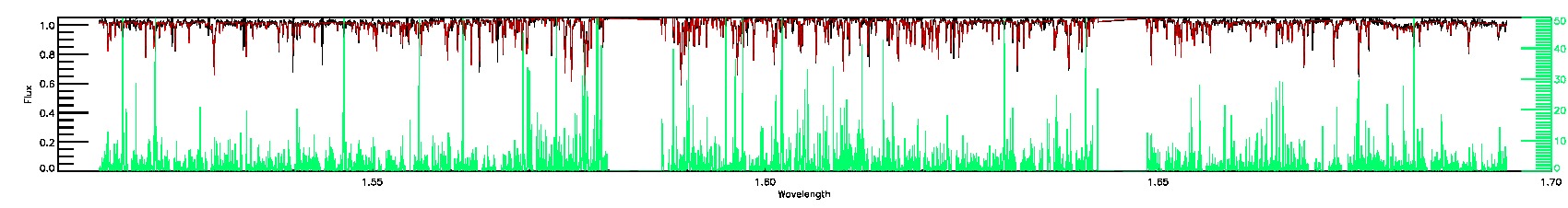
055-53-O
| 5451. | +/- | 33. |
| 5452. | +/- | 145. |
| 3.77 | +/- | 0. |
| 3.84 | +/- | 0. |
| 0.87 | +/- | 0. |
| 0.87 | +/- | -NaN |
| -0.55 | +/- | 0. |
| -0.55 | +/- | 0. |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -0.20 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.10 | +/- | 0. |
| 4.08 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.27 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.03 |
| -9999.00 |

055-53-O
| 5219. | +/- | 14. |
| 5223. | +/- | 119. |
| 4.59 | +/- | 0. |
| 4.59 | +/- | 0. |
| 1.65 | +/- | 0. |
| 1.65 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 4.88 | +/- | 0. |
| 0.69 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.53 |
| -9999.00 |

055-53-O
| 4619. | +/- | 7. |
| 4705. | +/- | 104. |
| 4.43 | +/- | 0. |
| 4.66 | +/- | 0. |
| 0.87 | +/- | 0. |
| 0.87 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -0.25 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.06 | +/- | 0. |
| 1.63 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.41 |
| -9999.00 |

055-53-O
| 3812. | +/- | 1. |
| 3923. | +/- | 68. |
| 0.99 | +/- | 0. |
| 0.90 | +/- | 0. |
| 1.96 | +/- | 0. |
| 1.96 | +/- | -NaN |
| -0.52 | +/- | 0. |
| -0.52 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.08 | +/- | 0. |
| 4.01 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.76 |
| -9999.00 |
055-53-O
| 5779. | +/- | 22. |
| 5724. | +/- | 141. |
| 4.09 | +/- | 0. |
| 4.04 | +/- | 0. |
| 1.23 | +/- | 0. |
| 1.23 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.18 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -2.49 |
| -9999.00 |

055-53-O
| 4838. | +/- | 14. |
| 4896. | +/- | 115. |
| 4.45 | +/- | 0. |
| 4.61 | +/- | 0. |
| 1.76 | +/- | 0. |
| 1.76 | +/- | -NaN |
| -0.21 | +/- | 0. |
| -0.20 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 1.86 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -1.69 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.36 |
| -9999.00 |

055-53-O
| 5772. | +/- | 12. |
| 5715. | +/- | 115. |
| 4.52 | +/- | 0. |
| 4.44 | +/- | 0. |
| 0.67 | +/- | 0. |
| 0.67 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 8.53 | +/- | 0. |
| 0.93 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.14 |
| -9999.00 |

055-53-O
| 4590. | +/- | 6. |
| 4693. | +/- | 104. |
| 2.60 | +/- | 0. |
| 2.46 | +/- | 0. |
| 1.20 | +/- | 0. |
| 1.20 | +/- | -NaN |
| -0.59 | +/- | 0. |
| -0.59 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.22 | +/- | 0. |
| 4.17 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.30 |
| -9999.00 |
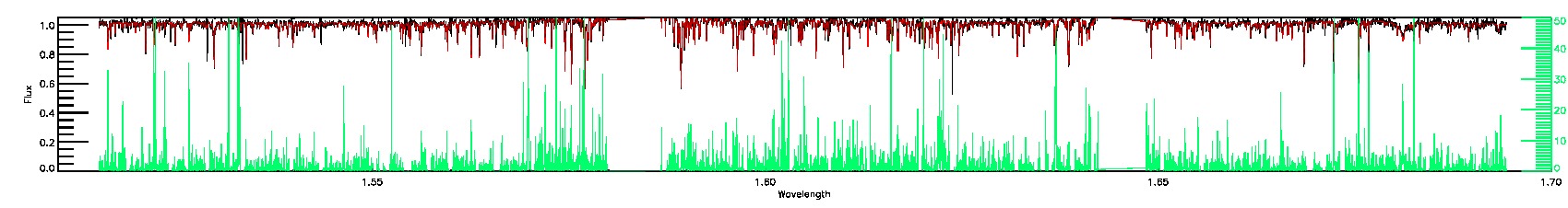
055-53-O
| 3863. | +/- | 5. |
| 3961. | +/- | 85. |
| 4.49 | +/- | 0. |
| 4.85 | +/- | 0. |
| 0.46 | +/- | 0. |
| 0.46 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -0.16 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.46 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.91 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
055-53-O
| 6412. | +/- | 35. |
| 6283. | +/- | 174. |
| 4.74 | +/- | 0. |
| 4.57 | +/- | 0. |
| 0.99 | +/- | 0. |
| 0.99 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.18 | +/- | 0. |
| -0.18 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 4.00 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| 0.06 |
| -9999.00 |
055-53-O
| 3897. | +/- | 1. |
| 4017. | +/- | 72. |
| 1.22 | +/- | 0. |
| 1.16 | +/- | 0. |
| 2.00 | +/- | 0. |
| 2.00 | +/- | -NaN |
| -0.73 | +/- | 0. |
| -0.73 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.26 | +/- | 0. |
| 4.54 | +/- | 0. |
| 0.66 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.94 |
| -9999.00 |
055-53-O
| 4447. | +/- | 3. |
| 4539. | +/- | 83. |
| 4.47 | +/- | 0. |
| 4.66 | +/- | 0. |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.00 | +/- | 0. |
| 4.22 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |

055-53-O
| 4760. | +/- | 8. |
| 4821. | +/- | 103. |
| 4.38 | +/- | 0. |
| 4.49 | +/- | 0. |
| 0.42 | +/- | 0. |
| 0.42 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 3.63 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.10 |
| -9999.00 |
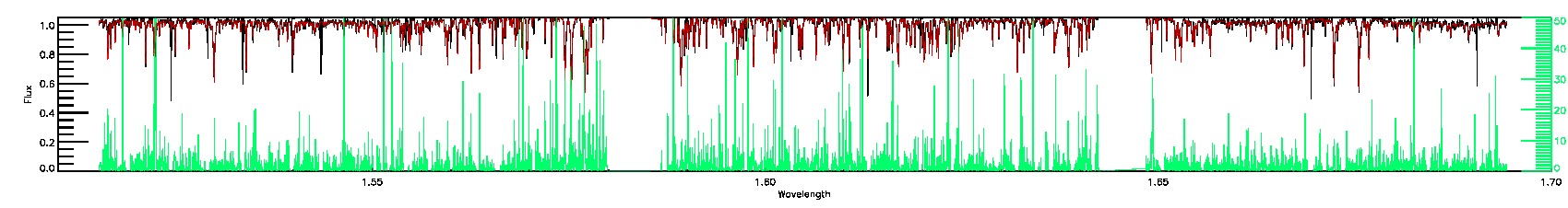
BRIGHT_NEIGHBOR
055-53-O
| 5268. | +/- | 17. |
| 5278. | +/- | 121. |
| 4.54 | +/- | 0. |
| 4.65 | +/- | 0. |
| 1.46 | +/- | 0. |
| 1.46 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -0.25 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 8.01 | +/- | 0. |
| 0.90 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.08 |
| -9999.00 |

055-53-O
| 5734. | +/- | 19. |
| 5678. | +/- | 133. |
| 4.08 | +/- | 0. |
| 3.99 | +/- | 0. |
| 1.49 | +/- | 0. |
| 1.49 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | 0. |
| -0.18 | +/- | 0. |
| -0.18 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.49 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.09 |
| -9999.00 |
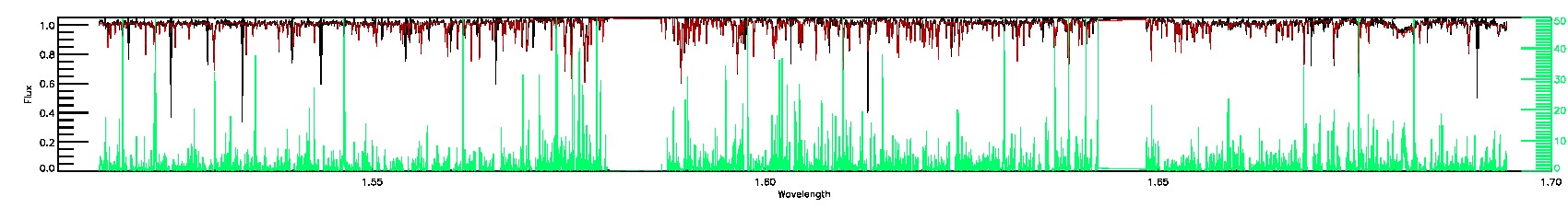
BRIGHT_NEIGHBOR
STAR_WARN,COLORTE_WARN
055-53-O
| 3649. | +/- | 5. |
| 3759. | +/- | 84. |
| 4.25 | +/- | 0. |
| 4.78 | +/- | 0. |
| 0.41 | +/- | 0. |
| 0.41 | +/- | -NaN |
| -0.52 | +/- | 0. |
| -0.52 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.06 | +/- | 0. |
| 6.41 | +/- | 0. |
| 0.81 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.69 |
| -9999.00 |
055-53-O
| 6123. | +/- | 14. |
| 6019. | +/- | 126. |
| 4.46 | +/- | 0. |
| 4.25 | +/- | 0. |
| 1.25 | +/- | 0. |
| 1.25 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 5.38 | +/- | 0. |
| 0.73 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.02 |
| -9999.00 |
055-53-O
| 4824. | +/- | 7. |
| 4863. | +/- | 97. |
| 3.71 | +/- | 0. |
| 3.60 | +/- | 0. |
| 0.40 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.53 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.43 |
| -9999.00 |
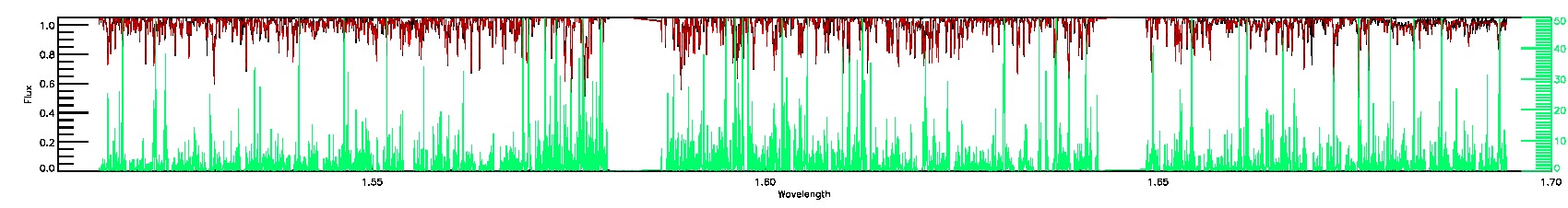
STAR_WARN,SN_WARN
055-53-O
| 4834. | +/- | 20. |
| 4894. | +/- | 121. |
| 3.50 | +/- | 0. |
| 3.38 | +/- | 0. |
| 0.72 | +/- | 0. |
| 0.72 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.24 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.09 | +/- | 0. |
| 3.41 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -1.16 |
| -9999.00 |

BRIGHT_NEIGHBOR
055-53-O
| 4741. | +/- | 9. |
| 4826. | +/- | 110. |
| 2.97 | +/- | 0. |
| 2.85 | +/- | 0. |
| 1.28 | +/- | 0. |
| 1.28 | +/- | -NaN |
| -0.57 | +/- | 0. |
| -0.57 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.26 | +/- | 0. |
| 4.12 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.42 |
| -9999.00 |

055-53-O
| 5996. | +/- | 16. |
| 5918. | +/- | 125. |
| 4.40 | +/- | 0. |
| 4.34 | +/- | 0. |
| 1.21 | +/- | 0. |
| 1.21 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.16 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.12 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.05 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.12 |
| -9999.00 |
055-53-O
| 5602. | +/- | 23. |
| 5591. | +/- | 140. |
| 3.86 | +/- | 0. |
| 4.00 | +/- | 0. |
| 1.06 | +/- | 0. |
| 1.06 | +/- | -NaN |
| -0.67 | +/- | 0. |
| -0.67 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.22 | +/- | 0. |
| 4.38 | +/- | 0. |
| 0.64 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -1.06 |
| -9999.00 |

055-53-O
| 5858. | +/- | 15. |
| 5804. | +/- | 123. |
| 4.40 | +/- | 0. |
| 4.44 | +/- | 0. |
| 1.17 | +/- | 0. |
| 1.17 | +/- | -NaN |
| -0.37 | +/- | 0. |
| -0.36 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.82 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.08 |
| -9999.00 |

055-53-O
| 4557. | +/- | 3. |
| 4635. | +/- | 86. |
| 4.41 | +/- | 0. |
| 4.51 | +/- | 0. |
| 0.96 | +/- | 0. |
| 0.96 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.00 | +/- | 0. |
| 1.67 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.18 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_WARN,COLORTE_WARN
055-53-O
| 3446. | +/- | 2. |
| 3536. | +/- | 58. |
| 4.72 | +/- | 0. |
| 5.08 | +/- | 0. |
| 2.20 | +/- | 0. |
| 2.20 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.33 | +/- | 0. |
| -0.33 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 20.94 | +/- | 0. |
| 1.32 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.50 |
| -9999.00 |
055-53-O
| 4876. | +/- | 15. |
| 4921. | +/- | 116. |
| 4.59 | +/- | 0. |
| 4.67 | +/- | 0. |
| 0.92 | +/- | 0. |
| 0.92 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 4.16 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.13 |
| -9999.00 |

055-53-O
| 6203. | +/- | 20. |
| 6102. | +/- | 133. |
| 4.79 | +/- | 0. |
| 4.69 | +/- | 0. |
| 0.65 | +/- | 0. |
| 0.65 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -0.18 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.12 | +/- | 0. |
| 14.54 | +/- | 0. |
| 1.16 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.18 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -1.12 |
| -9999.00 |
055-53-O
| 5048. | +/- | 15. |
| 5081. | +/- | 120. |
| 3.67 | +/- | 0. |
| 3.64 | +/- | 0. |
| 1.33 | +/- | 0. |
| 1.33 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -0.19 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.06 | +/- | 0. |
| 1.67 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.31 |
| -9999.00 |

055-53-O
| 4903. | +/- | 5. |
| 4928. | +/- | 85. |
| 3.79 | +/- | 0. |
| 3.66 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.36 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 0.45 |
| -9999.00 |

055-53-O
| 4818. | +/- | 5. |
| 4888. | +/- | 91. |
| 2.75 | +/- | 0. |
| 2.42 | +/- | 0. |
| 1.57 | +/- | 0. |
| 1.57 | +/- | -NaN |
| -0.43 | +/- | 0. |
| -0.43 | +/- | 0. |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.20 | +/- | 0. |
| 3.80 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.22 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.45 |
| -9999.00 |

BRIGHT_NEIGHBOR
055-53-O
| 6126. | +/- | 32. |
| 6025. | +/- | 161. |
| 4.43 | +/- | 0. |
| 4.26 | +/- | 0. |
| 1.24 | +/- | 0. |
| 1.24 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.37 | +/- | 0. |
| 0.37 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 5.66 | +/- | 0. |
| 0.75 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -1.77 |
| -9999.00 |
STAR_WARN,SN_WARN
055-53-O
| 4851. | +/- | 18. |
| 4910. | +/- | 121. |
| 4.52 | +/- | 0. |
| 4.71 | +/- | 0. |
| 0.89 | +/- | 0. |
| 0.89 | +/- | -NaN |
| -0.27 | +/- | 0. |
| -0.27 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.06 | +/- | 0. |
| 2.85 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.84 |
| -9999.00 |
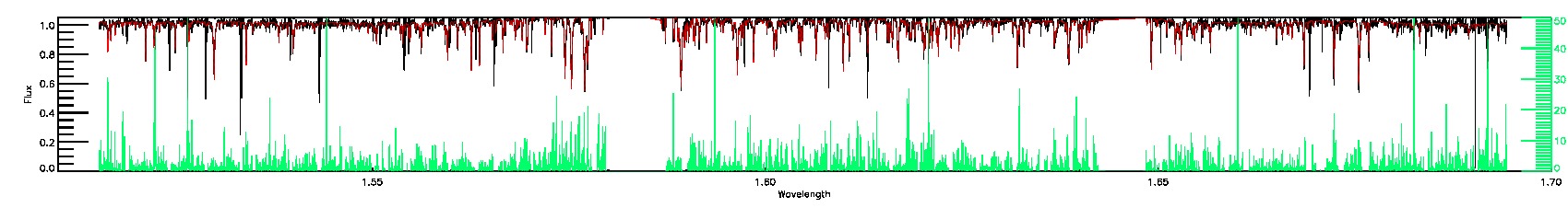
055-53-O
| 6248. | +/- | 14. |
| 6127. | +/- | 130. |
| 4.55 | +/- | 0. |
| 4.30 | +/- | 0. |
| 1.16 | +/- | 0. |
| 1.16 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | 0. |
| -0.13 | +/- | 0. |
| -0.13 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 7.87 | +/- | 0. |
| 0.90 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.38 |
| -9999.00 |
BRIGHT_NEIGHBOR
055-53-O
| 3947. | +/- | 3. |
| 4048. | +/- | 81. |
| 4.31 | +/- | 0. |
| 4.69 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -0.31 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.34 | +/- | 0. |
| -0.34 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 4.88 | +/- | 0. |
| 0.69 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.39 |
| -9999.00 |
055-53-O
| 5479. | +/- | 27. |
| 5464. | +/- | 140. |
| 4.13 | +/- | 0. |
| 4.20 | +/- | 0. |
| 1.61 | +/- | 0. |
| 1.61 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -0.25 | +/- | 0. |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.14 | +/- | 0. |
| 1.82 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.21 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.93 |
| -9999.00 |
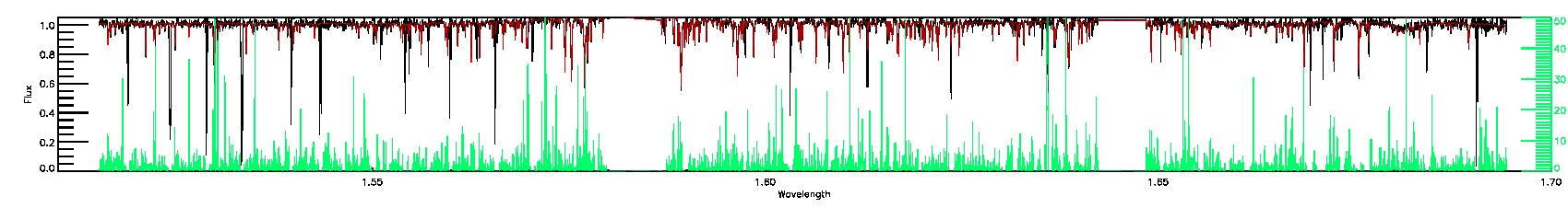
055-53-O
| 6017. | +/- | 25. |
| 5924. | +/- | 151. |
| 4.60 | +/- | 0. |
| 4.41 | +/- | 0. |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.06 | +/- | 0. |
| 6.74 | +/- | 0. |
| 0.83 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.48 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_WARN,COLORTE_WARN,SN_WARN
055-53-O
| 3186. | +/- | 9. |
| 3289. | +/- | 75. |
| 4.55 | +/- | 0. |
| 5.08 | +/- | 0. |
| 2.73 | +/- | 0. |
| 2.73 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -0.34 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.05 | +/- | 0. |
| 29.10 | +/- | 0. |
| 1.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.97 |
| -9999.00 |
055-53-O
| 5945. | +/- | 12. |
| 5867. | +/- | 121. |
| 4.40 | +/- | 0. |
| 4.29 | +/- | 0. |
| 1.40 | +/- | 0. |
| 1.40 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.22 | +/- | -NaN |
| 0.48 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.39 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.25 |
| -9999.00 |

055-53-O
| 5838. | +/- | 26. |
| 5781. | +/- | 147. |
| 4.38 | +/- | 0. |
| 4.37 | +/- | 0. |
| 1.19 | +/- | 0. |
| 1.19 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.27 | +/- | 0. |
| -0.27 | +/- | -NaN |
| 0.42 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 4.93 | +/- | 0. |
| 0.69 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.54 |
| -9999.00 |
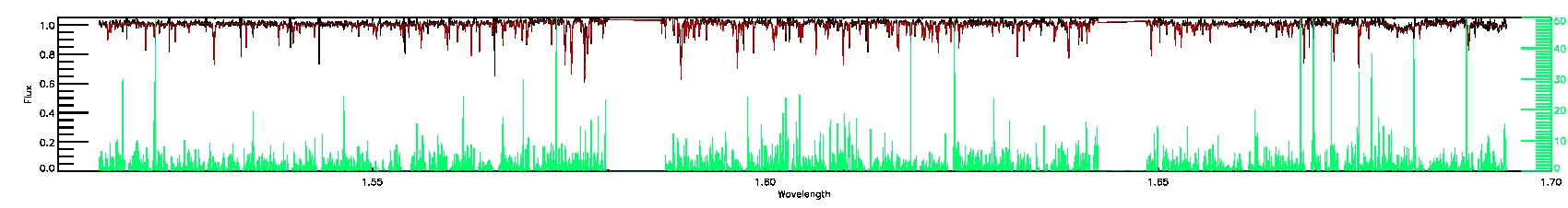
055-53-O
| 5853. | +/- | 11. |
| 5788. | +/- | 119. |
| 4.49 | +/- | 0. |
| 4.41 | +/- | 0. |
| 1.25 | +/- | 0. |
| 1.25 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.16 | +/- | 0. |
| -0.16 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 4.56 | +/- | 0. |
| 0.66 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.20 |
| -9999.00 |
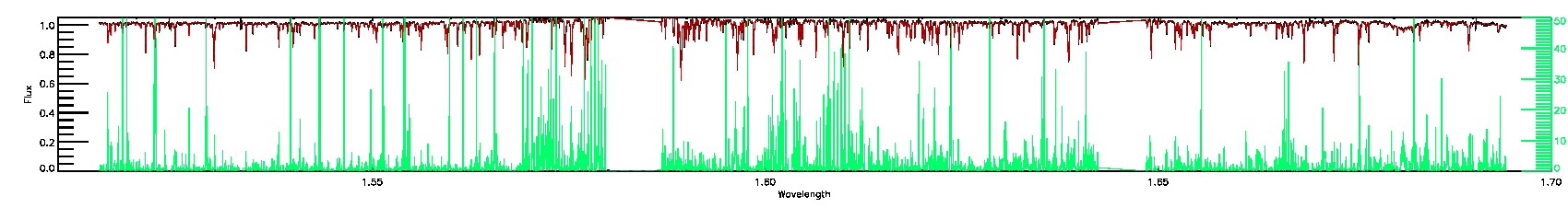
055-53-O
| 5944. | +/- | 22. |
| 5878. | +/- | 143. |
| 4.21 | +/- | 0. |
| 4.21 | +/- | 0. |
| 0.56 | +/- | 0. |
| 0.56 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -0.30 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.07 | +/- | 0. |
| 6.40 | +/- | 0. |
| 0.81 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.79 |
| -9999.00 |

055-53-O
| 4302. | +/- | 2. |
| 4410. | +/- | 78. |
| 2.17 | +/- | 0. |
| 1.99 | +/- | 0. |
| 1.49 | +/- | 0. |
| 1.49 | +/- | -NaN |
| -0.45 | +/- | 0. |
| -0.44 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.18 | +/- | 0. |
| 3.84 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.83 |
| -9999.00 |

055-53-O
| 4816. | +/- | 5. |
| 4880. | +/- | 89. |
| 2.77 | +/- | 0. |
| 2.43 | +/- | 0. |
| 1.48 | +/- | 0. |
| 1.48 | +/- | -NaN |
| -0.30 | +/- | 0. |
| -0.30 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.08 | +/- | 0. |
| 3.52 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.18 |
| -9999.00 |

BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
055-53-O
| 6571. | +/- | 26. |
| 6426. | +/- | 165. |
| 4.20 | +/- | 0. |
| 4.03 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.43 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.17 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.17 | +/- | 0. |
| 45.16 | +/- | 0. |
| 1.65 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -1.80 |
| -9999.00 |
055-53-O
| 6475. | +/- | 16. |
| 6345. | +/- | 145. |
| 4.25 | +/- | 0. |
| 4.12 | +/- | 0. |
| 1.82 | +/- | 0. |
| 1.82 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -0.24 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -0.29 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.07 | +/- | 0. |
| 11.72 | +/- | 0. |
| 1.07 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.18 |
| -9999.00 |
055-53-O
| 4133. | +/- | 3. |
| 4227. | +/- | 79. |
| 4.36 | +/- | 0. |
| 4.63 | +/- | 0. |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -0.13 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.24 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 6.11 | +/- | 0. |
| 0.79 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.79 |
| -9999.00 |

BRIGHT_NEIGHBOR
055-53-O
| 5555. | +/- | 35. |
| 5539. | +/- | 139. |
| 4.28 | +/- | 0. |
| 4.40 | +/- | 0. |
| 1.44 | +/- | 0. |
| 1.44 | +/- | -NaN |
| -0.41 | +/- | 0. |
| -0.41 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 14.76 | +/- | 0. |
| 1.17 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.63 |
| -9999.00 |

STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN
055-53-O
| 7909. | +/- | 13. |
| -10000. | +/- | -NaN |
| 4.59 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.00 | +/- | 0. |
| 2.00 | +/- | -NaN |
| 0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 84.45 | +/- | 0. |
| 1.93 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.45 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -1.91 |
| -9999.00 |
055-53-O
| 4677. | +/- | 10. |
| 4768. | +/- | 111. |
| 2.80 | +/- | 0. |
| 2.66 | +/- | 0. |
| 1.37 | +/- | 0. |
| 1.37 | +/- | -NaN |
| -0.55 | +/- | 0. |
| -0.55 | +/- | 0. |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.22 | +/- | 0. |
| 4.07 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.09 |
| -9999.00 |
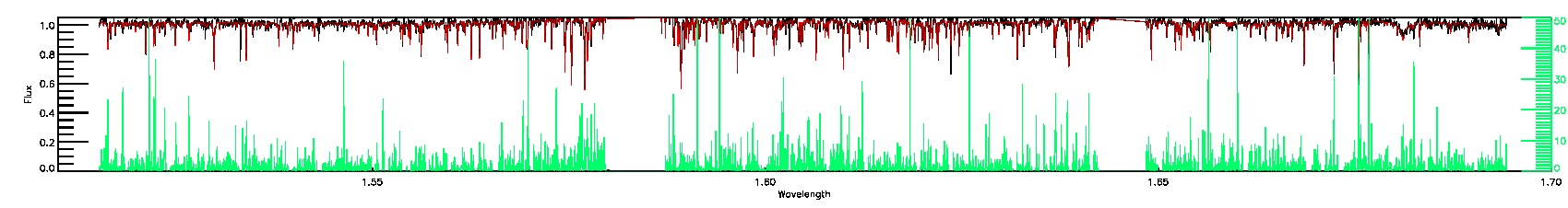
055-53-O
| 4906. | +/- | 6. |
| 4957. | +/- | 91. |
| 3.63 | +/- | 0. |
| 3.59 | +/- | 0. |
| 0.43 | +/- | 0. |
| 0.43 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.24 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.17 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.08 | +/- | 0. |
| 3.41 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.23 |
| -9999.00 |

055-53-O
| 4761. | +/- | 4. |
| 4827. | +/- | 87. |
| 2.76 | +/- | 0. |
| 2.43 | +/- | 0. |
| 1.45 | +/- | 0. |
| 1.45 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -0.18 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.28 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.16 |
| -9999.00 |

055-53-O
| 5574. | +/- | 9. |
| 5525. | +/- | 108. |
| 4.07 | +/- | 0. |
| 3.90 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 5.20 | +/- | 0. |
| 0.72 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.43 |
| -9999.00 |
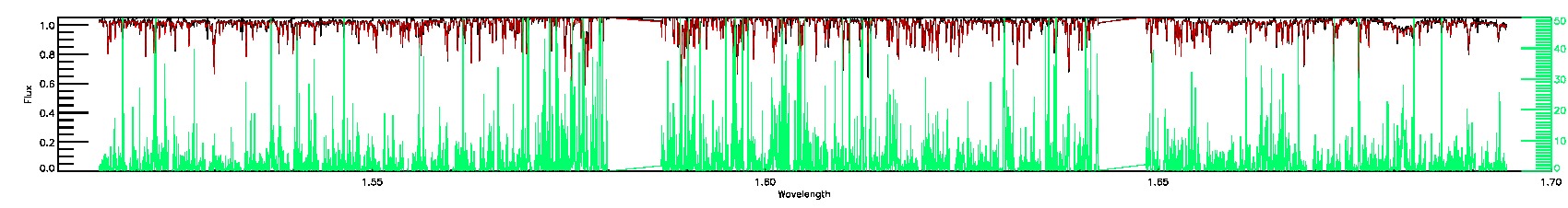
055-53-O
| 4817. | +/- | 11. |
| 4897. | +/- | 114. |
| 2.96 | +/- | 0. |
| 2.85 | +/- | 0. |
| 1.14 | +/- | 0. |
| 1.14 | +/- | -NaN |
| -0.66 | +/- | 0. |
| -0.66 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.23 | +/- | 0. |
| 4.36 | +/- | 0. |
| 0.64 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.05 |
| -9999.00 |

055-53-O
| 5755. | +/- | 20. |
| 5695. | +/- | 138. |
| 4.54 | +/- | 0. |
| 4.43 | +/- | 0. |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 6.91 | +/- | 0. |
| 0.84 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.43 |
| -9999.00 |

055-53-O
| 5058. | +/- | 14. |
| 5115. | +/- | 115. |
| 3.40 | +/- | 0. |
| 3.35 | +/- | 0. |
| 1.16 | +/- | 0. |
| 1.16 | +/- | -NaN |
| -0.79 | +/- | 0. |
| -0.79 | +/- | 0. |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -0.25 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.17 | +/- | 0. |
| 4.70 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.17 |
| -9999.00 |

055-53-O
| 5858. | +/- | 18. |
| 5792. | +/- | 138. |
| 4.57 | +/- | 0. |
| 4.48 | +/- | 0. |
| 0.53 | +/- | 0. |
| 0.53 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 4.55 | +/- | 0. |
| 0.66 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.12 |
| -9999.00 |

055-53-O
| 5401. | +/- | 9. |
| 5374. | +/- | 112. |
| 4.42 | +/- | 0. |
| 4.30 | +/- | 0. |
| 0.59 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.12 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.41 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.14 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.40 |
| -9999.00 |

055-53-O
| 5935. | +/- | 48. |
| 5881. | +/- | 167. |
| 4.39 | +/- | 0. |
| 4.49 | +/- | 0. |
| 1.06 | +/- | 0. |
| 1.06 | +/- | -NaN |
| -0.56 | +/- | 0. |
| -0.56 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.20 | +/- | 0. |
| 8.57 | +/- | 0. |
| 0.93 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.00 |
| -9999.00 |

055-53-O
| 4149. | +/- | 2. |
| 4251. | +/- | 74. |
| 4.33 | +/- | 0. |
| 4.68 | +/- | 0. |
| 0.48 | +/- | 0. |
| 0.48 | +/- | -NaN |
| -0.33 | +/- | 0. |
| -0.33 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 6.01 | +/- | 0. |
| 0.78 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.34 |
| -9999.00 |

055-53-O
| 5131. | +/- | 12. |
| 5155. | +/- | 112. |
| 4.56 | +/- | 0. |
| 4.67 | +/- | 0. |
| 1.24 | +/- | 0. |
| 1.24 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -0.20 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.12 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.04 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.17 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.60 |
| -9999.00 |

055-53-O
| 5299. | +/- | 8. |
| 5283. | +/- | 98. |
| 4.02 | +/- | 0. |
| 3.91 | +/- | 0. |
| 0.71 | +/- | 0. |
| 0.71 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.54 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 1.00 |
| -9999.00 |

055-53-O
| 4897. | +/- | 6. |
| 4962. | +/- | 94. |
| 2.71 | +/- | 0. |
| 2.39 | +/- | 0. |
| 1.74 | +/- | 0. |
| 1.74 | +/- | -NaN |
| -0.54 | +/- | 0. |
| -0.54 | +/- | 0. |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.18 | +/- | 0. |
| 4.06 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.20 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.41 |
| -9999.00 |

055-53-O
| 4883. | +/- | 12. |
| 4927. | +/- | 111. |
| 3.51 | +/- | 0. |
| 3.36 | +/- | 0. |
| 0.87 | +/- | 0. |
| 0.87 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.97 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.18 |
| -9999.00 |

055-53-O
| 5010. | +/- | 7. |
| 5039. | +/- | 92. |
| 3.68 | +/- | 0. |
| 3.61 | +/- | 0. |
| 1.09 | +/- | 0. |
| 1.09 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.97 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.04 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
055-53-O
| 3639. | +/- | 3. |
| 3737. | +/- | 71. |
| 4.40 | +/- | 0. |
| 4.80 | +/- | 0. |
| 0.70 | +/- | 0. |
| 0.70 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.23 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.10 | +/- | 0. |
| 4.76 | +/- | 0. |
| 0.68 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| 0.23 |
| -9999.00 |
055-53-O
| 5882. | +/- | 23. |
| 5821. | +/- | 145. |
| 4.33 | +/- | 0. |
| 4.32 | +/- | 0. |
| 0.57 | +/- | 0. |
| 0.57 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -0.26 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 6.87 | +/- | 0. |
| 0.84 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.57 |
| -9999.00 |
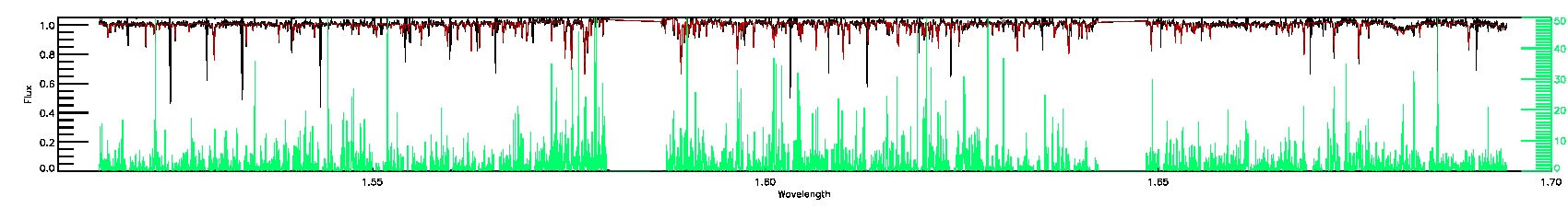
BRIGHT_NEIGHBOR
055-53-O
| 4860. | +/- | 10. |
| 4916. | +/- | 109. |
| 4.48 | +/- | 0. |
| 4.64 | +/- | 0. |
| 0.98 | +/- | 0. |
| 0.98 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 2.77 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.24 |
| -9999.00 |

BRIGHT_NEIGHBOR
055-53-O
| 4774. | +/- | 6. |
| 4851. | +/- | 93. |
| 2.69 | +/- | 0. |
| 2.38 | +/- | 0. |
| 1.58 | +/- | 0. |
| 1.58 | +/- | -NaN |
| -0.48 | +/- | 0. |
| -0.48 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.20 | +/- | 0. |
| 3.91 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.57 |
| -9999.00 |
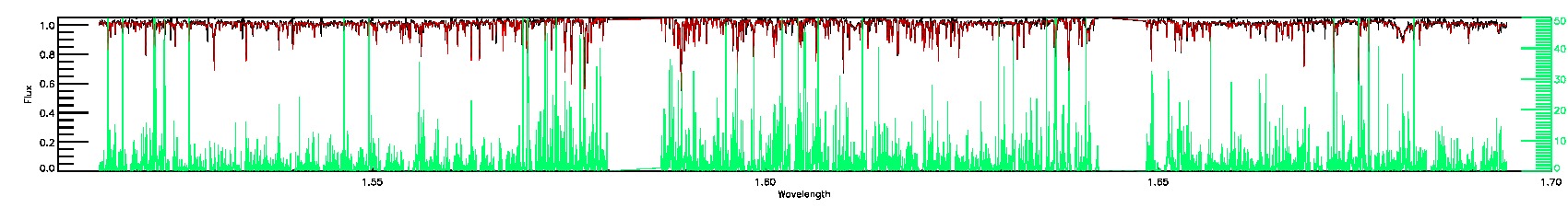
SUSPECT_BROAD_LINES
STAR_WARN,SN_WARN
055-53-O
| 5431. | +/- | 28. |
| 5410. | +/- | 138. |
| 4.46 | +/- | 0. |
| 4.42 | +/- | 0. |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.03 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 14.49 | +/- | 0. |
| 1.16 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.28 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
055-53-O
| 6410. | +/- | 19. |
| 6280. | +/- | 147. |
| 4.76 | +/- | 0. |
| 4.58 | +/- | 0. |
| 0.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 5.22 | +/- | 0. |
| 0.72 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.35 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
055-53-O
| 3446. | +/- | 3. |
| 3556. | +/- | 71. |
| 4.29 | +/- | 0. |
| 4.85 | +/- | 0. |
| 1.04 | +/- | 0. |
| 1.04 | +/- | -NaN |
| -0.52 | +/- | 0. |
| -0.52 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.09 | +/- | 0. |
| 6.94 | +/- | 0. |
| 0.84 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 1.06 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.48 |
| -9999.00 |
055-53-O
| 6117. | +/- | 39. |
| 6032. | +/- | 167. |
| 4.14 | +/- | 0. |
| 4.12 | +/- | 0. |
| 1.27 | +/- | 0. |
| 1.27 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -0.34 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.12 | +/- | 0. |
| 8.09 | +/- | 0. |
| 0.91 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -2.23 |
| -9999.00 |
055-53-O
| 5574. | +/- | 31. |
| 5547. | +/- | 143. |
| 4.01 | +/- | 0. |
| 4.05 | +/- | 0. |
| 1.00 | +/- | 0. |
| 1.00 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.23 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.07 | +/- | 0. |
| 4.16 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.47 |
| -9999.00 |

055-53-O
| 5807. | +/- | 23. |
| 5749. | +/- | 147. |
| 4.31 | +/- | 0. |
| 4.26 | +/- | 0. |
| 1.03 | +/- | 0. |
| 1.03 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.07 | +/- | 0. |
| 2.37 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.71 |
| -9999.00 |

055-53-O
| 5761. | +/- | 12. |
| 5704. | +/- | 115. |
| 4.31 | +/- | 0. |
| 4.24 | +/- | 0. |
| 1.08 | +/- | 0. |
| 1.08 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.08 | +/- | 0. |
| 3.22 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.08 |
| -9999.00 |

055-53-O
| 4584. | +/- | 6. |
| 4668. | +/- | 101. |
| 4.42 | +/- | 0. |
| 4.60 | +/- | 0. |
| 0.95 | +/- | 0. |
| 0.95 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 1.86 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.47 |
| -9999.00 |

055-53-O
| 5712. | +/- | 18. |
| 5660. | +/- | 134. |
| 4.32 | +/- | 0. |
| 4.24 | +/- | 0. |
| 0.67 | +/- | 0. |
| 0.67 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.06 | +/- | 0. |
| 3.59 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.68 |
| -9999.00 |

055-53-O
| 4848. | +/- | 5. |
| 4893. | +/- | 87. |
| 3.43 | +/- | 0. |
| 3.26 | +/- | 0. |
| 0.86 | +/- | 0. |
| 0.86 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.84 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.15 |
| -9999.00 |

055-53-O
| 5610. | +/- | 13. |
| 5573. | +/- | 118. |
| 4.47 | +/- | 0. |
| 4.44 | +/- | 0. |
| 1.04 | +/- | 0. |
| 1.04 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 3.23 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.40 |
| -9999.00 |
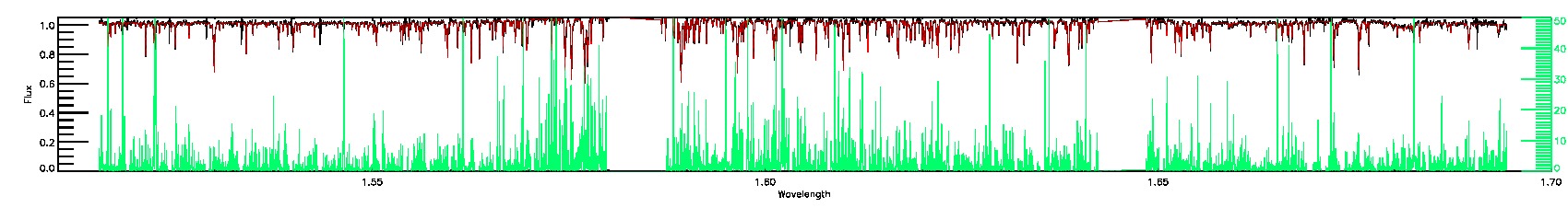
SUSPECT_BROAD_LINES
055-53-O
| 6273. | +/- | 16. |
| 6155. | +/- | 133. |
| 4.29 | +/- | 0. |
| 4.10 | +/- | 0. |
| 1.45 | +/- | 0. |
| 1.45 | +/- | -NaN |
| -0.00 | +/- | 0. |
| 0.00 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 18.68 | +/- | 0. |
| 1.27 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.00 |
| -9999.00 |
BRIGHT_NEIGHBOR
055-53-O
| 5247. | +/- | 14. |
| 5259. | +/- | 119. |
| 3.92 | +/- | 0. |
| 3.99 | +/- | 0. |
| 0.99 | +/- | 0. |
| 0.99 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.23 | +/- | 0. |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.16 | +/- | 0. |
| 3.39 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.00 |
| -9999.00 |

055-53-O
| 5832. | +/- | 10. |
| 5766. | +/- | 117. |
| 4.30 | +/- | 0. |
| 4.20 | +/- | 0. |
| 1.41 | +/- | 0. |
| 1.41 | +/- | -NaN |
| -0.00 | +/- | 0. |
| 0.00 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 3.34 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.98 |
| -9999.00 |

055-53-O
| 5147. | +/- | 10. |
| 5162. | +/- | 108. |
| 4.54 | +/- | 0. |
| 4.59 | +/- | 0. |
| 1.26 | +/- | 0. |
| 1.26 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 1.87 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.38 |
| -9999.00 |

055-53-O
| 5856. | +/- | 13. |
| 5792. | +/- | 119. |
| 4.46 | +/- | 0. |
| 4.40 | +/- | 0. |
| 0.74 | +/- | 0. |
| 0.74 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.13 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.11 | +/- | 0. |
| 1.88 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.14 |
| -9999.00 |
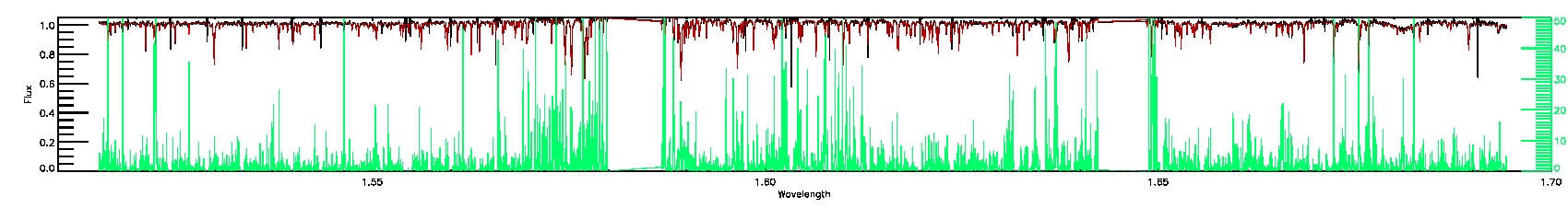
BRIGHT_NEIGHBOR
055-53-O
| 4905. | +/- | 13. |
| 4959. | +/- | 117. |
| 3.70 | +/- | 0. |
| 3.72 | +/- | 0. |
| 0.63 | +/- | 0. |
| 0.63 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -0.29 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.19 | +/- | 0. |
| 1.53 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.14 |
| -9999.00 |

055-53-O
| 5433. | +/- | 9. |
| 5415. | +/- | 104. |
| 4.65 | +/- | 0. |
| 4.64 | +/- | 0. |
| 1.50 | +/- | 0. |
| 1.50 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.05 | +/- | 0. |
| 5.05 | +/- | 0. |
| 0.70 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |

055-53-O
| 5792. | +/- | 8. |
| 5723. | +/- | 113. |
| 4.48 | +/- | 0. |
| 4.32 | +/- | 0. |
| 0.93 | +/- | 0. |
| 0.93 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.46 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.28 |
| -9999.00 |
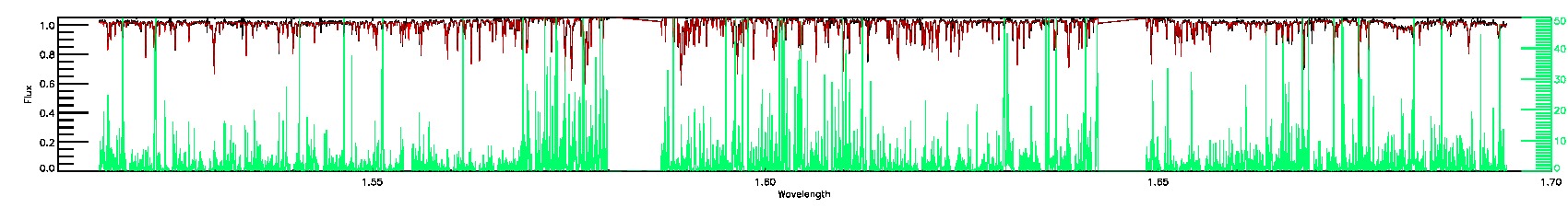
055-53-O
| 5359. | +/- | 12. |
| 5340. | +/- | 121. |
| 4.18 | +/- | 0. |
| 4.09 | +/- | 0. |
| 0.73 | +/- | 0. |
| 0.73 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 4.72 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.37 |
| -9999.00 |

BRIGHT_NEIGHBOR
055-53-O
| 5512. | +/- | 10. |
| 5482. | +/- | 108. |
| 4.39 | +/- | 0. |
| 4.33 | +/- | 0. |
| 1.05 | +/- | 0. |
| 1.05 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.69 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.20 |
| -9999.00 |
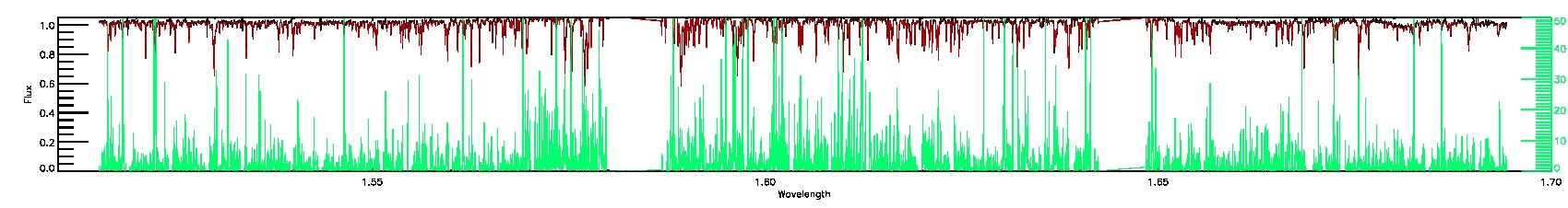
055-53-O
| 4191. | +/- | 1. |
| 4313. | +/- | 79. |
| 1.81 | +/- | 0. |
| 1.69 | +/- | 0. |
| 1.69 | +/- | 0. |
| 1.69 | +/- | -NaN |
| -0.77 | +/- | 0. |
| -0.77 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.19 | +/- | 0. |
| 4.64 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -1.12 |
| -9999.00 |
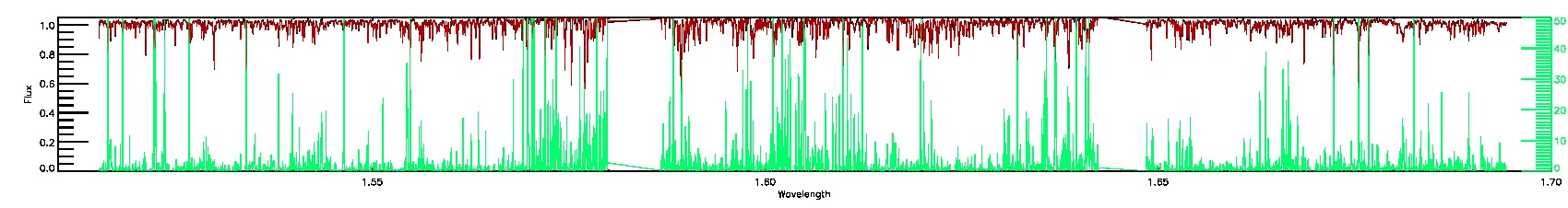
055-53-O
| 4650. | +/- | 7. |
| 4732. | +/- | 103. |
| 2.69 | +/- | 0. |
| 2.38 | +/- | 0. |
| 1.32 | +/- | 0. |
| 1.32 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -0.26 | +/- | 0. |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.20 | +/- | 0. |
| 3.43 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -1.06 |
| -9999.00 |

055-53-O
| 4610. | +/- | 9. |
| 4682. | +/- | 106. |
| 4.50 | +/- | 0. |
| 4.59 | +/- | 0. |
| 0.62 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.87 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 1.00 |
| -9999.00 |

055-53-O
| 3881. | +/- | 1. |
| 3984. | +/- | 68. |
| 1.23 | +/- | 0. |
| 1.12 | +/- | 0. |
| 1.66 | +/- | 0. |
| 1.66 | +/- | -NaN |
| -0.33 | +/- | 0. |
| -0.33 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.09 | +/- | 0. |
| 3.60 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.22 |
| -9999.00 |
055-53-O
| 4497. | +/- | 5. |
| 4583. | +/- | 97. |
| 4.47 | +/- | 0. |
| 4.60 | +/- | 0. |
| 0.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 4.30 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.02 |
| -9999.00 |

055-53-O
| 5910. | +/- | 20. |
| 5830. | +/- | 148. |
| 4.42 | +/- | 0. |
| 4.25 | +/- | 0. |
| 1.03 | +/- | 0. |
| 1.03 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 3.40 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| 0.10 |
| -9999.00 |
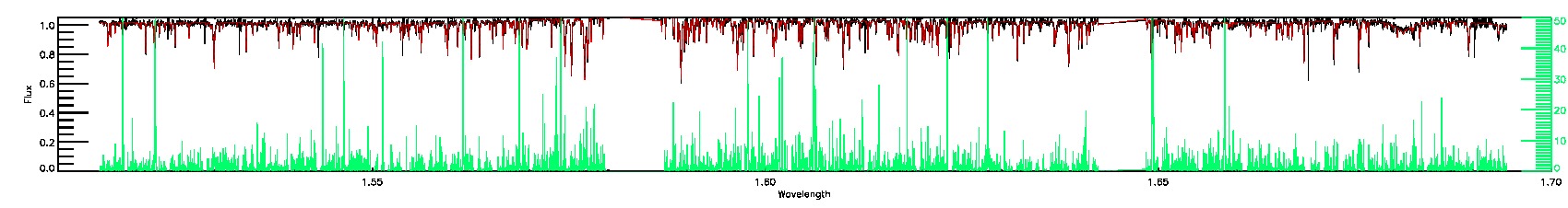
STAR_WARN,SN_WARN
055-53-O
| 4420. | +/- | 7. |
| 4516. | +/- | 105. |
| 4.34 | +/- | 0. |
| 4.58 | +/- | 0. |
| 1.01 | +/- | 0. |
| 1.01 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -0.18 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 5.72 | +/- | 0. |
| 0.76 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.27 |
| -9999.00 |

055-53-O
| 5925. | +/- | 10. |
| 5842. | +/- | 118. |
| 4.58 | +/- | 0. |
| 4.40 | +/- | 0. |
| 0.51 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 9.94 | +/- | 0. |
| 1.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.20 |
| -9999.00 |
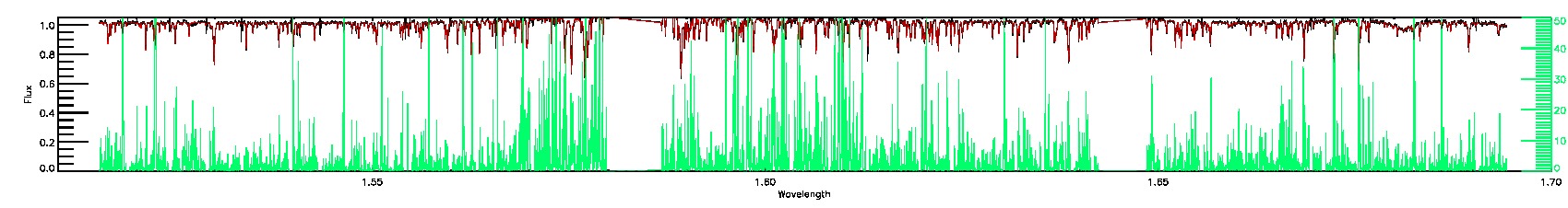
055-53-O
| 4655. | +/- | 5. |
| 4732. | +/- | 94. |
| 4.41 | +/- | 0. |
| 4.59 | +/- | 0. |
| 1.01 | +/- | 0. |
| 1.01 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.08 | +/- | 0. |
| 1.86 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_WARN,SN_WARN
055-53-O
| 4480. | +/- | 9. |
| 4562. | +/- | 103. |
| 4.55 | +/- | 0. |
| 4.64 | +/- | 0. |
| 0.76 | +/- | 0. |
| 0.76 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 17.36 | +/- | 0. |
| 1.24 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.87 |
| -9999.00 |
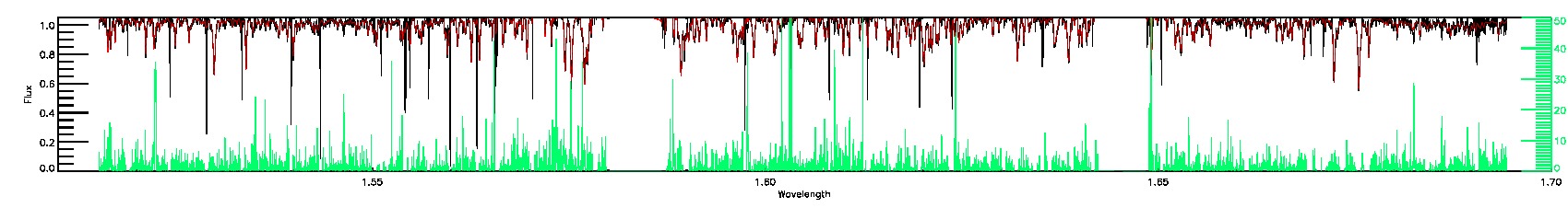
STAR_BAD
055-53-O
| 5940. | +/- | 12. |
| -10000. | +/- | -NaN |
| 3.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.16 | +/- | 0. |
| 1.16 | +/- | -NaN |
| -0.34 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.98 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.32 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -1.07 |
| -9999.00 |
055-53-O
| 5315. | +/- | 11. |
| 5307. | +/- | 115. |
| 4.52 | +/- | 0. |
| 4.50 | +/- | 0. |
| 1.02 | +/- | 0. |
| 1.02 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 1.67 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.13 |
| -9999.00 |

055-53-O
| 6039. | +/- | 31. |
| 5950. | +/- | 157. |
| 4.60 | +/- | 0. |
| 4.47 | +/- | 0. |
| 1.10 | +/- | 0. |
| 1.10 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 4.53 | +/- | 0. |
| 0.66 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.16 |
| -9999.00 |
055-53-O
| 5563. | +/- | 17. |
| 5526. | +/- | 131. |
| 4.46 | +/- | 0. |
| 4.39 | +/- | 0. |
| 0.64 | +/- | 0. |
| 0.64 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.06 | +/- | 0. |
| -0.18 | +/- | 0. |
| -0.18 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.67 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.14 |
| -9999.00 |

055-53-O
| 5117. | +/- | 11. |
| 5164. | +/- | 111. |
| 2.82 | +/- | 0. |
| 2.47 | +/- | 0. |
| 1.97 | +/- | 0. |
| 1.97 | +/- | -NaN |
| -0.70 | +/- | 0. |
| -0.70 | +/- | 0. |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.24 | +/- | 0. |
| 4.46 | +/- | 0. |
| 0.65 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| 0.42 |
| -9999.00 |

055-53-O
| 5096. | +/- | 19. |
| 5119. | +/- | 125. |
| 4.50 | +/- | 0. |
| 4.56 | +/- | 0. |
| 0.48 | +/- | 0. |
| 0.48 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.06 | +/- | 0. |
| 1.69 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.44 |
| -9999.00 |

055-53-O
| 5697. | +/- | 17. |
| 5663. | +/- | 117. |
| 4.58 | +/- | 0. |
| 4.66 | +/- | 0. |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| -0.38 | +/- | 0. |
| -0.37 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.13 | +/- | 0. |
| 10.20 | +/- | 0. |
| 1.01 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.75 |
| -9999.00 |

SUSPECT_BROAD_LINES
055-53-O
| 4344. | +/- | 2. |
| 4435. | +/- | 76. |
| 4.24 | +/- | 0. |
| 4.44 | +/- | 0. |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.03 | +/- | 0. |
| 14.81 | +/- | 0. |
| 1.17 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |

BRIGHT_NEIGHBOR
055-53-O
| 4430. | +/- | 4. |
| 4518. | +/- | 92. |
| 4.45 | +/- | 0. |
| 4.61 | +/- | 0. |
| 1.25 | +/- | 0. |
| 1.25 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.01 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 5.66 | +/- | 0. |
| 0.75 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.07 |
| -9999.00 |

055-53-O
| 5943. | +/- | 18. |
| 5872. | +/- | 134. |
| 4.41 | +/- | 0. |
| 4.37 | +/- | 0. |
| 0.77 | +/- | 0. |
| 0.77 | +/- | -NaN |
| -0.21 | +/- | 0. |
| -0.21 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.17 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 6.78 | +/- | 0. |
| 0.83 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.23 |
| -9999.00 |

055-53-O
| 5308. | +/- | 9. |
| 5308. | +/- | 101. |
| 4.41 | +/- | 0. |
| 4.46 | +/- | 0. |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.12 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 4.28 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.00 |
| -9999.00 |

055-53-O
| 6011. | +/- | 24. |
| 5931. | +/- | 144. |
| 4.38 | +/- | 0. |
| 4.31 | +/- | 0. |
| 0.92 | +/- | 0. |
| 0.92 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.17 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 2.69 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.11 |
| -9999.00 |
STAR_WARN,SN_WARN
055-53-O
| 4230. | +/- | 7. |
| 4335. | +/- | 101. |
| 4.31 | +/- | 0. |
| 4.67 | +/- | 0. |
| 0.80 | +/- | 0. |
| 0.80 | +/- | -NaN |
| -0.39 | +/- | 0. |
| -0.39 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 6.23 | +/- | 0. |
| 0.79 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -1.34 |
| -9999.00 |

STAR_WARN,SN_WARN
055-53-O
| 4908. | +/- | 23. |
| 4977. | +/- | 130. |
| 3.12 | +/- | 0. |
| 3.02 | +/- | 0. |
| 1.20 | +/- | 0. |
| 1.20 | +/- | -NaN |
| -0.67 | +/- | 0. |
| -0.67 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.30 | +/- | 0. |
| 4.37 | +/- | 0. |
| 0.64 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -1.55 |
| -9999.00 |
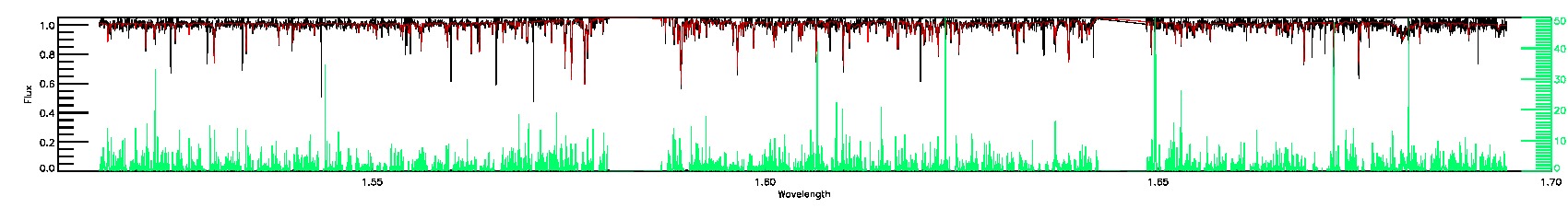
055-53-O
| 4712. | +/- | 7. |
| 4791. | +/- | 96. |
| 2.82 | +/- | 0. |
| 2.66 | +/- | 0. |
| 1.47 | +/- | 0. |
| 1.47 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -0.36 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.65 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.58 |
| -9999.00 |

055-53-O
| 4953. | +/- | 6. |
| 4989. | +/- | 90. |
| 3.46 | +/- | 0. |
| 3.30 | +/- | 0. |
| 1.26 | +/- | 0. |
| 1.26 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.96 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.19 |
| -9999.00 |

055-53-O
| 4069. | +/- | 5. |
| 4161. | +/- | 89. |
| 4.44 | +/- | 0. |
| 4.70 | +/- | 0. |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 4.54 | +/- | 0. |
| 0.66 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.12 |
| -9999.00 |

BRIGHT_NEIGHBOR
055-53-O
| 4645. | +/- | 4. |
| 4729. | +/- | 87. |
| 2.93 | +/- | 0. |
| 2.76 | +/- | 0. |
| 1.16 | +/- | 0. |
| 1.16 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -0.28 | +/- | 0. |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.17 | +/- | 0. |
| 3.49 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.25 |
| -9999.00 |

055-53-O
| 5643. | +/- | 35. |
| 5605. | +/- | 149. |
| 4.59 | +/- | 0. |
| 4.58 | +/- | 0. |
| 0.74 | +/- | 0. |
| 0.74 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -0.14 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 4.31 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| 0.98 |
| -9999.00 |

055-53-O
| 5730. | +/- | 25. |
| 5672. | +/- | 142. |
| 4.24 | +/- | 0. |
| 4.12 | +/- | 0. |
| 1.27 | +/- | 0. |
| 1.27 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.10 | +/- | 0. |
| -0.25 | +/- | 0. |
| -0.25 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 5.10 | +/- | 0. |
| 0.71 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.28 |
| -9999.00 |
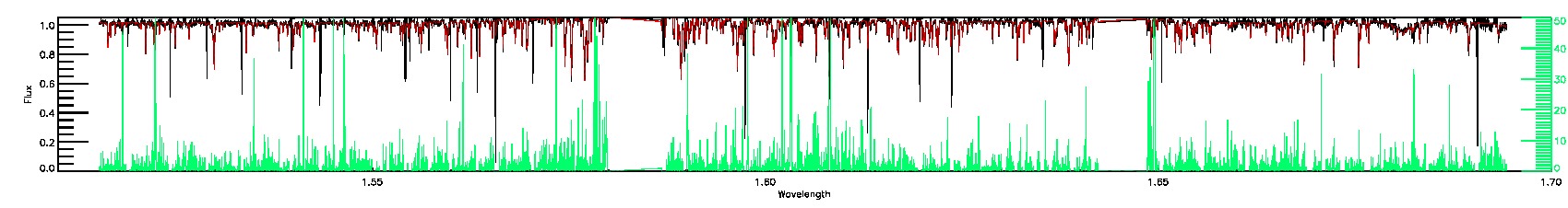
055-53-O
| 5411. | +/- | 14. |
| 5386. | +/- | 127. |
| 4.58 | +/- | 0. |
| 4.48 | +/- | 0. |
| 0.54 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 1.52 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.24 |
| -9999.00 |

055-53-O
| 4825. | +/- | 5. |
| 4888. | +/- | 90. |
| 2.81 | +/- | 0. |
| 2.46 | +/- | 0. |
| 1.51 | +/- | 0. |
| 1.51 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -0.29 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.10 | +/- | 0. |
| 3.50 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.52 |
| -9999.00 |

055-53-O
| 5855. | +/- | 11. |
| 5786. | +/- | 118. |
| 4.51 | +/- | 0. |
| 4.40 | +/- | 0. |
| 1.12 | +/- | 0. |
| 1.12 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.00 | +/- | 0. |
| 3.08 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.66 |
| -9999.00 |

055-53-O
| 6531. | +/- | 18. |
| 6389. | +/- | 146. |
| 4.66 | +/- | 0. |
| 4.47 | +/- | 0. |
| 1.14 | +/- | 0. |
| 1.14 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.14 | +/- | 0. |
| -0.14 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 9.05 | +/- | 0. |
| 0.96 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
STAR_WARN,SN_WARN
055-53-O
| 5748. | +/- | 58. |
| 5714. | +/- | 163. |
| 4.29 | +/- | 0. |
| 4.41 | +/- | 0. |
| 0.74 | +/- | 0. |
| 0.74 | +/- | -NaN |
| -0.53 | +/- | 0. |
| -0.53 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.25 | +/- | 0. |
| 5.27 | +/- | 0. |
| 0.72 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -0.61 |
| -9999.00 |

055-53-O
| 5529. | +/- | 32. |
| 5507. | +/- | 144. |
| 4.51 | +/- | 0. |
| 4.54 | +/- | 0. |
| 1.34 | +/- | 0. |
| 1.34 | +/- | -NaN |
| -0.21 | +/- | 0. |
| -0.21 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 2.33 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| 0.01 |
| -9999.00 |

055-53-O
| 5671. | +/- | 11. |
| 5625. | +/- | 113. |
| 4.48 | +/- | 0. |
| 4.41 | +/- | 0. |
| 0.85 | +/- | 0. |
| 0.85 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.05 | +/- | 0. |
| 1.87 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.17 |
| -9999.00 |

055-53-O
| 5868. | +/- | 20. |
| 5820. | +/- | 132. |
| 3.99 | +/- | 0. |
| 4.09 | +/- | 0. |
| 1.18 | +/- | 0. |
| 1.18 | +/- | -NaN |
| -0.52 | +/- | 0. |
| -0.52 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 4.01 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.04 |
| -9999.00 |

055-53-O
| 4816. | +/- | 7. |
| 4861. | +/- | 96. |
| 4.59 | +/- | 0. |
| 4.62 | +/- | 0. |
| 0.71 | +/- | 0. |
| 0.71 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.07 | +/- | 0. |
| 1.78 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.39 |
| -9999.00 |

055-53-O
| 4296. | +/- | 1. |
| 4376. | +/- | 72. |
| 2.41 | +/- | 0. |
| 2.15 | +/- | 0. |
| 1.39 | +/- | 0. |
| 1.39 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.65 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.09 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
055-53-O
| 3597. | +/- | 3. |
| 3711. | +/- | 65. |
| 4.37 | +/- | 0. |
| 4.93 | +/- | 0. |
| 0.94 | +/- | 0. |
| 0.94 | +/- | -NaN |
| -0.60 | +/- | 0. |
| -0.60 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 6.05 | +/- | 0. |
| 0.78 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.55 |
| -9999.00 |
055-53-O
| 4752. | +/- | 4. |
| 4804. | +/- | 83. |
| 3.45 | +/- | 0. |
| 3.26 | +/- | 0. |
| 0.48 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.66 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.26 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
055-53-O
| 3305. | +/- | 3. |
| 3410. | +/- | 64. |
| 4.56 | +/- | 0. |
| 5.09 | +/- | 0. |
| 2.21 | +/- | 0. |
| 2.21 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -0.39 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.05 | +/- | 0. |
| 10.26 | +/- | 0. |
| 1.01 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.35 |
| -9999.00 |
055-53-O
| 4872. | +/- | 6. |
| 4929. | +/- | 91. |
| 3.13 | +/- | 0. |
| 2.98 | +/- | 0. |
| 1.31 | +/- | 0. |
| 1.31 | +/- | -NaN |
| -0.27 | +/- | 0. |
| -0.27 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.46 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.39 |
| -9999.00 |

055-53-O
| 5814. | +/- | 34. |
| 5761. | +/- | 152. |
| 4.19 | +/- | 0. |
| 4.20 | +/- | 0. |
| 0.67 | +/- | 0. |
| 0.67 | +/- | -NaN |
| -0.27 | +/- | 0. |
| -0.26 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.09 | +/- | 0. |
| 3.45 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -2.37 |
| -9999.00 |

055-53-O
| 5452. | +/- | 19. |
| 5440. | +/- | 124. |
| 4.24 | +/- | 0. |
| 4.30 | +/- | 0. |
| 0.92 | +/- | 0. |
| 0.92 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.13 | +/- | 0. |
| -0.13 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 5.51 | +/- | 0. |
| 0.74 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.99 |
| -9999.00 |
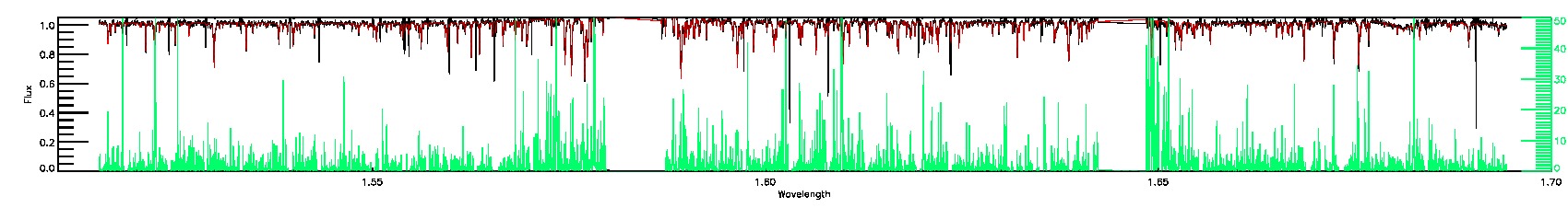
STAR_WARN,COLORTE_WARN
055-53-O
| 4678. | +/- | 5. |
| 4769. | +/- | 88. |
| 3.60 | +/- | 0. |
| 3.61 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| -0.53 | +/- | 0. |
| -0.53 | +/- | 0. |
| -0.20 | +/- | 0. |
| -0.20 | +/- | -NaN |
| -0.30 | +/- | 0. |
| -0.30 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.02 | +/- | 0. |
| 4.04 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.23 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.05 |
| -9999.00 |

055-53-O
| 4479. | +/- | 3. |
| 4588. | +/- | 83. |
| 2.49 | +/- | 0. |
| 2.33 | +/- | 0. |
| 1.40 | +/- | 0. |
| 1.40 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -0.49 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.20 | +/- | 0. |
| 3.94 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.63 |
| -9999.00 |
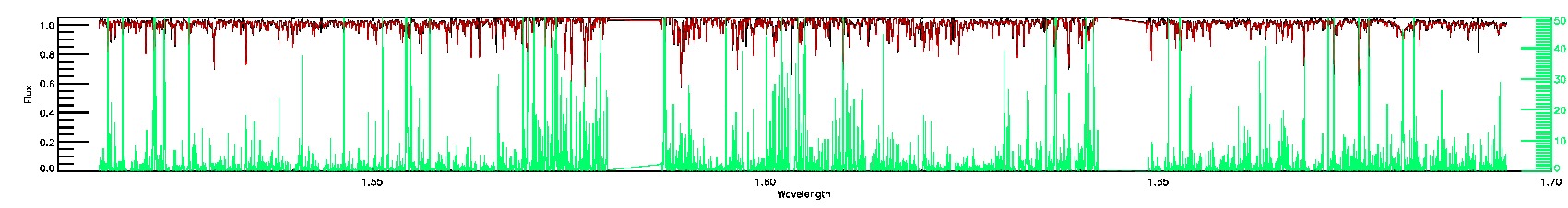
055-53-O
| 4598. | +/- | 8. |
| 4690. | +/- | 109. |
| 2.68 | +/- | 0. |
| 2.51 | +/- | 0. |
| 1.22 | +/- | 0. |
| 1.22 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -0.35 | +/- | 0. |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.23 | +/- | 0. |
| 3.64 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.45 |
| -9999.00 |

055-53-O
| 5405. | +/- | 24. |
| 5391. | +/- | 135. |
| 4.48 | +/- | 0. |
| 4.48 | +/- | 0. |
| 1.31 | +/- | 0. |
| 1.31 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.27 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.01 |
| -9999.00 |
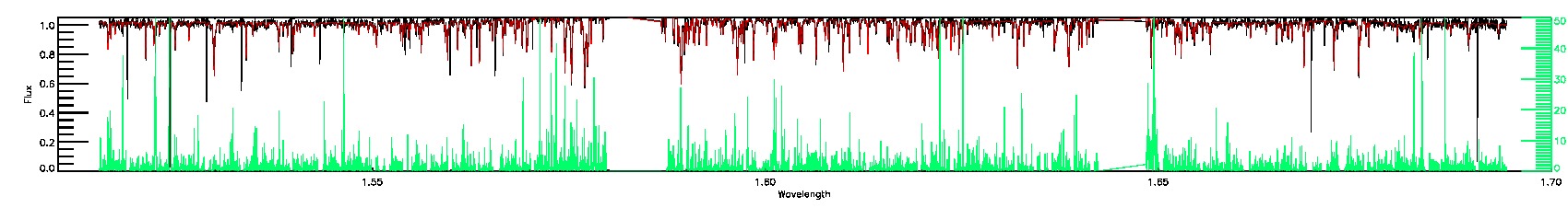
BRIGHT_NEIGHBOR
055-53-O
| 5468. | +/- | 21. |
| 5448. | +/- | 131. |
| 4.49 | +/- | 0. |
| 4.49 | +/- | 0. |
| 0.57 | +/- | 0. |
| 0.57 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.00 | +/- | 0. |
| 6.10 | +/- | 0. |
| 0.79 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 1.00 |
| -9999.00 |

055-53-O
| 5521. | +/- | 13. |
| 5482. | +/- | 124. |
| 4.35 | +/- | 0. |
| 4.23 | +/- | 0. |
| 0.48 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.20 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 3.42 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.19 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
055-53-O
| 3590. | +/- | 5. |
| 3702. | +/- | 81. |
| 4.27 | +/- | 0. |
| 4.80 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| -0.53 | +/- | 0. |
| -0.53 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 4.30 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.24 |
| -9999.00 |
055-53-O
| 4415. | +/- | 3. |
| 4504. | +/- | 88. |
| 4.41 | +/- | 0. |
| 4.58 | +/- | 0. |
| 0.47 | +/- | 0. |
| 0.47 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.41 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |

BRIGHT_NEIGHBOR
055-53-O
| 6052. | +/- | 19. |
| 5970. | +/- | 148. |
| 4.66 | +/- | 0. |
| 4.61 | +/- | 0. |
| 0.76 | +/- | 0. |
| 0.76 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.24 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.11 | +/- | 0. |
| 1.76 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.27 |
| -9999.00 |
055-53-O
| 5592. | +/- | 23. |
| 5563. | +/- | 139. |
| 4.48 | +/- | 0. |
| 4.51 | +/- | 0. |
| 0.92 | +/- | 0. |
| 0.92 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -0.21 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.14 | +/- | 0. |
| 1.54 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.38 |
| -9999.00 |

BRIGHT_NEIGHBOR
055-53-O
| 5916. | +/- | 15. |
| 5837. | +/- | 134. |
| 4.34 | +/- | 0. |
| 4.19 | +/- | 0. |
| 0.47 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.08 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 5.89 | +/- | 0. |
| 0.77 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.42 |
| -9999.00 |
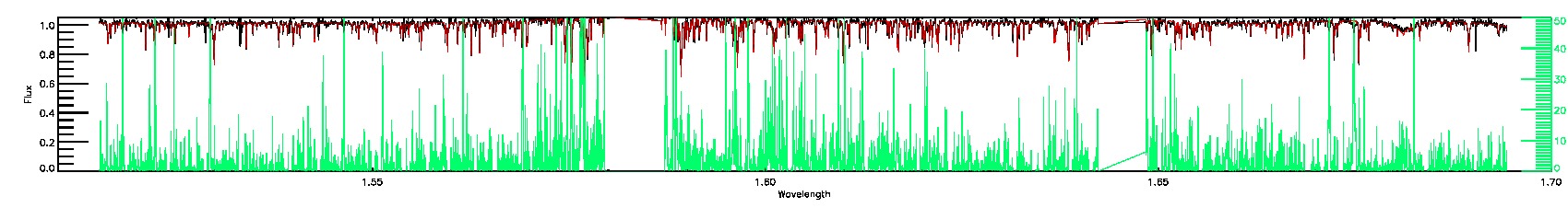
BRIGHT_NEIGHBOR
055-53-O
| 6337. | +/- | 21. |
| 6213. | +/- | 144. |
| 4.39 | +/- | 0. |
| 4.20 | +/- | 0. |
| 0.75 | +/- | 0. |
| 0.75 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.12 | +/- | -NaN |
| 0.49 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 11.61 | +/- | 0. |
| 1.06 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.14 |
| -9999.00 |
055-53-O
| 5597. | +/- | 11. |
| 5558. | +/- | 111. |
| 4.05 | +/- | 0. |
| 3.98 | +/- | 0. |
| 1.36 | +/- | 0. |
| 1.36 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.02 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.94 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.28 |
| -9999.00 |

055-53-O
| 4538. | +/- | 4. |
| 4638. | +/- | 95. |
| 2.72 | +/- | 0. |
| 2.55 | +/- | 0. |
| 1.24 | +/- | 0. |
| 1.24 | +/- | -NaN |
| -0.38 | +/- | 0. |
| -0.38 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.17 | +/- | 0. |
| 3.69 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.98 |
| -9999.00 |

055-53-O
| 5732. | +/- | 30. |
| 5680. | +/- | 146. |
| 4.23 | +/- | 0. |
| 4.16 | +/- | 0. |
| 0.85 | +/- | 0. |
| 0.85 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.23 | +/- | 0. |
| -0.23 | +/- | -NaN |
| 0.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.10 | +/- | 0. |
| 4.65 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| 0.84 |
| -9999.00 |

055-53-O
| 4976. | +/- | 8. |
| 5014. | +/- | 100. |
| 3.85 | +/- | 0. |
| 3.88 | +/- | 0. |
| 0.95 | +/- | 0. |
| 0.95 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | 0. |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.16 | +/- | 0. |
| 3.18 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.15 |
| -9999.00 |

055-53-O
| 4928. | +/- | 15. |
| 4969. | +/- | 117. |
| 4.54 | +/- | 0. |
| 4.63 | +/- | 0. |
| 1.42 | +/- | 0. |
| 1.42 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.11 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| -0.10 |
| -9999.00 |

055-53-O
| 5599. | +/- | 19. |
| 5555. | +/- | 132. |
| 4.48 | +/- | 0. |
| 4.37 | +/- | 0. |
| 0.63 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.28 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.03 |
| -9999.00 |

SUSPECT_BROAD_LINES
055-53-O
| 7011. | +/- | 17. |
| 6812. | +/- | 167. |
| 4.78 | +/- | 0. |
| 4.48 | +/- | 0. |
| 1.27 | +/- | 0. |
| 1.27 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -0.16 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 45.28 | +/- | 0. |
| 1.66 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -1.06 |
| -9999.00 |
055-53-O
| 4427. | +/- | 5. |
| 4522. | +/- | 103. |
| 4.35 | +/- | 0. |
| 4.57 | +/- | 0. |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.14 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.87 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.01 |
| -9999.00 |

055-53-O
| 5026. | +/- | 13. |
| 5055. | +/- | 113. |
| 3.96 | +/- | 0. |
| 4.00 | +/- | 0. |
| 0.64 | +/- | 0. |
| 0.64 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | 0. |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.07 | +/- | 0. |
| 3.04 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.99 |
| -9999.00 |

055-53-O
| 4800. | +/- | 7. |
| 4887. | +/- | 98. |
| 2.84 | +/- | 0. |
| 2.74 | +/- | 0. |
| 1.42 | +/- | 0. |
| 1.42 | +/- | -NaN |
| -0.78 | +/- | 0. |
| -0.78 | +/- | 0. |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.21 | +/- | 0. |
| 4.65 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -1.14 |
| -9999.00 |

STAR_BAD
055-53-O
| 4966. | +/- | 12. |
| -10000. | +/- | -NaN |
| 4.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.39 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.30 | +/- | 0. |
| 0.86 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.41 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.66 |
| -9999.00 |

055-53-O
| 5573. | +/- | 17. |
| 5533. | +/- | 133. |
| 4.55 | +/- | 0. |
| 4.45 | +/- | 0. |
| 1.05 | +/- | 0. |
| 1.05 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 1.57 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.44 |
| -9999.00 |

055-53-O
| 5517. | +/- | 27. |
| 5488. | +/- | 140. |
| 4.38 | +/- | 0. |
| 4.34 | +/- | 0. |
| 0.41 | +/- | 0. |
| 0.41 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.05 | +/- | 0. |
| 3.19 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.48 |
| -9999.00 |

055-53-O
| 3816. | +/- | 1. |
| 3969. | +/- | 88. |
| 0.34 | +/- | 0. |
| 0.39 | +/- | 0. |
| 2.60 | +/- | 0. |
| 2.60 | +/- | -NaN |
| -1.51 | +/- | 0. |
| -1.51 | +/- | 0. |
| -0.55 | +/- | 0. |
| -0.55 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.11 | +/- | 0. |
| 7.15 | +/- | 0. |
| 0.85 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.53 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -0.61 |
| -9999.00 |
055-53-O
| 6512. | +/- | 18. |
| 6376. | +/- | 147. |
| 4.40 | +/- | 0. |
| 4.26 | +/- | 0. |
| 1.99 | +/- | 0. |
| 1.99 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -0.22 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 11.41 | +/- | 0. |
| 1.06 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -0.69 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_WARN,SN_WARN
055-53-O
| 5506. | +/- | 26. |
| 5470. | +/- | 141. |
| 4.15 | +/- | 0. |
| 4.03 | +/- | 0. |
| 0.86 | +/- | 0. |
| 0.86 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.00 | +/- | 0. |
| 3.46 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.31 |
| -9999.00 |
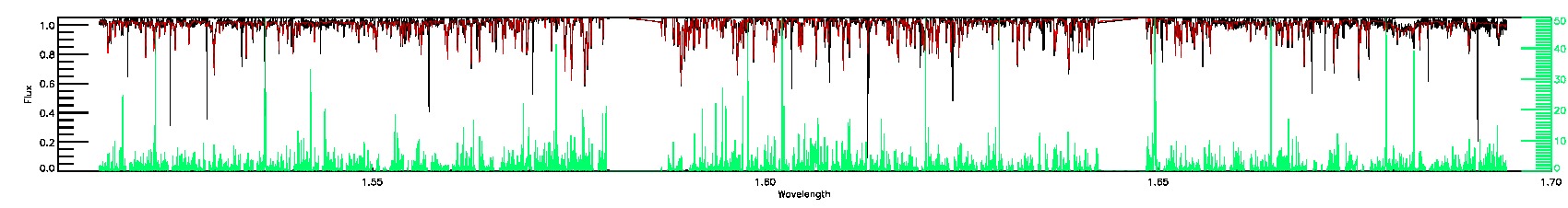
055-53-O
| 5046. | +/- | 7. |
| 5058. | +/- | 95. |
| 3.95 | +/- | 0. |
| 3.87 | +/- | 0. |
| 0.41 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.04 | +/- | 0. |
| 2.49 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.42 |
| -9999.00 |

055-53-O
| 6061. | +/- | 19. |
| 5983. | +/- | 134. |
| 4.30 | +/- | 0. |
| 4.29 | +/- | 0. |
| 0.81 | +/- | 0. |
| 0.81 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -0.35 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 5.35 | +/- | 0. |
| 0.73 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.30 |
| -9999.00 |
055-53-O
| 4660. | +/- | 12. |
| 4759. | +/- | 117. |
| 2.75 | +/- | 0. |
| 2.63 | +/- | 0. |
| 1.32 | +/- | 0. |
| 1.32 | +/- | -NaN |
| -0.68 | +/- | 0. |
| -0.68 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.26 | +/- | 0. |
| 4.41 | +/- | 0. |
| 0.64 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -0.85 |
| -9999.00 |
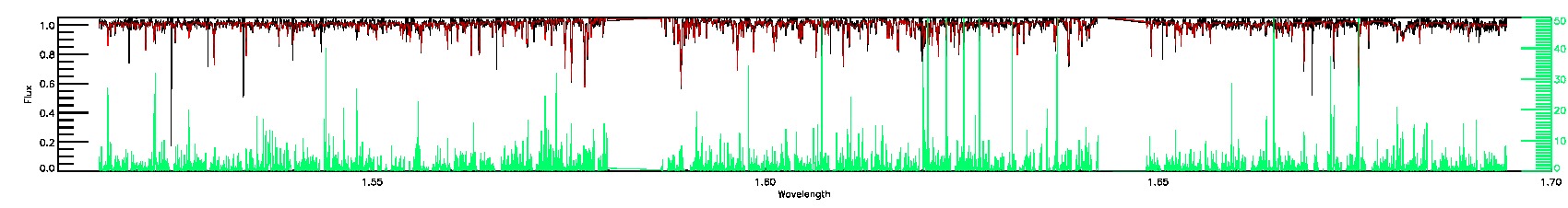
BRIGHT_NEIGHBOR
055-53-O
| 4775. | +/- | 10. |
| 4826. | +/- | 107. |
| 4.54 | +/- | 0. |
| 4.58 | +/- | 0. |
| 0.70 | +/- | 0. |
| 0.70 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.00 | +/- | 0. |
| 1.75 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.26 |
| -9999.00 |

BRIGHT_NEIGHBOR
055-53-O
| 4972. | +/- | 16. |
| 5009. | +/- | 119. |
| 4.28 | +/- | 0. |
| 4.38 | +/- | 0. |
| 0.66 | +/- | 0. |
| 0.66 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 7.07 | +/- | 0. |
| 0.85 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -0.14 |
| -9999.00 |

055-53-O
| 6074. | +/- | 27. |
| 5977. | +/- | 155. |
| 4.47 | +/- | 0. |
| 4.30 | +/- | 0. |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.12 | +/- | -NaN |
| 0.43 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 4.63 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| -0.19 |
| -9999.00 |
055-53-O
| 4855. | +/- | 5. |
| 4922. | +/- | 92. |
| 2.80 | +/- | 0. |
| 2.45 | +/- | 0. |
| 1.62 | +/- | 0. |
| 1.62 | +/- | -NaN |
| -0.47 | +/- | 0. |
| -0.47 | +/- | 0. |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.19 | +/- | 0. |
| 3.90 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.21 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.84 |
| -9999.00 |

STAR_WARN,SN_WARN
055-53-O
| 5154. | +/- | 18. |
| 5158. | +/- | 125. |
| 4.01 | +/- | 0. |
| 3.96 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | 0. |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.65 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.08 |
| -9999.00 |
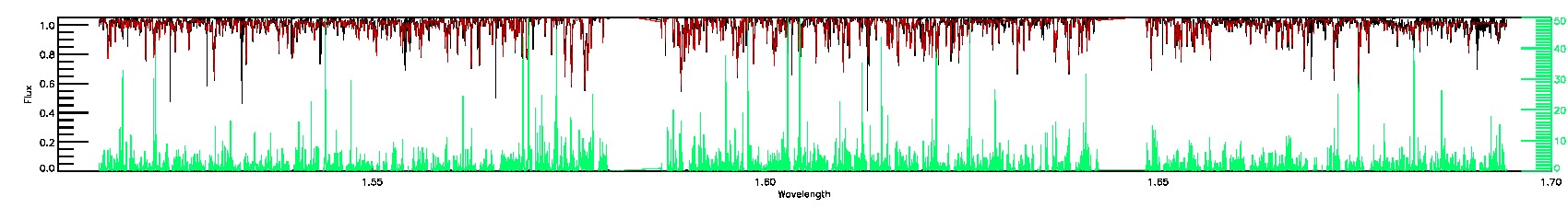
055-53-O
| 5357. | +/- | 13. |
| 5335. | +/- | 124. |
| 4.47 | +/- | 0. |
| 4.36 | +/- | 0. |
| 0.41 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 1.80 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.13 |
| -9999.00 |

055-53-O
| 4003. | +/- | 3. |
| 4106. | +/- | 81. |
| 4.33 | +/- | 0. |
| 4.71 | +/- | 0. |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -0.34 | +/- | 0. |
| -0.34 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -0.35 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 4.86 | +/- | 0. |
| 0.69 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.38 |
| -9999.00 |
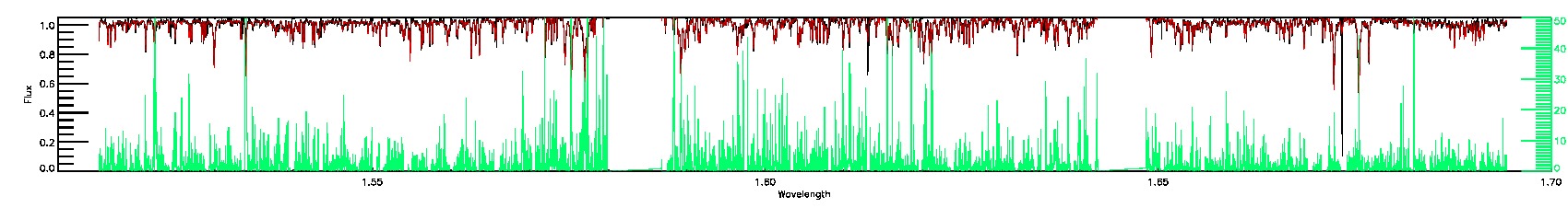
055-53-O
| 5472. | +/- | 9. |
| 5446. | +/- | 105. |
| 4.67 | +/- | 0. |
| 4.62 | +/- | 0. |
| 1.72 | +/- | 0. |
| 1.72 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.04 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.12 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 5.88 | +/- | 0. |
| 0.77 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.17 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |

055-53-O
| 4682. | +/- | 8. |
| 4756. | +/- | 106. |
| 4.29 | +/- | 0. |
| 4.46 | +/- | 0. |
| 0.42 | +/- | 0. |
| 0.42 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -0.14 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 6.47 | +/- | 0. |
| 0.81 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.11 |
| -9999.00 |

055-53-O
| 4142. | +/- | 3. |
| 4231. | +/- | 84. |
| 4.50 | +/- | 0. |
| 4.72 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 3.46 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.15 |
| -9999.00 |

055-53-O
| 4292. | +/- | 2. |
| 4388. | +/- | 75. |
| 2.28 | +/- | 0. |
| 2.07 | +/- | 0. |
| 1.35 | +/- | 0. |
| 1.35 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -0.18 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.14 | +/- | 0. |
| 3.28 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.49 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
055-53-O
| 3712. | +/- | 5. |
| 3819. | +/- | 85. |
| 4.37 | +/- | 0. |
| 4.85 | +/- | 0. |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -0.44 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 4.15 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.44 |
| -9999.00 |
STAR_WARN,SN_WARN
055-53-O
| 5475. | +/- | 45. |
| 5460. | +/- | 148. |
| 4.30 | +/- | 0. |
| 4.36 | +/- | 0. |
| 1.62 | +/- | 0. |
| 1.62 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.06 | +/- | 0. |
| 11.13 | +/- | 0. |
| 1.05 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.17 |
| -9999.00 |
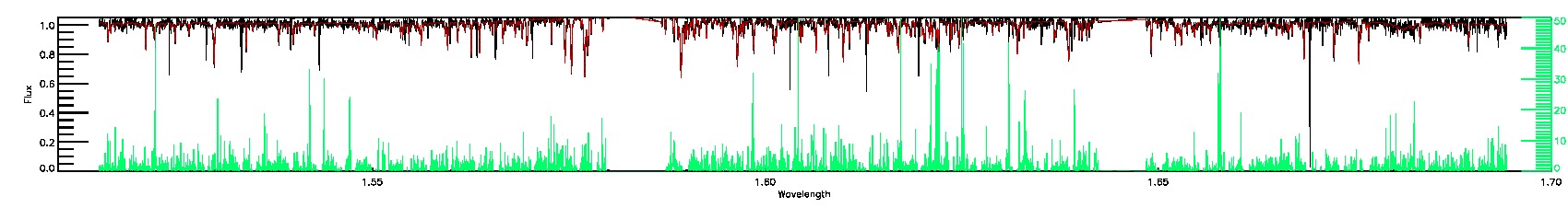
055-53-O
| 6012. | +/- | 18. |
| 5930. | +/- | 134. |
| 4.51 | +/- | 0. |
| 4.41 | +/- | 0. |
| 0.95 | +/- | 0. |
| 0.95 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.17 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.03 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.31 |
| -9999.00 |
055-53-O
| 5552. | +/- | 15. |
| 5527. | +/- | 119. |
| 4.46 | +/- | 0. |
| 4.49 | +/- | 0. |
| 1.11 | +/- | 0. |
| 1.11 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -0.19 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.06 | +/- | 0. |
| 2.71 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.54 |
| -9999.00 |

055-53-O
| 4568. | +/- | 3. |
| 4648. | +/- | 80. |
| 2.72 | +/- | 0. |
| 2.40 | +/- | 0. |
| 1.23 | +/- | 0. |
| 1.23 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.10 | +/- | 0. |
| 2.94 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -0.10 |
| -9999.00 |

055-53-O
| 6026. | +/- | 28. |
| 5947. | +/- | 157. |
| 4.62 | +/- | 0. |
| 4.57 | +/- | 0. |
| 0.79 | +/- | 0. |
| 0.79 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.18 | +/- | 0. |
| -0.18 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 1.71 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -2.38 |
| -9999.00 |
STAR_WARN,SN_WARN
055-53-O
| 6220. | +/- | 72. |
| 6125. | +/- | 187. |
| 4.42 | +/- | 0. |
| 4.40 | +/- | 0. |
| 1.65 | +/- | 0. |
| 1.65 | +/- | -NaN |
| -0.39 | +/- | 0. |
| -0.38 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.12 | +/- | -NaN |
| 1.01 | +/- | 0. |
| 1.01 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 5.08 | +/- | 0. |
| 0.71 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -2.23 |
| -9999.00 |
STAR_WARN,SN_WARN
055-53-O
| 4724. | +/- | 22. |
| 4802. | +/- | 125. |
| 4.42 | +/- | 0. |
| 4.68 | +/- | 0. |
| 0.44 | +/- | 0. |
| 0.44 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -0.36 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -0.31 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.00 | +/- | 0. |
| 1.72 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.57 |
| -9999.00 |
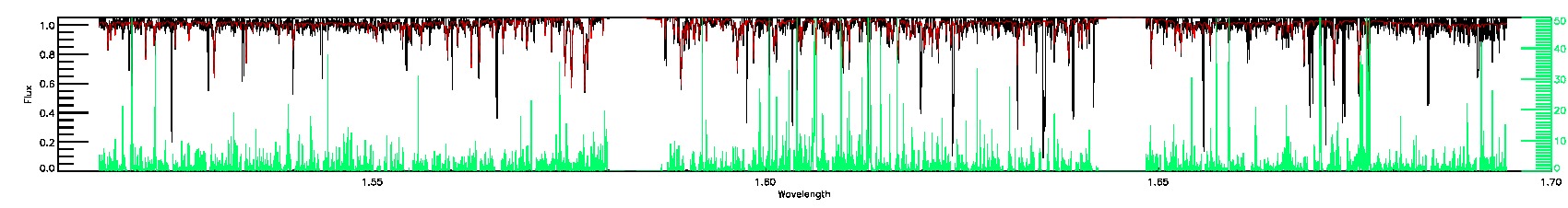
055-53-O
| 5002. | +/- | 5. |
| 5020. | +/- | 89. |
| 4.55 | +/- | 0. |
| 4.49 | +/- | 0. |
| 0.89 | +/- | 0. |
| 0.89 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.27 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.09 | +/- | 0. |
| 2.21 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.23 |
| -9999.00 |

055-53-O
| 5863. | +/- | 11. |
| 5789. | +/- | 121. |
| 4.42 | +/- | 0. |
| 4.27 | +/- | 0. |
| 1.30 | +/- | 0. |
| 1.30 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 6.40 | +/- | 0. |
| 0.81 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.07 |
| -9999.00 |

055-53-O
| 5747. | +/- | 26. |
| 5697. | +/- | 148. |
| 4.27 | +/- | 0. |
| 4.24 | +/- | 0. |
| 1.17 | +/- | 0. |
| 1.17 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.13 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.11 | +/- | 0. |
| 1.72 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| 0.11 |
| -9999.00 |

055-53-O
| 6047. | +/- | 33. |
| 5966. | +/- | 156. |
| 4.46 | +/- | 0. |
| 4.41 | +/- | 0. |
| 1.36 | +/- | 0. |
| 1.36 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -0.23 | +/- | 0. |
| -0.21 | +/- | 0. |
| -0.21 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -0.31 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 4.27 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.28 |
| -9999.00 |
055-53-O
| 6323. | +/- | 21. |
| 6202. | +/- | 150. |
| 4.66 | +/- | 0. |
| 4.48 | +/- | 0. |
| 0.47 | +/- | 0. |
| 0.47 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.27 | +/- | 0. |
| -0.27 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -0.25 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 4.34 | +/- | 0. |
| 0.64 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.41 |
| -9999.00 |
055-53-O
| 4482. | +/- | 7. |
| 4580. | +/- | 105. |
| 4.26 | +/- | 0. |
| 4.50 | +/- | 0. |
| 0.80 | +/- | 0. |
| 0.80 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 6.88 | +/- | 0. |
| 0.84 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.10 |
| -9999.00 |

STAR_WARN,SN_WARN
055-53-O
| 5819. | +/- | 37. |
| 5762. | +/- | 160. |
| 4.49 | +/- | 0. |
| 4.46 | +/- | 0. |
| 1.24 | +/- | 0. |
| 1.24 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -0.18 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 1.63 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -2.24 |
| -9999.00 |

055-53-O
| 4920. | +/- | 17. |
| 4979. | +/- | 123. |
| 3.48 | +/- | 0. |
| 3.39 | +/- | 0. |
| 0.96 | +/- | 0. |
| 0.96 | +/- | -NaN |
| -0.46 | +/- | 0. |
| -0.46 | +/- | 0. |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| -0.21 | +/- | 0. |
| -0.21 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.19 | +/- | 0. |
| 3.87 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.05 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
055-53-O
| 6212. | +/- | 17. |
| 6105. | +/- | 132. |
| 4.75 | +/- | 0. |
| 4.61 | +/- | 0. |
| 0.53 | +/- | 0. |
| 0.53 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.17 | +/- | 0. |
| -0.17 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 5.83 | +/- | 0. |
| 0.77 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.01 |
| -9999.00 |
SUSPECT_BROAD_LINES
055-53-O
| 5774. | +/- | 51. |
| 5737. | +/- | 161. |
| 4.28 | +/- | 0. |
| 4.40 | +/- | 0. |
| 0.88 | +/- | 0. |
| 0.88 | +/- | -NaN |
| -0.52 | +/- | 0. |
| -0.52 | +/- | 0. |
| -0.21 | +/- | 0. |
| -0.21 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -0.25 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.15 | +/- | 0. |
| 17.03 | +/- | 0. |
| 1.23 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| -0.69 |
| -9999.00 |

055-53-O
| 5614. | +/- | 22. |
| 5568. | +/- | 140. |
| 4.36 | +/- | 0. |
| 4.25 | +/- | 0. |
| 0.84 | +/- | 0. |
| 0.84 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.15 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.01 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.14 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.04 |
| -9999.00 |

STAR_WARN,SN_WARN
055-53-O
| 5713. | +/- | 40. |
| 5675. | +/- | 157. |
| 4.20 | +/- | 0. |
| 4.25 | +/- | 0. |
| 1.34 | +/- | 0. |
| 1.34 | +/- | -NaN |
| -0.34 | +/- | 0. |
| -0.33 | +/- | 0. |
| -0.32 | +/- | 0. |
| -0.32 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.19 | +/- | 0. |
| 2.04 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.85 |
| -9999.00 |
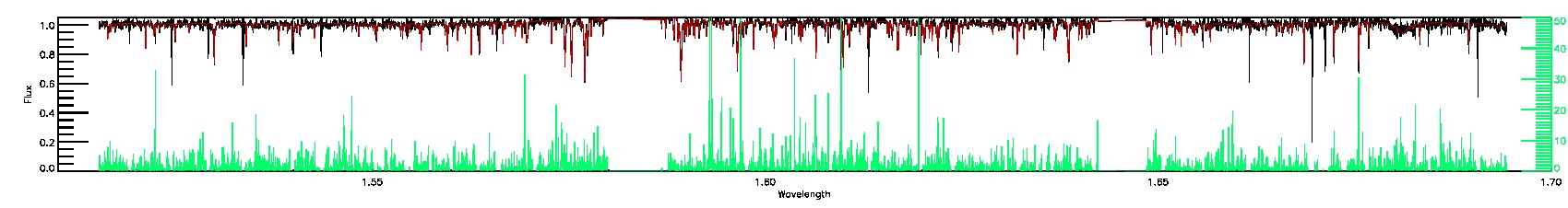
055-53-O
| 4558. | +/- | 8. |
| 4663. | +/- | 113. |
| 2.62 | +/- | 0. |
| 2.47 | +/- | 0. |
| 1.29 | +/- | 0. |
| 1.29 | +/- | -NaN |
| -0.55 | +/- | 0. |
| -0.55 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.24 | +/- | 0. |
| 4.08 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -1.41 |
| -9999.00 |

055-53-O
| 4600. | +/- | 6. |
| 4669. | +/- | 96. |
| 4.30 | +/- | 0. |
| 4.35 | +/- | 0. |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.10 | +/- | 0. |
| 3.43 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.21 |
| -9999.00 |
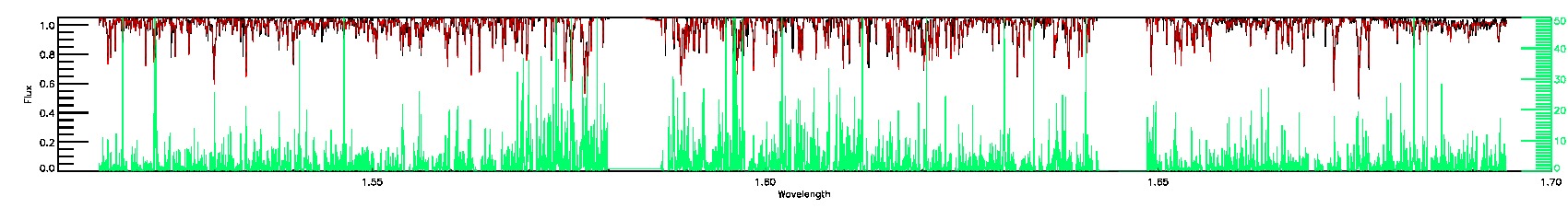
STAR_WARN,SN_WARN
055-53-O
| 4292. | +/- | 7. |
| 4376. | +/- | 97. |
| 4.32 | +/- | 0. |
| 4.46 | +/- | 0. |
| 0.51 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.09 | +/- | 0. |
| 5.58 | +/- | 0. |
| 0.75 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.08 |
| -9999.00 |
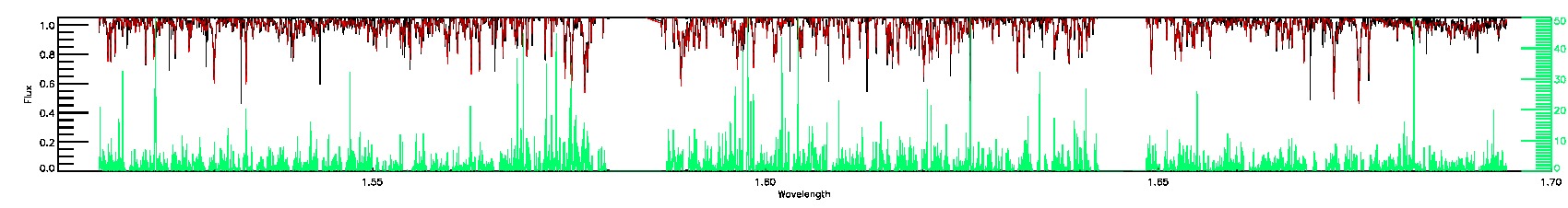
STAR_WARN,SN_WARN
055-53-O
| 5692. | +/- | 46. |
| 5647. | +/- | 158. |
| 4.29 | +/- | 0. |
| 4.26 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 2.19 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -1.82 |
| -9999.00 |

STAR_WARN,SN_WARN
055-53-O
| 3986. | +/- | 7. |
| 4110. | +/- | 106. |
| 4.18 | +/- | 0. |
| 4.77 | +/- | 0. |
| 0.65 | +/- | 0. |
| 0.65 | +/- | -NaN |
| -0.85 | +/- | 0. |
| -0.84 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.22 | +/- | 0. |
| 4.76 | +/- | 0. |
| 0.68 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.64 |
| -9999.00 |
STAR_WARN,COLORTE_WARN,SN_WARN
055-53-O
| 4456. | +/- | 17. |
| 4595. | +/- | 121. |
| 3.33 | +/- | 0. |
| 3.32 | +/- | 0. |
| 0.59 | +/- | 0. |
| 0.59 | +/- | -NaN |
| -1.21 | +/- | 0. |
| -1.21 | +/- | 0. |
| -0.20 | +/- | 0. |
| -0.20 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -0.49 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.04 | +/- | 0. |
| 5.99 | +/- | 0. |
| 0.78 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.27 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.74 |
| -9999.00 |
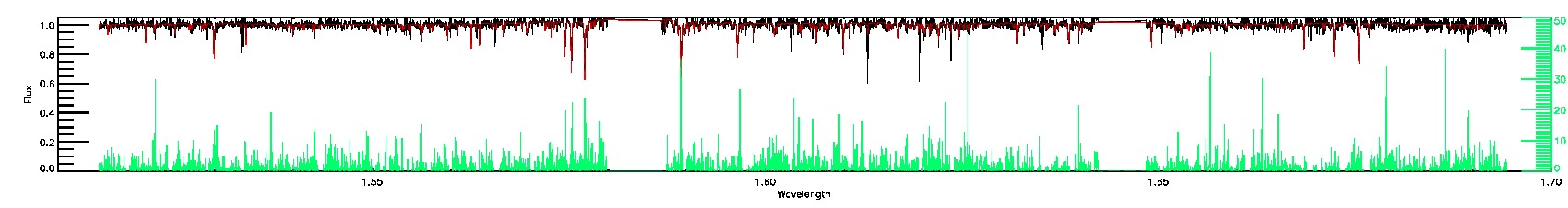
STAR_BAD
055-53-O
| 6094. | +/- | 25. |
| -10000. | +/- | -NaN |
| 4.41 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.15 | +/- | 0. |
| 1.15 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.30 | +/- | 0. |
| 0.80 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -2.16 |
| -9999.00 |
STAR_BAD
055-53-O
| 4363. | +/- | 12. |
| -10000. | +/- | -NaN |
| 0.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.08 | +/- | 0. |
| 2.08 | +/- | -NaN |
| -2.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.59 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.27 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.48 |
| -9999.00 |

STAR_WARN,SN_WARN
055-53-O
| 4880. | +/- | 20. |
| 4942. | +/- | 123. |
| 4.43 | +/- | 0. |
| 4.68 | +/- | 0. |
| 1.90 | +/- | 0. |
| 1.90 | +/- | -NaN |
| -0.42 | +/- | 0. |
| -0.41 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.07 | +/- | 0. |
| 4.42 | +/- | 0. |
| 0.64 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.36 |
| -9999.00 |
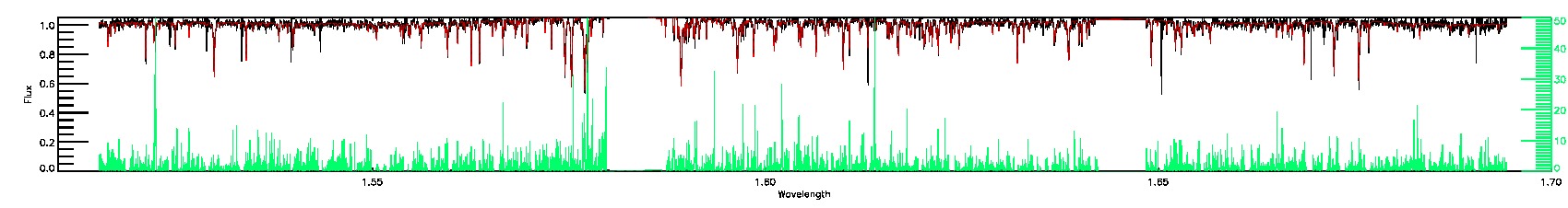
055-53-O
| 5114. | +/- | 12. |
| 5166. | +/- | 113. |
| 2.72 | +/- | 0. |
| 2.40 | +/- | 0. |
| 1.67 | +/- | 0. |
| 1.67 | +/- | -NaN |
| -0.82 | +/- | 0. |
| -0.82 | +/- | 0. |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.27 | +/- | 0. |
| 4.77 | +/- | 0. |
| 0.68 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.20 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -1.65 |
| -9999.00 |

055-53-O
| 4477. | +/- | 2. |
| 4562. | +/- | 77. |
| 2.71 | +/- | 0. |
| 2.48 | +/- | 0. |
| 1.19 | +/- | 0. |
| 1.19 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.79 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.15 |
| -9999.00 |

STAR_WARN,SN_WARN
055-53-O
| 5677. | +/- | 47. |
| 5643. | +/- | 156. |
| 4.34 | +/- | 0. |
| 4.41 | +/- | 0. |
| 1.07 | +/- | 0. |
| 1.07 | +/- | -NaN |
| -0.34 | +/- | 0. |
| -0.34 | +/- | 0. |
| -0.24 | +/- | 0. |
| -0.24 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -0.25 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 4.89 | +/- | 0. |
| 0.69 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.16 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.26 |
| -9999.00 |

055-53-O
| 4573. | +/- | 4. |
| 4673. | +/- | 94. |
| 2.65 | +/- | 0. |
| 2.49 | +/- | 0. |
| 1.42 | +/- | 0. |
| 1.42 | +/- | -NaN |
| -0.47 | +/- | 0. |
| -0.47 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.20 | +/- | 0. |
| 3.89 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -0.69 |
| -9999.00 |

BRIGHT_NEIGHBOR
055-53-O
| 4052. | +/- | 2. |
| 4182. | +/- | 85. |
| 1.29 | +/- | 0. |
| 1.27 | +/- | 0. |
| 1.89 | +/- | 0. |
| 1.89 | +/- | -NaN |
| -0.99 | +/- | 0. |
| -0.99 | +/- | 0. |
| -0.25 | +/- | 0. |
| -0.25 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.16 | +/- | 0. |
| 5.27 | +/- | 0. |
| 0.72 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.25 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -2.02 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_WARN,SN_WARN
055-53-O
| 5205. | +/- | 40. |
| 5218. | +/- | 133. |
| 4.81 | +/- | 0. |
| 4.88 | +/- | 0. |
| 0.43 | +/- | 0. |
| 0.43 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.46 | +/- | 0. |
| -0.46 | +/- | -NaN |
| 0.67 | +/- | 0. |
| 0.67 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.05 | +/- | 0. |
| 39.62 | +/- | 0. |
| 1.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.46 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 1.00 |
| -9999.00 |
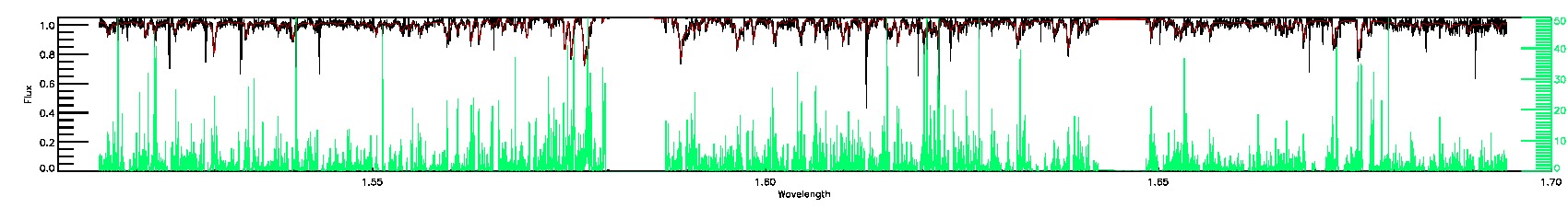
STAR_WARN,SN_WARN
055-53-O
| 5706. | +/- | 39. |
| 5675. | +/- | 160. |
| 4.09 | +/- | 0. |
| 4.21 | +/- | 0. |
| 0.75 | +/- | 0. |
| 0.75 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -0.49 | +/- | 0. |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -0.31 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.17 | +/- | 0. |
| 3.94 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.75 |
| -9999.00 |

055-53-O
| 5685. | +/- | 27. |
| 5656. | +/- | 145. |
| 4.38 | +/- | 0. |
| 4.50 | +/- | 0. |
| 0.80 | +/- | 0. |
| 0.80 | +/- | -NaN |
| -0.48 | +/- | 0. |
| -0.47 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.16 | +/- | 0. |
| 2.23 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.39 |
| -9999.00 |
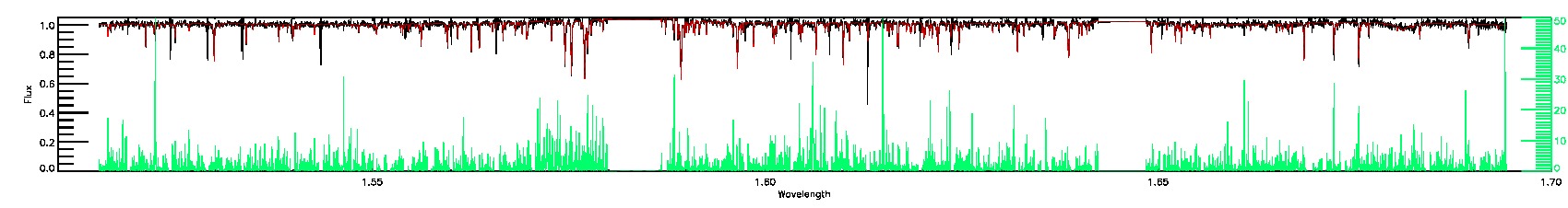
STAR_WARN,SN_WARN
055-53-O
| 5596. | +/- | 43. |
| 5555. | +/- | 155. |
| 4.66 | +/- | 0. |
| 4.57 | +/- | 0. |
| 0.45 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.01 | +/- | 0. |
| 1.66 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| -0.08 |
| -9999.00 |

STAR_WARN,SN_WARN
055-53-O
| 5121. | +/- | 39. |
| 5159. | +/- | 139. |
| 4.33 | +/- | 0. |
| 4.56 | +/- | 0. |
| 0.58 | +/- | 0. |
| 0.58 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -0.50 | +/- | 0. |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.24 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.15 | +/- | 0. |
| 1.90 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.24 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.10 |
| -9999.00 |

055-53-O
| 4608. | +/- | 5. |
| 4689. | +/- | 96. |
| 4.18 | +/- | 0. |
| 4.36 | +/- | 0. |
| 0.44 | +/- | 0. |
| 0.44 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.13 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.12 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 6.37 | +/- | 0. |
| 0.80 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.02 |
| -9999.00 |

