120-61-O
| 5678. | +/- | 15. |
| 5628. | +/- | 127. |
| 4.20 | +/- | 0. |
| 4.11 | +/- | 0. |
| 1.18 | +/- | 0. |
| 1.18 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | 0. |
| -0.25 | +/- | 0. |
| -0.25 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 1.79 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.13 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_WARN,SN_WARN
120-61-O
| 4100. | +/- | 8. |
| 4202. | +/- | 99. |
| 4.44 | +/- | 0. |
| 4.79 | +/- | 0. |
| 0.40 | +/- | 0. |
| 0.40 | +/- | -NaN |
| -0.32 | +/- | 0. |
| -0.31 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 5.50 | +/- | 0. |
| 0.74 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| -0.40 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
120-61-O
| 3563. | +/- | 4. |
| 3652. | +/- | 75. |
| 4.51 | +/- | 0. |
| 4.84 | +/- | 0. |
| 0.89 | +/- | 0. |
| 0.89 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -0.14 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.05 | +/- | 0. |
| 2.69 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.33 |
| -9999.00 |
120-61-O
| 5627. | +/- | 18. |
| 5584. | +/- | 137. |
| 4.44 | +/- | 0. |
| 4.37 | +/- | 0. |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.03 | +/- | 0. |
| -0.17 | +/- | 0. |
| -0.17 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.00 | +/- | 0. |
| 1.62 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.11 |
| -9999.00 |

STAR_WARN,SN_WARN
120-61-O
| 4842. | +/- | 22. |
| 4910. | +/- | 127. |
| 4.48 | +/- | 0. |
| 4.74 | +/- | 0. |
| 0.51 | +/- | 0. |
| 0.51 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -0.44 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.18 | +/- | 0. |
| 1.64 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.37 |
| -9999.00 |
STAR_WARN,SN_WARN
120-61-O
| 5020. | +/- | 21. |
| 5050. | +/- | 125. |
| 4.55 | +/- | 0. |
| 4.61 | +/- | 0. |
| 1.45 | +/- | 0. |
| 1.45 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.00 | +/- | 0. |
| 1.81 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.21 |
| -9999.00 |

120-61-O
| 5234. | +/- | 14. |
| 5242. | +/- | 116. |
| 3.72 | +/- | 0. |
| 3.68 | +/- | 0. |
| 1.25 | +/- | 0. |
| 1.25 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.13 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 3.18 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.22 |
| -9999.00 |
120-61-O
| 5850. | +/- | 34. |
| 5794. | +/- | 157. |
| 4.18 | +/- | 0. |
| 4.18 | +/- | 0. |
| 1.10 | +/- | 0. |
| 1.10 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -0.28 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.06 | +/- | 0. |
| 1.98 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.48 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,ROTATION_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
120-61-O
| 4293. | +/- | 22. |
| -10000. | +/- | -NaN |
| 4.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.49 | +/- | 0. |
| 0.49 | +/- | -NaN |
| -1.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.75 | +/- | 0. |
| 0.76 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
120-61-O
| 5954. | +/- | 33. |
| 5904. | +/- | 148. |
| 4.21 | +/- | 0. |
| 4.37 | +/- | 0. |
| 0.88 | +/- | 0. |
| 0.88 | +/- | -NaN |
| -0.71 | +/- | 0. |
| -0.71 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.11 | +/- | 0. |
| 3.58 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -2.35 |
| -9999.00 |

BRIGHT_NEIGHBOR
120-61-O
| 5795. | +/- | 39. |
| 5759. | +/- | 153. |
| 4.31 | +/- | 0. |
| 4.46 | +/- | 0. |
| 0.54 | +/- | 0. |
| 0.54 | +/- | -NaN |
| -0.60 | +/- | 0. |
| -0.60 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -0.19 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.19 | +/- | 0. |
| 8.17 | +/- | 0. |
| 0.91 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.59 |
| -9999.00 |
120-61-O
| 5619. | +/- | 9. |
| 5575. | +/- | 109. |
| 4.45 | +/- | 0. |
| 4.36 | +/- | 0. |
| 0.50 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 2.03 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |

120-61-O
| 4927. | +/- | 6. |
| 4985. | +/- | 94. |
| 2.75 | +/- | 0. |
| 2.42 | +/- | 0. |
| 1.75 | +/- | 0. |
| 1.75 | +/- | -NaN |
| -0.45 | +/- | 0. |
| -0.45 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.08 | +/- | 0. |
| 3.85 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.98 |
| -9999.00 |

120-61-O
| 5536. | +/- | 26. |
| 5527. | +/- | 141. |
| 3.93 | +/- | 0. |
| 4.07 | +/- | 0. |
| 0.98 | +/- | 0. |
| 0.98 | +/- | -NaN |
| -0.54 | +/- | 0. |
| -0.54 | +/- | 0. |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.13 | +/- | 0. |
| 4.06 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.53 |
| -9999.00 |
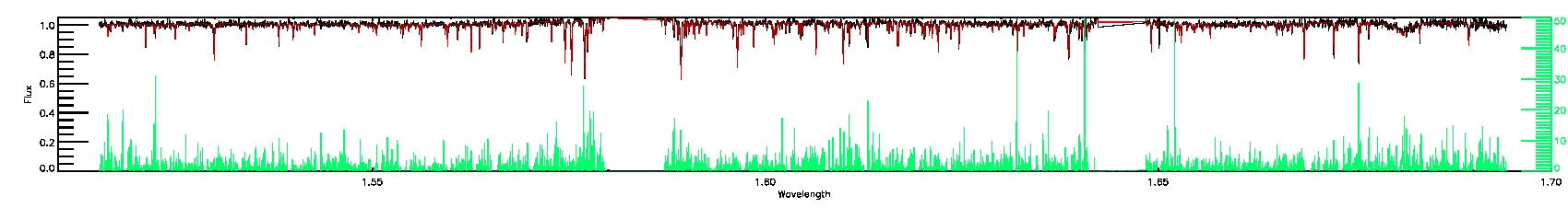
120-61-O
| 5235. | +/- | 28. |
| 5251. | +/- | 134. |
| 4.50 | +/- | 0. |
| 4.63 | +/- | 0. |
| 0.67 | +/- | 0. |
| 0.67 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -0.31 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.07 | +/- | 0. |
| 1.56 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.19 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.83 |
| -9999.00 |
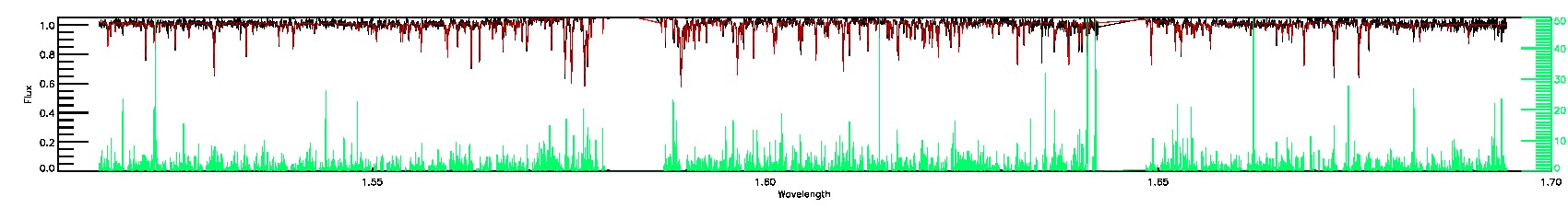
STAR_WARN,COLORTE_WARN,SN_WARN
120-61-O
| 3647. | +/- | 6. |
| 3749. | +/- | 84. |
| 4.41 | +/- | 0. |
| 4.84 | +/- | 0. |
| 0.70 | +/- | 0. |
| 0.70 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -0.31 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.11 | +/- | 0. |
| 5.10 | +/- | 0. |
| 0.71 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| 0.12 |
| -9999.00 |
SUSPECT_BROAD_LINES
120-61-O
| 3913. | +/- | 2. |
| 4000. | +/- | 66. |
| 4.52 | +/- | 0. |
| 4.77 | +/- | 0. |
| 0.94 | +/- | 0. |
| 0.94 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 42.87 | +/- | 0. |
| 1.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.21 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
120-61-O
| 3374. | +/- | 6. |
| 3473. | +/- | 77. |
| 4.42 | +/- | 0. |
| 4.87 | +/- | 0. |
| 1.56 | +/- | 0. |
| 1.56 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.24 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.04 | +/- | 0. |
| 6.36 | +/- | 0. |
| 0.80 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.25 |
| -9999.00 |
BRIGHT_NEIGHBOR
120-61-O
| 4624. | +/- | 7. |
| 4717. | +/- | 105. |
| 2.73 | +/- | 0. |
| 2.58 | +/- | 0. |
| 1.21 | +/- | 0. |
| 1.21 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -0.44 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.23 | +/- | 0. |
| 3.82 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.61 |
| -9999.00 |

BRIGHT_NEIGHBOR
STAR_WARN,SN_WARN
120-61-O
| 4026. | +/- | 8. |
| 4114. | +/- | 94. |
| 4.58 | +/- | 0. |
| 4.81 | +/- | 0. |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -0.22 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 3.53 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.23 |
| -9999.00 |
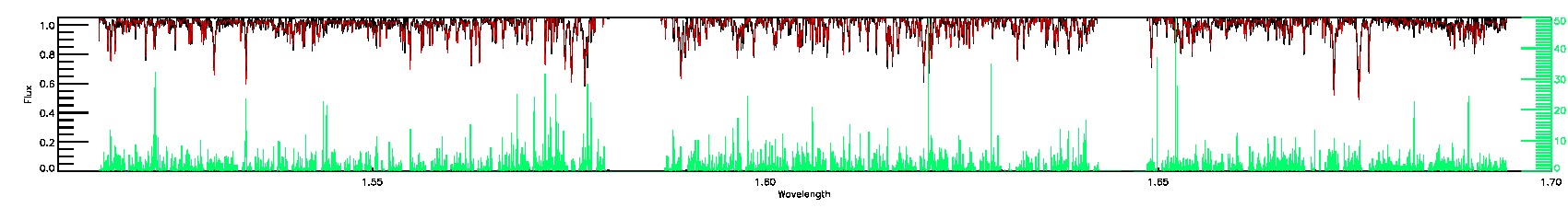
STAR_WARN,SN_WARN
120-61-O
| 5008. | +/- | 28. |
| 5058. | +/- | 131. |
| 4.44 | +/- | 0. |
| 4.68 | +/- | 0. |
| 1.64 | +/- | 0. |
| 1.64 | +/- | -NaN |
| -0.47 | +/- | 0. |
| -0.47 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.15 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.06 | +/- | 0. |
| 1.90 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.15 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.43 |
| -9999.00 |

BRIGHT_NEIGHBOR
120-61-O
| 6118. | +/- | 44. |
| 6026. | +/- | 169. |
| 4.17 | +/- | 0. |
| 4.08 | +/- | 0. |
| 1.33 | +/- | 0. |
| 1.33 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.17 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 7.10 | +/- | 0. |
| 0.85 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.29 |
| -9999.00 |
120-61-O
| 6069. | +/- | 15. |
| 5974. | +/- | 127. |
| 4.65 | +/- | 0. |
| 4.48 | +/- | 0. |
| 0.62 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.03 | +/- | 0. |
| -0.17 | +/- | 0. |
| -0.17 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 4.39 | +/- | 0. |
| 0.64 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.09 |
| -9999.00 |
120-61-O
| 5366. | +/- | 14. |
| 5343. | +/- | 122. |
| 4.29 | +/- | 0. |
| 4.17 | +/- | 0. |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.26 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 8.04 | +/- | 0. |
| 0.91 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.55 |
| -9999.00 |

LOW_SNR
STAR_WARN,SN_WARN
120-61-O
| 4974. | +/- | 26. |
| 5019. | +/- | 129. |
| 3.60 | +/- | 0. |
| 3.54 | +/- | 0. |
| 1.17 | +/- | 0. |
| 1.17 | +/- | -NaN |
| -0.27 | +/- | 0. |
| -0.27 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 1.61 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -0.32 |
| -9999.00 |

120-61-O
| 5061. | +/- | 7. |
| 5087. | +/- | 93. |
| 3.53 | +/- | 0. |
| 3.40 | +/- | 0. |
| 1.15 | +/- | 0. |
| 1.15 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| 3.04 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |

BRIGHT_NEIGHBOR
STAR_WARN,SN_WARN
120-61-O
| 5316. | +/- | 28. |
| 5313. | +/- | 139. |
| 4.54 | +/- | 0. |
| 4.57 | +/- | 0. |
| 0.55 | +/- | 0. |
| 0.55 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.07 | +/- | 0. |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.12 | +/- | 0. |
| 1.92 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.31 |
| -9999.00 |

120-61-O
| 5532. | +/- | 10. |
| 5502. | +/- | 109. |
| 4.48 | +/- | 0. |
| 4.45 | +/- | 0. |
| 1.15 | +/- | 0. |
| 1.15 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 1.55 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.43 |
| -9999.00 |

STAR_WARN,SN_WARN
120-61-O
| 5225. | +/- | 45. |
| 5255. | +/- | 147. |
| 3.87 | +/- | 0. |
| 4.04 | +/- | 0. |
| 0.51 | +/- | 0. |
| 0.51 | +/- | -NaN |
| -0.60 | +/- | 0. |
| -0.60 | +/- | 0. |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -0.45 | +/- | 0. |
| -0.45 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.23 | +/- | 0. |
| 4.21 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.24 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -1.15 |
| -9999.00 |

120-61-O
| 4333. | +/- | 4. |
| 4433. | +/- | 95. |
| 4.30 | +/- | 0. |
| 4.59 | +/- | 0. |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -0.25 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.11 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.65 |
| -9999.00 |

120-61-O
| 4597. | +/- | 3. |
| 4673. | +/- | 81. |
| 2.70 | +/- | 0. |
| 2.39 | +/- | 0. |
| 1.24 | +/- | 0. |
| 1.24 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 2.91 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.04 |
| -9999.00 |

120-61-O
| 5572. | +/- | 21. |
| 5557. | +/- | 130. |
| 4.15 | +/- | 0. |
| 4.29 | +/- | 0. |
| 0.93 | +/- | 0. |
| 0.93 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -0.49 | +/- | 0. |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| -0.42 | +/- | 0. |
| -0.42 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.21 | +/- | 0. |
| 3.95 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.86 |
| -9999.00 |

120-61-O
| 6154. | +/- | 26. |
| 6059. | +/- | 161. |
| 4.68 | +/- | 0. |
| 4.59 | +/- | 0. |
| 0.52 | +/- | 0. |
| 0.52 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -0.19 | +/- | 0. |
| -0.16 | +/- | 0. |
| -0.16 | +/- | -NaN |
| 0.53 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 1.72 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -0.39 |
| -9999.00 |
STAR_WARN,SN_WARN
120-61-O
| 5692. | +/- | 40. |
| 5652. | +/- | 155. |
| 4.56 | +/- | 0. |
| 4.58 | +/- | 0. |
| 0.81 | +/- | 0. |
| 0.81 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -0.25 | +/- | 0. |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 2.29 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -0.12 |
| -9999.00 |
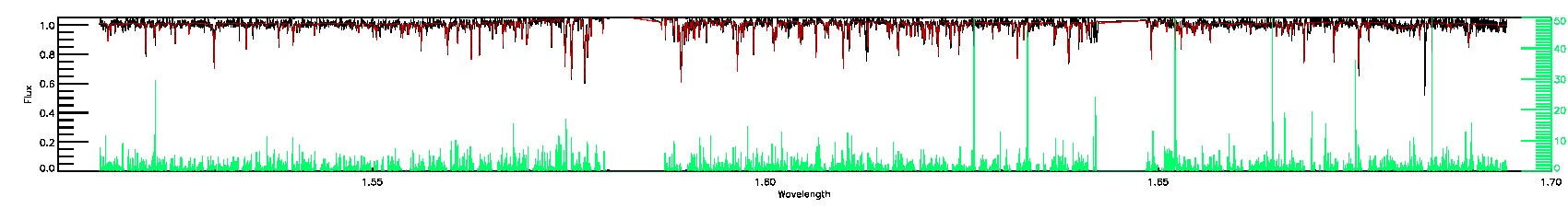
BRIGHT_NEIGHBOR
STAR_WARN,SN_WARN
120-61-O
| 5817. | +/- | 53. |
| 5762. | +/- | 165. |
| 4.49 | +/- | 0. |
| 4.48 | +/- | 0. |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -0.21 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.09 | +/- | 0. |
| 6.02 | +/- | 0. |
| 0.78 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.12 |
| -9999.00 |
120-61-O
| 5080. | +/- | 7. |
| 5108. | +/- | 95. |
| 3.53 | +/- | 0. |
| 3.41 | +/- | 0. |
| 1.22 | +/- | 0. |
| 1.22 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.17 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 3.27 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.13 |
| -9999.00 |

STAR_BAD
STAR_WARN,SN_WARN
120-61-O
| 4997. | +/- | 30. |
| -10000. | +/- | -NaN |
| 4.32 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.74 | +/- | 0. |
| 1.74 | +/- | -NaN |
| -0.62 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.26 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.64 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.24 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.26 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
120-61-O
| 3504. | +/- | 3. |
| 3585. | +/- | 64. |
| 4.69 | +/- | 0. |
| 4.96 | +/- | 0. |
| 1.26 | +/- | 0. |
| 1.26 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| -0.30 | +/- | 0. |
| -0.30 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.06 | +/- | 0. |
| 5.03 | +/- | 0. |
| 0.70 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.29 |
| -9999.00 |
120-61-O
| 5218. | +/- | 10. |
| 5226. | +/- | 113. |
| 4.63 | +/- | 0. |
| 4.67 | +/- | 0. |
| 0.55 | +/- | 0. |
| 0.55 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 1.55 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.31 |
| -9999.00 |

120-61-O
| 5603. | +/- | 17. |
| 5561. | +/- | 128. |
| 4.26 | +/- | 0. |
| 4.18 | +/- | 0. |
| 1.15 | +/- | 0. |
| 1.15 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 4.81 | +/- | 0. |
| 0.68 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.25 |
| -9999.00 |
STAR_WARN,SN_WARN
120-61-O
| 5690. | +/- | 47. |
| 5658. | +/- | 157. |
| 4.06 | +/- | 0. |
| 4.15 | +/- | 0. |
| 1.66 | +/- | 0. |
| 1.66 | +/- | -NaN |
| -0.42 | +/- | 0. |
| -0.41 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 1.62 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.53 |
| -9999.00 |

120-61-O
| 6144. | +/- | 33. |
| 6060. | +/- | 159. |
| 4.82 | +/- | 0. |
| 4.83 | +/- | 0. |
| 1.10 | +/- | 0. |
| 1.10 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -0.43 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.13 | +/- | 0. |
| 8.28 | +/- | 0. |
| 0.92 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.20 |
| -9999.00 |
120-61-O
| 5767. | +/- | 21. |
| 5707. | +/- | 141. |
| 4.44 | +/- | 0. |
| 4.34 | +/- | 0. |
| 1.17 | +/- | 0. |
| 1.17 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.15 | +/- | 0. |
| 0.33 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.09 |
| -9999.00 |

SUSPECT_BROAD_LINES
120-61-O
| 6403. | +/- | 15. |
| 6258. | +/- | 134. |
| 4.45 | +/- | 0. |
| 4.12 | +/- | 0. |
| 1.72 | +/- | 0. |
| 1.72 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | 0. |
| -0.16 | +/- | 0. |
| -0.16 | +/- | -NaN |
| 0.50 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 13.67 | +/- | 0. |
| 1.14 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.21 |
| -9999.00 |
SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES
STAR_BAD,ROTATION_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
120-61-O
| 4377. | +/- | 14. |
| -10000. | +/- | -NaN |
| 4.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.59 | +/- | 0. |
| 1.59 | +/- | -NaN |
| -0.98 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.24 | +/- | 0. |
| 0.72 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -2.14 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.77 |
| -9999.00 |
120-61-O
| 4837. | +/- | 11. |
| 4882. | +/- | 110. |
| 4.51 | +/- | 0. |
| 4.55 | +/- | 0. |
| 0.94 | +/- | 0. |
| 0.94 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 1.75 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.16 |
| -9999.00 |

120-61-O
| 5942. | +/- | 12. |
| 5857. | +/- | 131. |
| 4.38 | +/- | 0. |
| 4.19 | +/- | 0. |
| 1.29 | +/- | 0. |
| 1.29 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.46 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.28 |
| -9999.00 |

120-61-O
| 6234. | +/- | 15. |
| 6119. | +/- | 131. |
| 4.36 | +/- | 0. |
| 4.16 | +/- | 0. |
| 1.40 | +/- | 0. |
| 1.40 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | 0. |
| -0.17 | +/- | 0. |
| -0.17 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.00 | +/- | 0. |
| 8.46 | +/- | 0. |
| 0.93 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.08 |
| -9999.00 |
STAR_WARN,SN_WARN
120-61-O
| 5622. | +/- | 48. |
| 5595. | +/- | 156. |
| 4.32 | +/- | 0. |
| 4.39 | +/- | 0. |
| 1.73 | +/- | 0. |
| 1.73 | +/- | -NaN |
| -0.34 | +/- | 0. |
| -0.34 | +/- | 0. |
| -0.16 | +/- | 0. |
| -0.16 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -0.35 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.09 | +/- | 0. |
| 1.50 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -1.05 |
| -9999.00 |

120-61-O
| 5944. | +/- | 19. |
| 5884. | +/- | 127. |
| 4.51 | +/- | 0. |
| 4.57 | +/- | 0. |
| 0.41 | +/- | 0. |
| 0.41 | +/- | -NaN |
| -0.46 | +/- | 0. |
| -0.46 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 4.49 | +/- | 0. |
| 0.65 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.61 |
| -9999.00 |
120-61-O
| 5073. | +/- | 13. |
| 5089. | +/- | 117. |
| 4.61 | +/- | 0. |
| 4.59 | +/- | 0. |
| 1.19 | +/- | 0. |
| 1.19 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 1.89 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.04 |
| -9999.00 |

BRIGHT_NEIGHBOR
120-61-O
| 5542. | +/- | 9. |
| 5500. | +/- | 106. |
| 4.18 | +/- | 0. |
| 4.05 | +/- | 0. |
| 0.70 | +/- | 0. |
| 0.70 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 3.51 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.13 |
| -9999.00 |
STAR_WARN,SN_WARN
120-61-O
| 4384. | +/- | 11. |
| 4486. | +/- | 109. |
| 4.40 | +/- | 0. |
| 4.69 | +/- | 0. |
| 0.74 | +/- | 0. |
| 0.74 | +/- | -NaN |
| -0.30 | +/- | 0. |
| -0.30 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 5.98 | +/- | 0. |
| 0.78 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -0.52 |
| -9999.00 |
120-61-O
| 4836. | +/- | 7. |
| 4897. | +/- | 94. |
| 3.21 | +/- | 0. |
| 3.06 | +/- | 0. |
| 1.38 | +/- | 0. |
| 1.38 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -0.25 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.07 | +/- | 0. |
| 3.42 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.56 |
| -9999.00 |

120-61-O
| 5763. | +/- | 12. |
| 5716. | +/- | 118. |
| 4.08 | +/- | 0. |
| 4.10 | +/- | 0. |
| 1.16 | +/- | 0. |
| 1.16 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -0.27 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.07 | +/- | 0. |
| 3.48 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.03 |
| -9999.00 |

120-61-O
| 5406. | +/- | 35. |
| 5452. | +/- | 154. |
| 3.34 | +/- | 0. |
| 3.37 | +/- | 0. |
| 0.48 | +/- | 0. |
| 0.48 | +/- | -NaN |
| -1.48 | +/- | 0. |
| -1.48 | +/- | 0. |
| 0.80 | +/- | 0. |
| 0.80 | +/- | -NaN |
| 0.52 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.22 | +/- | 0. |
| 7.04 | +/- | 0. |
| 0.85 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.89 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -2.19 |
| -9999.00 |
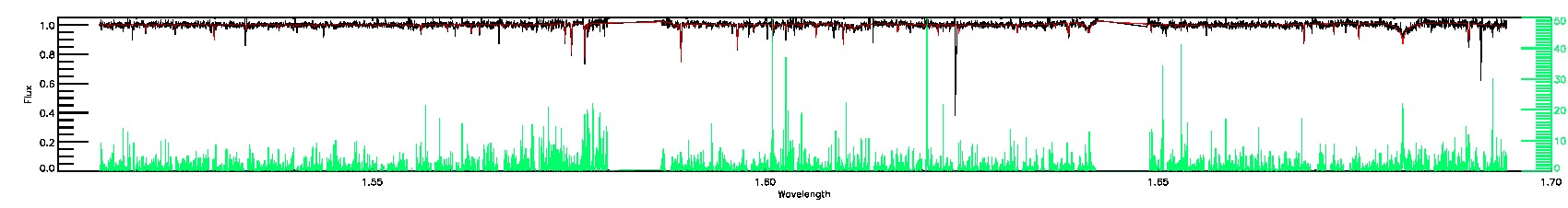
STAR_WARN,SN_WARN
120-61-O
| 4712. | +/- | 13. |
| 4790. | +/- | 117. |
| 4.46 | +/- | 0. |
| 4.70 | +/- | 0. |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| -0.32 | +/- | 0. |
| -0.32 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.14 | +/- | 0. |
| 3.46 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.26 |
| -9999.00 |

120-61-O
| 5764. | +/- | 20. |
| 5706. | +/- | 139. |
| 4.35 | +/- | 0. |
| 4.26 | +/- | 0. |
| 1.13 | +/- | 0. |
| 1.13 | +/- | -NaN |
| -0.00 | +/- | 0. |
| 0.00 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 1.69 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.05 |
| -9999.00 |

120-61-O
| 6220. | +/- | 18. |
| 6118. | +/- | 134. |
| 4.33 | +/- | 0. |
| 4.24 | +/- | 0. |
| 1.52 | +/- | 0. |
| 1.52 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.13 | +/- | 0. |
| -0.13 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 5.04 | +/- | 0. |
| 0.70 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.17 |
| -9999.00 |
120-61-O
| 4530. | +/- | 2. |
| 4616. | +/- | 79. |
| 2.53 | +/- | 0. |
| 2.29 | +/- | 0. |
| 1.32 | +/- | 0. |
| 1.32 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | 0. |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.11 | +/- | 0. |
| 2.97 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.21 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.13 |
| -9999.00 |

120-61-O
| 5225. | +/- | 21. |
| 5228. | +/- | 129. |
| 4.41 | +/- | 0. |
| 4.41 | +/- | 0. |
| 0.89 | +/- | 0. |
| 0.89 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 7.26 | +/- | 0. |
| 0.86 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.22 |
| -9999.00 |

120-61-O
| 4802. | +/- | 4. |
| 4849. | +/- | 85. |
| 3.34 | +/- | 0. |
| 3.16 | +/- | 0. |
| 1.04 | +/- | 0. |
| 1.04 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.70 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.20 |
| -9999.00 |
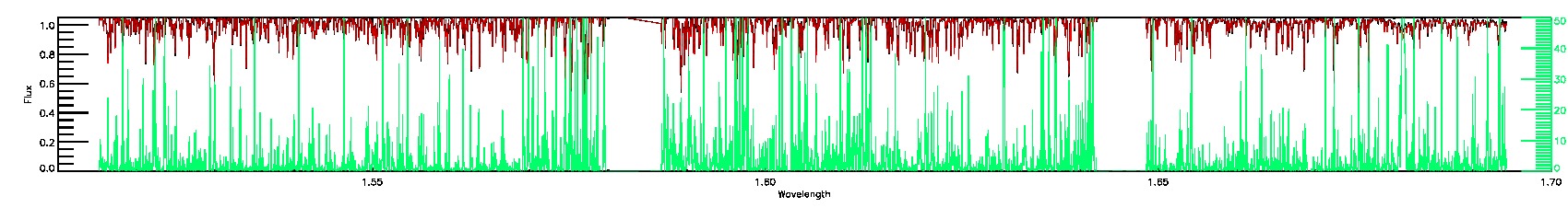
STAR_WARN,COLORTE_WARN
120-61-O
| 3752. | +/- | 4. |
| 3853. | +/- | 82. |
| 4.48 | +/- | 0. |
| 4.90 | +/- | 0. |
| 1.05 | +/- | 0. |
| 1.05 | +/- | -NaN |
| -0.30 | +/- | 0. |
| -0.30 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -0.20 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 1.78 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.13 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_WARN,COLORTE_WARN,SN_WARN
120-61-O
| 3743. | +/- | 8. |
| 3841. | +/- | 87. |
| 4.49 | +/- | 0. |
| 4.87 | +/- | 0. |
| 0.84 | +/- | 0. |
| 0.84 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -0.26 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 11.55 | +/- | 0. |
| 1.06 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.61 |
| -9999.00 |
STAR_WARN,SN_WARN
120-61-O
| 5684. | +/- | 34. |
| 5631. | +/- | 153. |
| 4.55 | +/- | 0. |
| 4.44 | +/- | 0. |
| 0.96 | +/- | 0. |
| 0.96 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 1.69 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.14 |
| -9999.00 |
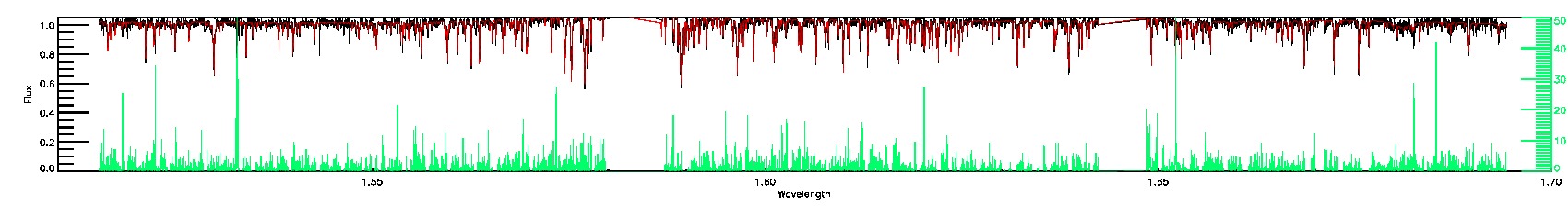
120-61-O
| 6245. | +/- | 28. |
| 6135. | +/- | 165. |
| 4.70 | +/- | 0. |
| 4.56 | +/- | 0. |
| 0.46 | +/- | 0. |
| 0.46 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.09 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.45 |
| -9999.00 |
120-61-O
| 4352. | +/- | 2. |
| 4467. | +/- | 81. |
| 2.21 | +/- | 0. |
| 2.05 | +/- | 0. |
| 1.49 | +/- | 0. |
| 1.49 | +/- | -NaN |
| -0.61 | +/- | 0. |
| -0.61 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.25 | +/- | 0. |
| 4.23 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -1.48 |
| -9999.00 |
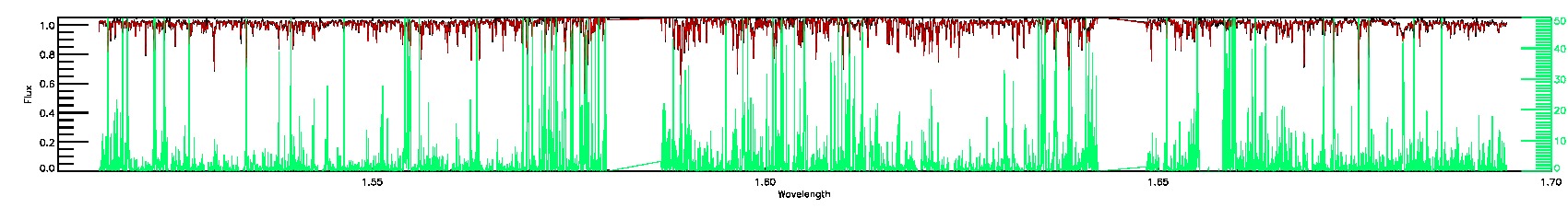
STAR_WARN,SN_WARN
120-61-O
| 5747. | +/- | 53. |
| 5691. | +/- | 160. |
| 4.44 | +/- | 0. |
| 4.35 | +/- | 0. |
| 1.39 | +/- | 0. |
| 1.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.01 | +/- | 0. |
| -0.20 | +/- | 0. |
| -0.20 | +/- | -NaN |
| 0.45 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 2.24 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.14 |
| -9999.00 |
BRIGHT_NEIGHBOR
120-61-O
| 5542. | +/- | 16. |
| 5501. | +/- | 130. |
| 4.43 | +/- | 0. |
| 4.31 | +/- | 0. |
| 0.76 | +/- | 0. |
| 0.76 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 1.74 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.27 |
| -9999.00 |
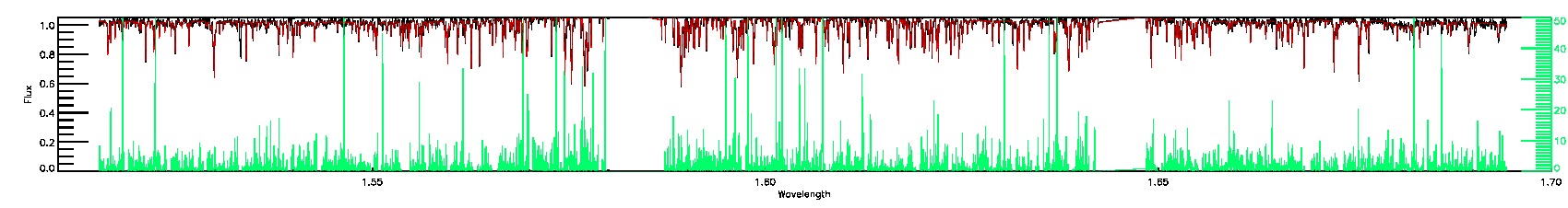
120-61-O
| 5552. | +/- | 8. |
| 5512. | +/- | 106. |
| 4.67 | +/- | 0. |
| 4.56 | +/- | 0. |
| 1.55 | +/- | 0. |
| 1.55 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.05 | +/- | 0. |
| 5.14 | +/- | 0. |
| 0.71 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.03 |
| -9999.00 |

BRIGHT_NEIGHBOR
120-61-O
| 4708. | +/- | 11. |
| 4777. | +/- | 110. |
| 4.52 | +/- | 0. |
| 4.68 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.43 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.11 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -0.14 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 1.52 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.23 |
| -9999.00 |

120-61-O
| 4983. | +/- | 6. |
| 5017. | +/- | 91. |
| 4.49 | +/- | 0. |
| 4.55 | +/- | 0. |
| 1.34 | +/- | 0. |
| 1.34 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 1.59 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |

BRIGHT_NEIGHBOR
STAR_WARN,SN_WARN
120-61-O
| 5094. | +/- | 22. |
| 5106. | +/- | 126. |
| 4.54 | +/- | 0. |
| 4.51 | +/- | 0. |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.16 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.61 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.82 |
| -9999.00 |

BRIGHT_NEIGHBOR
120-61-O
| 5750. | +/- | 38. |
| 5698. | +/- | 153. |
| 4.14 | +/- | 0. |
| 4.10 | +/- | 0. |
| 1.38 | +/- | 0. |
| 1.38 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.11 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| 0.84 |
| -9999.00 |
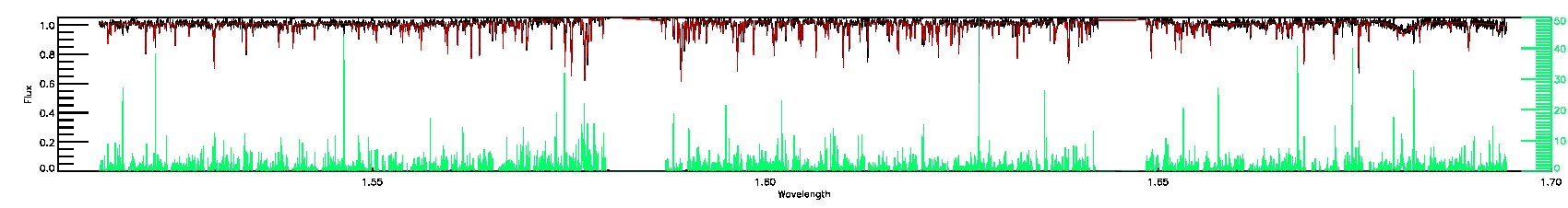
SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,COLORTE_WARN
120-61-O
| 6168. | +/- | 17. |
| -10000. | +/- | -NaN |
| 3.63 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.55 | +/- | 0. |
| 0.55 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 94.91 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.36 |
| -9999.00 |
120-61-O
| 5781. | +/- | 26. |
| 5725. | +/- | 149. |
| 4.64 | +/- | 0. |
| 4.59 | +/- | 0. |
| 1.40 | +/- | 0. |
| 1.40 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 2.19 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.64 |
| -9999.00 |

120-61-O
| 5235. | +/- | 12. |
| 5239. | +/- | 119. |
| 4.50 | +/- | 0. |
| 4.51 | +/- | 0. |
| 0.69 | +/- | 0. |
| 0.69 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 1.83 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.08 |
| -9999.00 |

120-61-O
| 5678. | +/- | 14. |
| 5642. | +/- | 115. |
| 4.47 | +/- | 0. |
| 4.51 | +/- | 0. |
| 1.08 | +/- | 0. |
| 1.08 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -0.27 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 2.11 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.27 |
| -9999.00 |

STAR_WARN,SN_WARN
120-61-O
| 4288. | +/- | 10. |
| 4394. | +/- | 109. |
| 4.21 | +/- | 0. |
| 4.56 | +/- | 0. |
| 0.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -0.40 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| -0.30 | +/- | 0. |
| -0.30 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 6.62 | +/- | 0. |
| 0.82 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -2.14 |
| -9999.00 |
| -0.68 |
| -9999.00 |

BRIGHT_NEIGHBOR
120-61-O
| 5142. | +/- | 9. |
| 5156. | +/- | 107. |
| 4.44 | +/- | 0. |
| 4.47 | +/- | 0. |
| 0.54 | +/- | 0. |
| 0.54 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.78 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
120-61-O
| 7781. | +/- | 12. |
| 7653. | +/- | 254. |
| 4.69 | +/- | 0. |
| 4.97 | +/- | 0. |
| 0.46 | +/- | 0. |
| 0.46 | +/- | -NaN |
| -1.72 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| 0.01 | +/- | 1. |
| 0.01 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.57 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -1.36 |
| -9999.00 |
120-61-O
| 5983. | +/- | 21. |
| 5905. | +/- | 138. |
| 4.54 | +/- | 0. |
| 4.46 | +/- | 0. |
| 0.70 | +/- | 0. |
| 0.70 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -0.14 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 9.43 | +/- | 0. |
| 0.97 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.23 |
| -9999.00 |
120-61-O
| 4704. | +/- | 9. |
| 4779. | +/- | 107. |
| 3.07 | +/- | 0. |
| 2.91 | +/- | 0. |
| 1.11 | +/- | 0. |
| 1.11 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -0.25 | +/- | 0. |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.18 | +/- | 0. |
| 3.43 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.53 |
| -9999.00 |

120-61-O
| 6439. | +/- | 17. |
| 6302. | +/- | 140. |
| 4.62 | +/- | 0. |
| 4.39 | +/- | 0. |
| 1.13 | +/- | 0. |
| 1.13 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 13.66 | +/- | 0. |
| 1.14 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
120-61-O
| 7053. | +/- | 18. |
| 6857. | +/- | 178. |
| 4.70 | +/- | 0. |
| 4.42 | +/- | 0. |
| 1.31 | +/- | 0. |
| 1.31 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.10 | +/- | 0. |
| 12.76 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.52 |
| -9999.00 |
STAR_BAD
120-61-O
| 3689. | +/- | 8. |
| -10000. | +/- | -NaN |
| 5.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.53 | +/- | 0. |
| 0.53 | +/- | -NaN |
| -0.81 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.41 | +/- | 0. |
| 0.92 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.96 |
| -9999.00 |
120-61-O
| 5728. | +/- | 19. |
| 5689. | +/- | 123. |
| 4.47 | +/- | 0. |
| 4.54 | +/- | 0. |
| 0.46 | +/- | 0. |
| 0.46 | +/- | -NaN |
| -0.37 | +/- | 0. |
| -0.37 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -0.31 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.14 | +/- | 0. |
| 10.27 | +/- | 0. |
| 1.01 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.71 |
| -9999.00 |
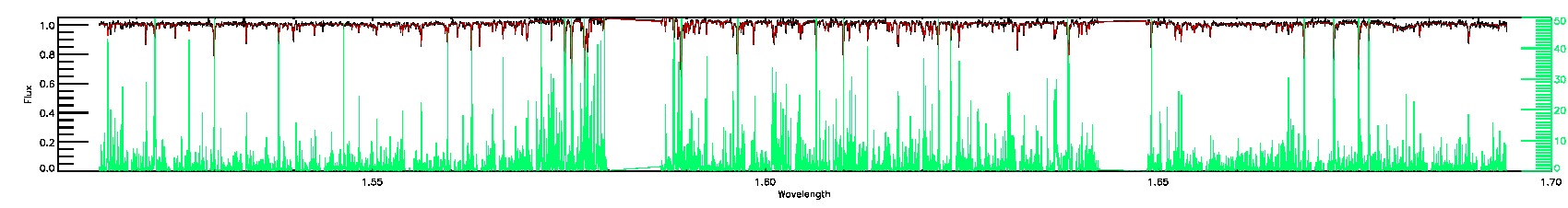
120-61-O
| 4521. | +/- | 8. |
| 4610. | +/- | 106. |
| 4.50 | +/- | 0. |
| 4.68 | +/- | 0. |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.98 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.08 |
| -9999.00 |

120-61-O
| 5654. | +/- | 17. |
| 5629. | +/- | 120. |
| 3.86 | +/- | 0. |
| 3.92 | +/- | 0. |
| 1.34 | +/- | 0. |
| 1.34 | +/- | -NaN |
| -0.46 | +/- | 0. |
| -0.46 | +/- | 0. |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 2.88 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.33 |
| -9999.00 |

120-61-O
| 3865. | +/- | 4. |
| 3957. | +/- | 84. |
| 4.46 | +/- | 0. |
| 4.76 | +/- | 0. |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -0.22 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 1.83 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
120-61-O
| 4344. | +/- | 3. |
| 4439. | +/- | 82. |
| 4.21 | +/- | 0. |
| 4.44 | +/- | 0. |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.14 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 6.57 | +/- | 0. |
| 0.82 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.03 |
| -9999.00 |

120-61-O
| 6105. | +/- | 42. |
| 6025. | +/- | 165. |
| 4.39 | +/- | 0. |
| 4.41 | +/- | 0. |
| 1.18 | +/- | 0. |
| 1.18 | +/- | -NaN |
| -0.43 | +/- | 0. |
| -0.42 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.09 | +/- | 0. |
| 10.84 | +/- | 0. |
| 1.03 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.24 |
| -9999.00 |
120-61-O
| 5118. | +/- | 14. |
| 5142. | +/- | 117. |
| 4.43 | +/- | 0. |
| 4.54 | +/- | 0. |
| 0.74 | +/- | 0. |
| 0.74 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -0.18 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 4.28 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.04 |
| -9999.00 |

120-61-O
| 5142. | +/- | 16. |
| 5179. | +/- | 121. |
| 4.41 | +/- | 0. |
| 4.66 | +/- | 0. |
| 0.60 | +/- | 0. |
| 0.60 | +/- | -NaN |
| -0.55 | +/- | 0. |
| -0.55 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -0.20 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.14 | +/- | 0. |
| 1.63 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.88 |
| -9999.00 |
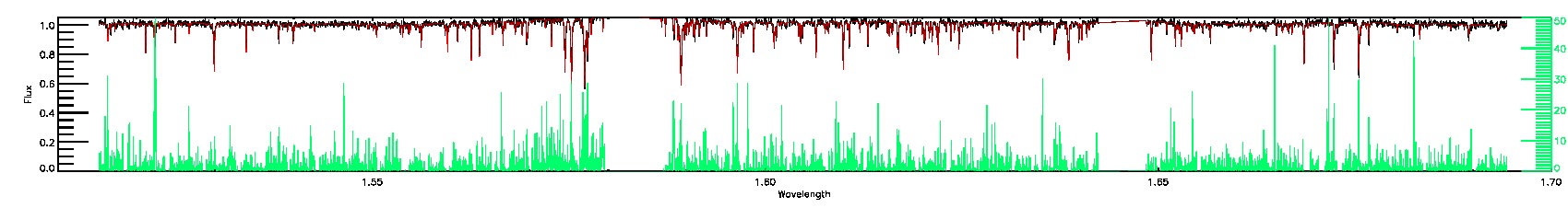
STAR_WARN,SN_WARN
120-61-O
| 4043. | +/- | 7. |
| 4136. | +/- | 93. |
| 4.40 | +/- | 0. |
| 4.67 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.06 | +/- | 0. |
| 3.24 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.13 |
| -9999.00 |

120-61-O
| 6241. | +/- | 18. |
| 6145. | +/- | 138. |
| 4.33 | +/- | 0. |
| 4.32 | +/- | 0. |
| 0.43 | +/- | 0. |
| 0.43 | +/- | -NaN |
| -0.41 | +/- | 0. |
| -0.40 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 9.54 | +/- | 0. |
| 0.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -1.01 |
| -9999.00 |
STAR_WARN,SN_WARN
120-61-O
| 5131. | +/- | 22. |
| 5153. | +/- | 129. |
| 3.90 | +/- | 0. |
| 3.95 | +/- | 0. |
| 1.04 | +/- | 0. |
| 1.04 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -0.16 | +/- | 0. |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.14 | +/- | 0. |
| 3.26 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.21 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -2.42 |
| -9999.00 |

BRIGHT_NEIGHBOR
120-61-O
| 5503. | +/- | 9. |
| 5463. | +/- | 110. |
| 4.45 | +/- | 0. |
| 4.29 | +/- | 0. |
| 0.57 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.29 | +/- | 0. |
| -0.14 | +/- | 0. |
| -0.14 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 3.16 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.14 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.22 |
| -9999.00 |

120-61-O
| 4508. | +/- | 2. |
| 4593. | +/- | 78. |
| 2.65 | +/- | 0. |
| 2.35 | +/- | 0. |
| 1.27 | +/- | 0. |
| 1.27 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.09 | +/- | 0. |
| 2.84 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.12 |
| -9999.00 |

120-61-O
| 5752. | +/- | 22. |
| 5716. | +/- | 129. |
| 3.95 | +/- | 0. |
| 4.04 | +/- | 0. |
| 1.17 | +/- | 0. |
| 1.17 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -0.49 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.04 | +/- | 0. |
| 5.77 | +/- | 0. |
| 0.76 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -1.28 |
| -9999.00 |

120-61-O
| 6166. | +/- | 14. |
| 6058. | +/- | 128. |
| 4.29 | +/- | 0. |
| 4.09 | +/- | 0. |
| 1.37 | +/- | 0. |
| 1.37 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 6.63 | +/- | 0. |
| 0.82 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.30 |
| -9999.00 |
STAR_WARN,SN_WARN
120-61-O
| 5868. | +/- | 47. |
| 5808. | +/- | 166. |
| 4.52 | +/- | 0. |
| 4.50 | +/- | 0. |
| 0.58 | +/- | 0. |
| 0.58 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -0.23 | +/- | 0. |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.10 | +/- | 0. |
| 1.64 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -0.47 |
| -9999.00 |
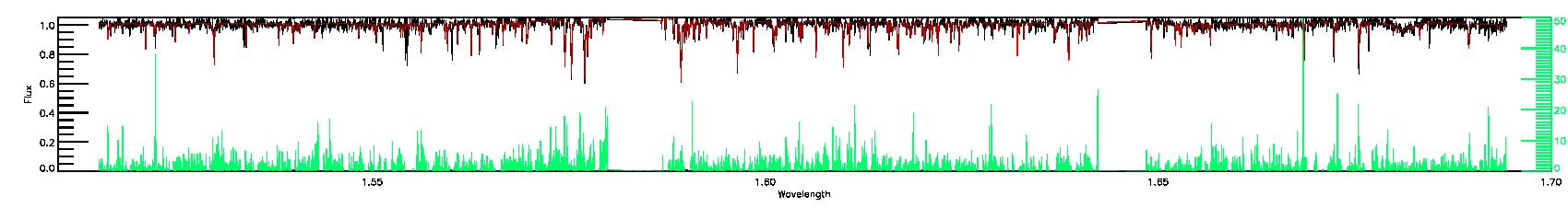
SUSPECT_BROAD_LINES
STAR_WARN,SN_WARN
120-61-O
| 3709. | +/- | 9. |
| 3804. | +/- | 87. |
| 4.40 | +/- | 0. |
| 4.75 | +/- | 0. |
| 1.23 | +/- | 0. |
| 1.23 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 21.04 | +/- | 0. |
| 1.32 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.36 |
| -9999.00 |
STAR_WARN,SN_WARN
120-61-O
| 4955. | +/- | 19. |
| 4983. | +/- | 121. |
| 4.66 | +/- | 0. |
| 4.64 | +/- | 0. |
| 1.68 | +/- | 0. |
| 1.68 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.19 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 1.65 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
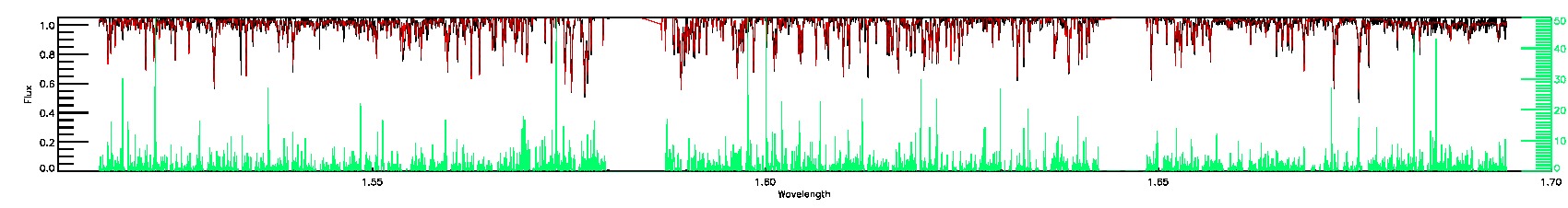
120-61-O
| 4059. | +/- | 5. |
| 4151. | +/- | 92. |
| 4.36 | +/- | 0. |
| 4.62 | +/- | 0. |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.07 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.02 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_WARN,SN_WARN
120-61-O
| 4993. | +/- | 23. |
| 5020. | +/- | 123. |
| 4.60 | +/- | 0. |
| 4.61 | +/- | 0. |
| 1.51 | +/- | 0. |
| 1.51 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 18.38 | +/- | 0. |
| 1.26 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.49 |
| -9999.00 |

120-61-O
| 5436. | +/- | 24. |
| 5420. | +/- | 137. |
| 4.49 | +/- | 0. |
| 4.50 | +/- | 0. |
| 1.24 | +/- | 0. |
| 1.24 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.35 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.49 |
| -9999.00 |
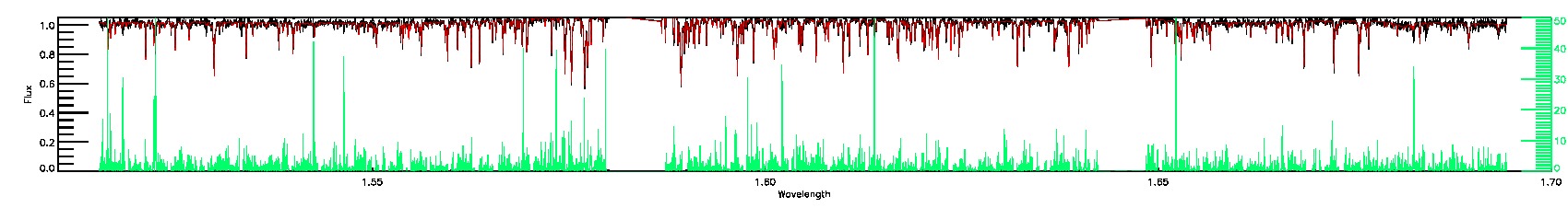
BRIGHT_NEIGHBOR
120-61-O
| 4807. | +/- | 17. |
| 4925. | +/- | 130. |
| 2.26 | +/- | 0. |
| 2.23 | +/- | 0. |
| 0.61 | +/- | 0. |
| 0.61 | +/- | -NaN |
| -1.53 | +/- | 0. |
| -1.53 | +/- | 0. |
| 0.82 | +/- | 0. |
| 0.82 | +/- | -NaN |
| -0.30 | +/- | 0. |
| -0.30 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.28 | +/- | 0. |
| 7.22 | +/- | 0. |
| 0.86 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.93 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -2.19 |
| -9999.00 |
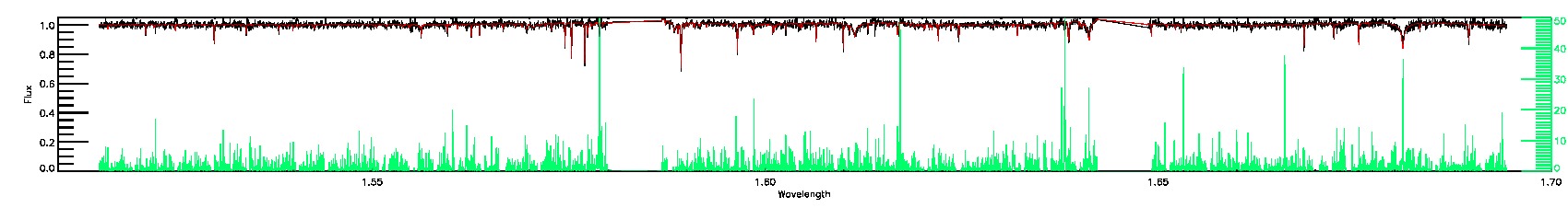
120-61-O
| 6275. | +/- | 15. |
| 6163. | +/- | 135. |
| 4.66 | +/- | 0. |
| 4.52 | +/- | 0. |
| 0.47 | +/- | 0. |
| 0.47 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.56 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 4.25 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.16 |
| -9999.00 |
120-61-O
| 4959. | +/- | 9. |
| 4989. | +/- | 108. |
| 4.51 | +/- | 0. |
| 4.51 | +/- | 0. |
| 0.42 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.14 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 1.69 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.30 |
| -9999.00 |

STAR_WARN,SN_WARN
120-61-O
| 5191. | +/- | 23. |
| 5195. | +/- | 130. |
| 4.32 | +/- | 0. |
| 4.30 | +/- | 0. |
| 0.37 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 6.63 | +/- | 0. |
| 0.82 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.14 |
| -9999.00 |

120-61-O
| 5577. | +/- | 25. |
| 5539. | +/- | 143. |
| 4.68 | +/- | 0. |
| 4.61 | +/- | 0. |
| 0.76 | +/- | 0. |
| 0.76 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 1.91 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| 0.03 |
| -9999.00 |
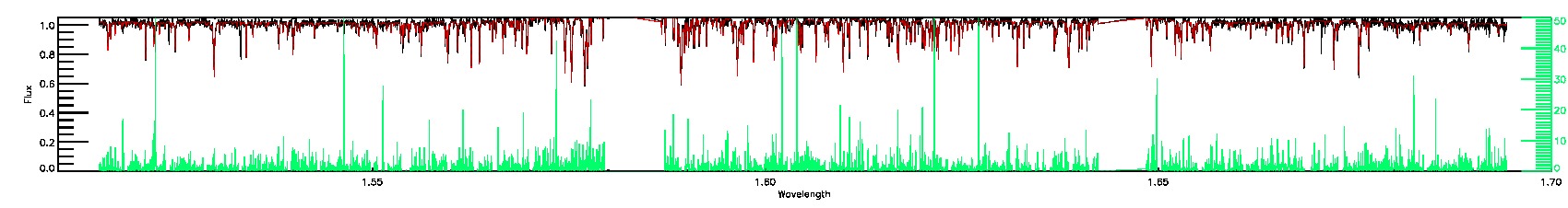
120-61-O
| 5166. | +/- | 9. |
| 5181. | +/- | 107. |
| 4.48 | +/- | 0. |
| 4.53 | +/- | 0. |
| 1.61 | +/- | 0. |
| 1.61 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 1.68 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.31 |
| -9999.00 |
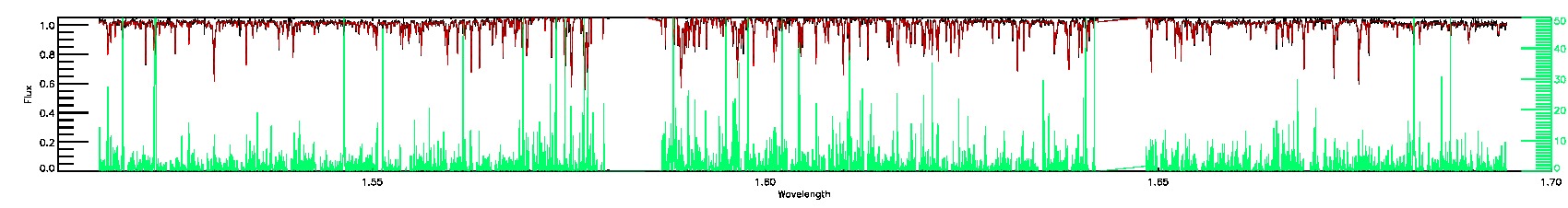
STAR_WARN,SN_WARN
120-61-O
| 5775. | +/- | 65. |
| 5738. | +/- | 166. |
| 4.18 | +/- | 0. |
| 4.29 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| -0.51 | +/- | 0. |
| -0.51 | +/- | 0. |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.07 | +/- | 0. |
| 4.00 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.20 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.67 |
| -9999.00 |
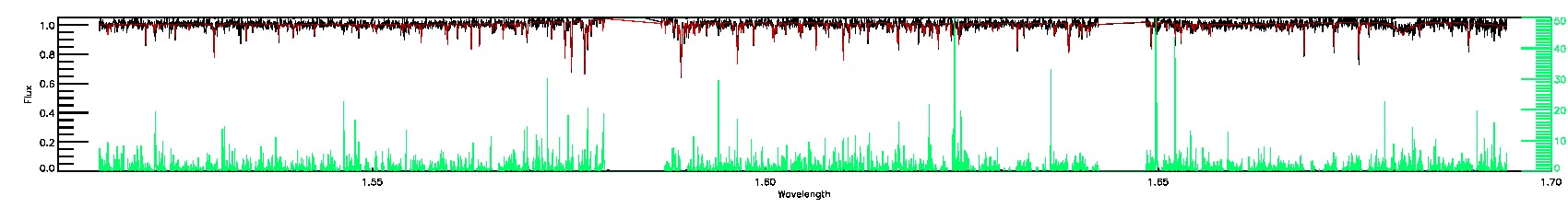
STAR_WARN,SN_WARN
120-61-O
| 5777. | +/- | 40. |
| 5724. | +/- | 159. |
| 4.39 | +/- | 0. |
| 4.36 | +/- | 0. |
| 0.81 | +/- | 0. |
| 0.81 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.17 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.45 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.20 |
| -9999.00 |

120-61-O
| 5293. | +/- | 14. |
| 5280. | +/- | 124. |
| 4.31 | +/- | 0. |
| 4.22 | +/- | 0. |
| 0.44 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 1.81 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| 0.30 |
| -9999.00 |

120-61-O
| 5730. | +/- | 22. |
| 5678. | +/- | 139. |
| 4.41 | +/- | 0. |
| 4.36 | +/- | 0. |
| 1.27 | +/- | 0. |
| 1.27 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.16 | +/- | 0. |
| -0.16 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.61 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.02 |
| -9999.00 |
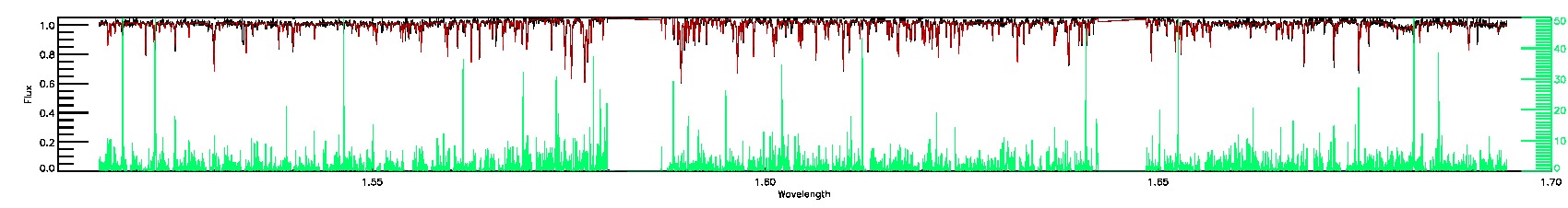
SUSPECT_BROAD_LINES
120-61-O
| 6395. | +/- | 16. |
| 6264. | +/- | 138. |
| 4.15 | +/- | 0. |
| 3.94 | +/- | 0. |
| 1.95 | +/- | 0. |
| 1.95 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.06 | +/- | 0. |
| 31.22 | +/- | 0. |
| 1.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.21 |
| -9999.00 |
120-61-O
| 5493. | +/- | 7. |
| 5452. | +/- | 102. |
| 4.28 | +/- | 0. |
| 4.11 | +/- | 0. |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | 0. |
| -0.13 | +/- | 0. |
| -0.13 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.11 | +/- | 0. |
| 0.33 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.32 |
| -9999.00 |

STAR_WARN,SN_WARN
120-61-O
| 6282. | +/- | 57. |
| 6172. | +/- | 183. |
| 4.57 | +/- | 0. |
| 4.46 | +/- | 0. |
| 0.78 | +/- | 0. |
| 0.78 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -0.19 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 7.35 | +/- | 0. |
| 0.87 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.76 |
| -9999.00 |
STAR_WARN,SN_WARN
120-61-O
| 4373. | +/- | 7. |
| 4454. | +/- | 99. |
| 4.51 | +/- | 0. |
| 4.61 | +/- | 0. |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.07 | +/- | 0. |
| 3.21 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.45 |
| -9999.00 |

120-61-O
| 4879. | +/- | 15. |
| 4928. | +/- | 118. |
| 3.42 | +/- | 0. |
| 3.27 | +/- | 0. |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.13 | +/- | 0. |
| 3.13 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.13 |
| -9999.00 |
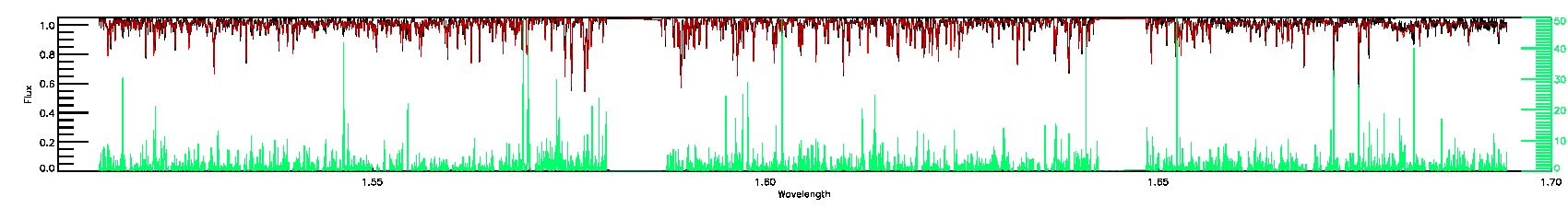
STAR_WARN,SN_WARN
120-61-O
| 4951. | +/- | 17. |
| 4985. | +/- | 120. |
| 4.64 | +/- | 0. |
| 4.68 | +/- | 0. |
| 1.58 | +/- | 0. |
| 1.58 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 3.70 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.07 |
| -9999.00 |
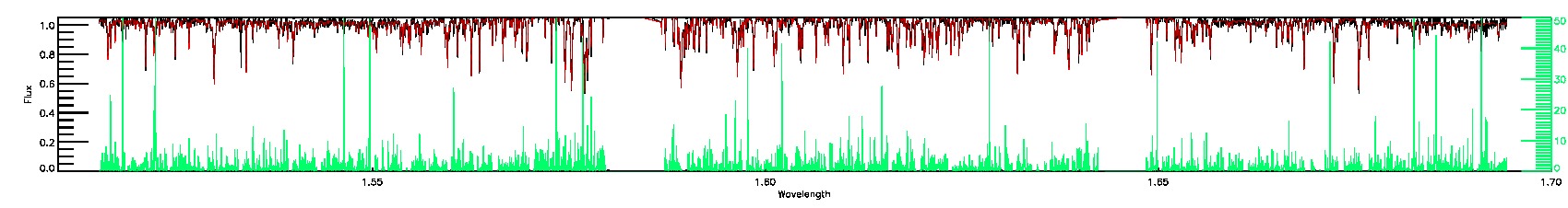
STAR_WARN,SN_WARN
120-61-O
| 5054. | +/- | 25. |
| 5076. | +/- | 128. |
| 4.43 | +/- | 0. |
| 4.45 | +/- | 0. |
| 0.43 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.79 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.10 |
| -9999.00 |

120-61-O
| 5854. | +/- | 20. |
| 5802. | +/- | 134. |
| 4.37 | +/- | 0. |
| 4.42 | +/- | 0. |
| 0.88 | +/- | 0. |
| 0.88 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -0.40 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 4.91 | +/- | 0. |
| 0.69 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.44 |
| -9999.00 |

SUSPECT_RV_COMBINATION
120-61-O
| 5419. | +/- | 43. |
| 5429. | +/- | 146. |
| 4.19 | +/- | 0. |
| 4.44 | +/- | 0. |
| 0.82 | +/- | 0. |
| 0.82 | +/- | -NaN |
| -0.67 | +/- | 0. |
| -0.67 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.17 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.15 | +/- | 0. |
| 3.87 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -2.21 |
| -9999.00 |

120-61-O
| 4179. | +/- | 2. |
| 4276. | +/- | 73. |
| 4.28 | +/- | 0. |
| 4.57 | +/- | 0. |
| 0.46 | +/- | 0. |
| 0.46 | +/- | -NaN |
| -0.21 | +/- | 0. |
| -0.21 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -0.20 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 6.28 | +/- | 0. |
| 0.80 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.11 |
| -9999.00 |

120-61-O
| 5758. | +/- | 39. |
| 5717. | +/- | 156. |
| 4.12 | +/- | 0. |
| 4.19 | +/- | 0. |
| 0.69 | +/- | 0. |
| 0.69 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -0.40 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.14 | +/- | 0. |
| 3.61 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -2.42 |
| -9999.00 |
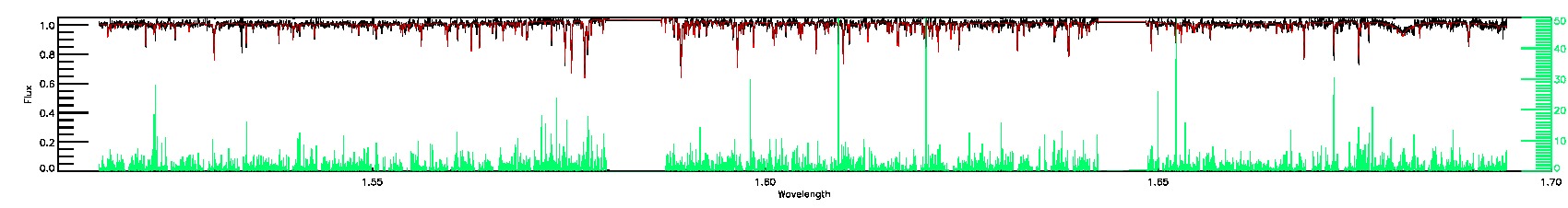
120-61-O
| 5883. | +/- | 19. |
| 5811. | +/- | 141. |
| 4.44 | +/- | 0. |
| 4.33 | +/- | 0. |
| 0.97 | +/- | 0. |
| 0.97 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | 0. |
| -0.13 | +/- | 0. |
| -0.13 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.01 | +/- | 0. |
| 11.15 | +/- | 0. |
| 1.05 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.16 |
| -9999.00 |

STAR_WARN,SN_WARN
120-61-O
| 5539. | +/- | 49. |
| 5515. | +/- | 151. |
| 4.20 | +/- | 0. |
| 4.23 | +/- | 0. |
| 1.08 | +/- | 0. |
| 1.08 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -0.19 | +/- | 0. |
| -0.31 | +/- | 0. |
| -0.31 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 5.38 | +/- | 0. |
| 0.73 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.26 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| 0.00 |
| -9999.00 |

120-61-O
| 5090. | +/- | 13. |
| 5121. | +/- | 114. |
| 3.65 | +/- | 0. |
| 3.62 | +/- | 0. |
| 1.18 | +/- | 0. |
| 1.18 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -0.28 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.47 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.56 |
| -9999.00 |
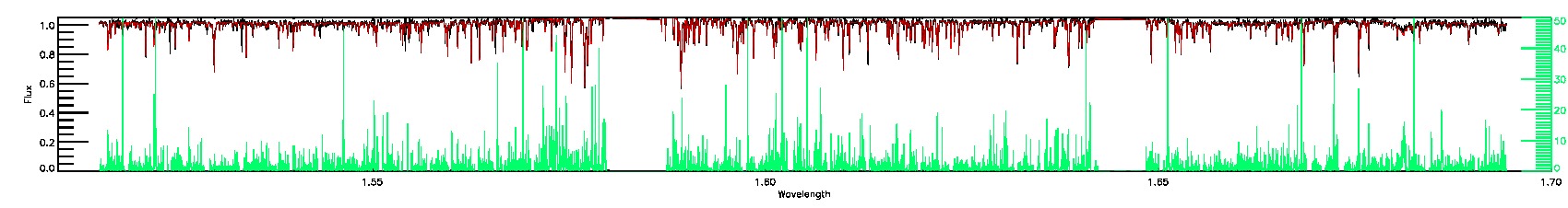
STAR_WARN,SN_WARN
120-61-O
| 4768. | +/- | 16. |
| 4827. | +/- | 117. |
| 4.56 | +/- | 0. |
| 4.68 | +/- | 0. |
| 0.98 | +/- | 0. |
| 0.98 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -0.14 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 1.96 | +/- | 0. |
| 0.29 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.10 |
| -9999.00 |

120-61-O
| 5311. | +/- | 15. |
| 5301. | +/- | 125. |
| 4.49 | +/- | 0. |
| 4.44 | +/- | 0. |
| 1.31 | +/- | 0. |
| 1.31 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.10 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 1.52 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.51 |
| -9999.00 |
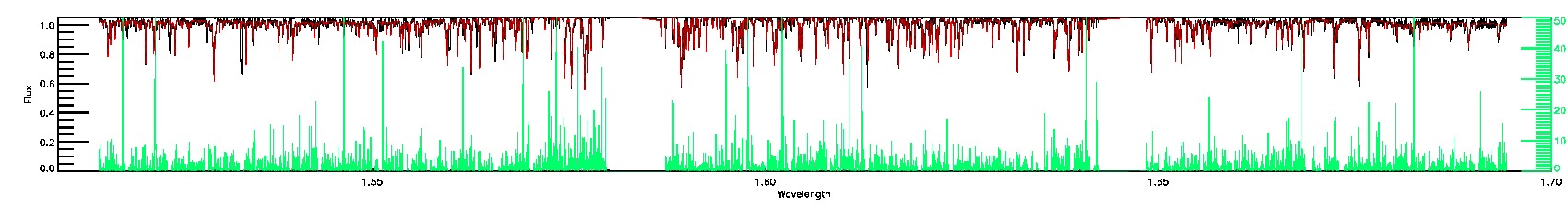
SUSPECT_BROAD_LINES
120-61-O
| 4260. | +/- | 6. |
| 4335. | +/- | 92. |
| 4.46 | +/- | 0. |
| 4.52 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.09 | +/- | 0. |
| 24.61 | +/- | 0. |
| 1.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 0.22 |
| -9999.00 |
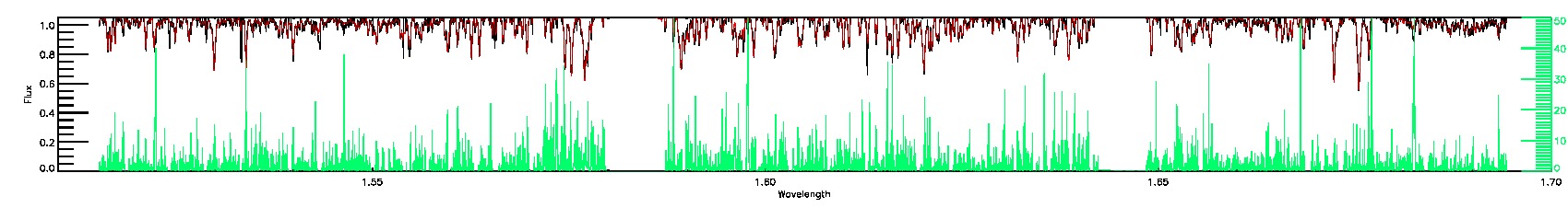
120-61-O
| 5394. | +/- | 28. |
| 5394. | +/- | 137. |
| 4.41 | +/- | 0. |
| 4.54 | +/- | 0. |
| 0.80 | +/- | 0. |
| 0.80 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -0.35 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.27 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.99 |
| -9999.00 |

120-61-O
| 4559. | +/- | 6. |
| 4628. | +/- | 96. |
| 4.53 | +/- | 0. |
| 4.54 | +/- | 0. |
| 0.76 | +/- | 0. |
| 0.76 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.10 | +/- | 0. |
| 1.64 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.31 |
| -9999.00 |

SUSPECT_BROAD_LINES
120-61-O
| 3763. | +/- | 3. |
| 3853. | +/- | 63. |
| 4.47 | +/- | 0. |
| 4.76 | +/- | 0. |
| 1.41 | +/- | 0. |
| 1.41 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 68.19 | +/- | 0. |
| 1.83 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.01 |
| -9999.00 |
120-61-O
| 5755. | +/- | 29. |
| 5702. | +/- | 147. |
| 4.33 | +/- | 0. |
| 4.28 | +/- | 0. |
| 1.10 | +/- | 0. |
| 1.10 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.55 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.33 |
| -9999.00 |

120-61-O
| 5675. | +/- | 16. |
| 5632. | +/- | 133. |
| 4.53 | +/- | 0. |
| 4.51 | +/- | 0. |
| 0.52 | +/- | 0. |
| 0.52 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.12 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.10 | +/- | 0. |
| 1.93 | +/- | 0. |
| 0.29 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.16 |
| -9999.00 |

120-61-O
| 5021. | +/- | 13. |
| 5077. | +/- | 115. |
| 3.53 | +/- | 0. |
| 3.49 | +/- | 0. |
| 1.13 | +/- | 0. |
| 1.13 | +/- | -NaN |
| -0.66 | +/- | 0. |
| -0.66 | +/- | 0. |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -0.25 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.20 | +/- | 0. |
| 4.36 | +/- | 0. |
| 0.64 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.29 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -2.42 |
| -9999.00 |

120-61-O
| 4956. | +/- | 6. |
| 4987. | +/- | 92. |
| 3.72 | +/- | 0. |
| 3.64 | +/- | 0. |
| 1.13 | +/- | 0. |
| 1.13 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.79 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.24 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
120-61-O
| 3498. | +/- | 4. |
| 3582. | +/- | 71. |
| 4.56 | +/- | 0. |
| 4.86 | +/- | 0. |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.07 | +/- | 0. |
| 6.81 | +/- | 0. |
| 0.83 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.88 |
| -9999.00 |
120-61-O
| 3882. | +/- | 4. |
| 3990. | +/- | 89. |
| 4.43 | +/- | 0. |
| 4.88 | +/- | 0. |
| 1.13 | +/- | 0. |
| 1.13 | +/- | -NaN |
| -0.45 | +/- | 0. |
| -0.45 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.20 | +/- | 0. |
| 3.40 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.00 |
| -9999.00 |
STAR_WARN,SN_WARN
120-61-O
| 5392. | +/- | 42. |
| 5388. | +/- | 146. |
| 4.49 | +/- | 0. |
| 4.57 | +/- | 0. |
| 1.79 | +/- | 0. |
| 1.79 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -0.25 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.19 | +/- | 0. |
| 1.66 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.99 |
| -9999.00 |

120-61-O
| 5882. | +/- | 14. |
| 5823. | +/- | 123. |
| 4.19 | +/- | 0. |
| 4.20 | +/- | 0. |
| 1.07 | +/- | 0. |
| 1.07 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -0.28 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 1.82 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.35 |
| -9999.00 |
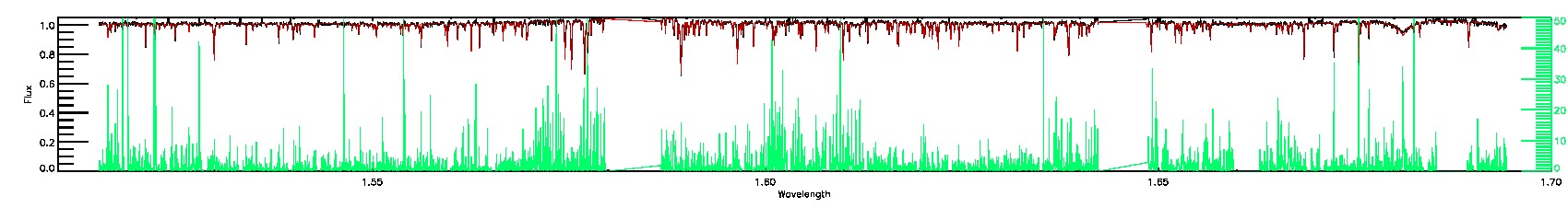
120-61-O
| 5932. | +/- | 30. |
| 5856. | +/- | 157. |
| 4.76 | +/- | 0. |
| 4.65 | +/- | 0. |
| 0.45 | +/- | 0. |
| 0.45 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.69 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.45 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
120-61-O
| 3463. | +/- | 4. |
| 3558. | +/- | 74. |
| 4.58 | +/- | 0. |
| 4.98 | +/- | 0. |
| 1.75 | +/- | 0. |
| 1.75 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.14 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 7.57 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.16 |
| -9999.00 |
120-61-O
| 4795. | +/- | 13. |
| 4851. | +/- | 112. |
| 4.51 | +/- | 0. |
| 4.62 | +/- | 0. |
| 0.77 | +/- | 0. |
| 0.77 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 1.68 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.08 |
| -9999.00 |
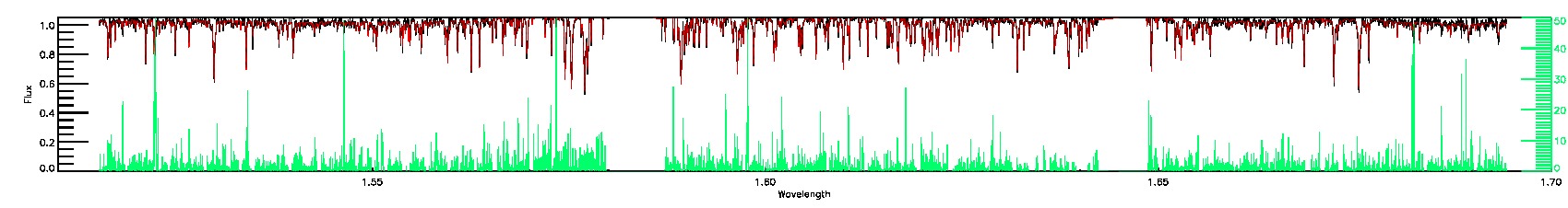
STAR_WARN,SN_WARN
120-61-O
| 4162. | +/- | 7. |
| 4256. | +/- | 97. |
| 4.30 | +/- | 0. |
| 4.56 | +/- | 0. |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.13 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -0.23 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.06 | +/- | 0. |
| 6.94 | +/- | 0. |
| 0.84 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -1.37 |
| -9999.00 |

120-61-O
| 4706. | +/- | 11. |
| 4769. | +/- | 109. |
| 4.42 | +/- | 0. |
| 4.51 | +/- | 0. |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.05 | +/- | 0. |
| 4.23 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.24 |
| -9999.00 |

STAR_WARN,SN_WARN
120-61-O
| 4403. | +/- | 8. |
| 4506. | +/- | 108. |
| 4.37 | +/- | 0. |
| 4.68 | +/- | 0. |
| 0.75 | +/- | 0. |
| 0.75 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -0.34 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.16 | +/- | 0. |
| 3.31 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.45 |
| -9999.00 |

STAR_WARN,SN_WARN
120-61-O
| 4356. | +/- | 11. |
| 4444. | +/- | 104. |
| 4.47 | +/- | 0. |
| 4.64 | +/- | 0. |
| 0.84 | +/- | 0. |
| 0.84 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 5.31 | +/- | 0. |
| 0.72 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.07 |
| -9999.00 |

STAR_WARN,SN_WARN
120-61-O
| 5312. | +/- | 18. |
| 5289. | +/- | 128. |
| 4.10 | +/- | 0. |
| 3.93 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.40 | +/- | 0. |
| 0.40 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 3.75 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.58 |
| -9999.00 |

120-61-O
| 5932. | +/- | 26. |
| 5865. | +/- | 144. |
| 4.15 | +/- | 0. |
| 4.13 | +/- | 0. |
| 0.88 | +/- | 0. |
| 0.88 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.23 | +/- | 0. |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.39 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.21 |
| -9999.00 |

STAR_WARN,SN_WARN
120-61-O
| 4904. | +/- | 27. |
| 4973. | +/- | 131. |
| 3.15 | +/- | 0. |
| 3.05 | +/- | 0. |
| 1.14 | +/- | 0. |
| 1.14 | +/- | -NaN |
| -0.64 | +/- | 0. |
| -0.64 | +/- | 0. |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.13 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.25 | +/- | 0. |
| 4.31 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -1.93 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -1.45 |
| -9999.00 |
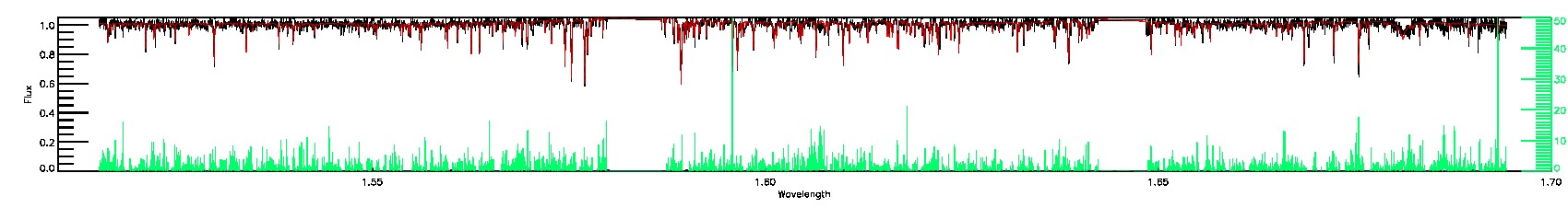
STAR_WARN,COLORTE_WARN
120-61-O
| 3709. | +/- | 2. |
| 3813. | +/- | 65. |
| 4.42 | +/- | 0. |
| 4.87 | +/- | 0. |
| 0.62 | +/- | 0. |
| 0.62 | +/- | -NaN |
| -0.38 | +/- | 0. |
| -0.37 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.12 | +/- | 0. |
| 3.65 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -0.36 |
| -9999.00 |
STAR_WARN,SN_WARN
120-61-O
| 5361. | +/- | 29. |
| 5340. | +/- | 136. |
| 4.26 | +/- | 0. |
| 4.15 | +/- | 0. |
| 0.50 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 3.14 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| 0.11 |
| -9999.00 |

120-61-O
| 5063. | +/- | 8. |
| 5069. | +/- | 106. |
| 4.38 | +/- | 0. |
| 4.26 | +/- | 0. |
| 0.42 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.40 | +/- | 0. |
| 0.40 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.07 | +/- | 0. |
| 1.92 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.22 |
| -9999.00 |

STAR_WARN,SN_WARN
120-61-O
| 4631. | +/- | 18. |
| 4712. | +/- | 117. |
| 4.49 | +/- | 0. |
| 4.68 | +/- | 0. |
| 0.84 | +/- | 0. |
| 0.84 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -0.17 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -0.14 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.59 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.39 |
| -9999.00 |

LOW_SNR
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
120-61-O
| 5516. | +/- | 92. |
| -10000. | +/- | -NaN |
| 4.57 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.86 | +/- | 0. |
| 1.86 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.26 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.78 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -9999.00 |
| -9999.00 |

STAR_BAD
STAR_WARN,COLORTE_WARN
120-61-O
| 3724. | +/- | 4. |
| -10000. | +/- | -NaN |
| 4.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.84 | +/- | 0. |
| 0.84 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.19 | +/- | 0. |
| 0.72 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.03 |
| -9999.00 |
STAR_WARN,SN_WARN
120-61-O
| 4523. | +/- | 9. |
| 4606. | +/- | 105. |
| 4.45 | +/- | 0. |
| 4.56 | +/- | 0. |
| 0.52 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 5.09 | +/- | 0. |
| 0.71 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.16 |
| -9999.00 |

SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES
120-61-O
| 5464. | +/- | 16. |
| 5451. | +/- | 108. |
| 4.54 | +/- | 0. |
| 4.61 | +/- | 0. |
| 1.14 | +/- | 0. |
| 1.14 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -0.25 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 19.36 | +/- | 0. |
| 1.29 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.21 |
| -9999.00 |

120-61-O
| 4654. | +/- | 8. |
| 4726. | +/- | 104. |
| 2.73 | +/- | 0. |
| 2.40 | +/- | 0. |
| 1.33 | +/- | 0. |
| 1.33 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.06 | +/- | 0. |
| 3.01 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -0.02 |
| -9999.00 |

STAR_WARN,SN_WARN
120-61-O
| 5826. | +/- | 33. |
| 5759. | +/- | 158. |
| 4.44 | +/- | 0. |
| 4.32 | +/- | 0. |
| 0.88 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.04 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.00 | +/- | 0. |
| 1.62 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| -1.14 |
| -9999.00 |

STAR_WARN,SN_WARN
120-61-O
| 5808. | +/- | 56. |
| 5756. | +/- | 166. |
| 4.54 | +/- | 0. |
| 4.55 | +/- | 0. |
| 0.88 | +/- | 0. |
| 0.88 | +/- | -NaN |
| -0.27 | +/- | 0. |
| -0.27 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.12 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.11 | +/- | 0. |
| 1.73 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.31 |
| -9999.00 |
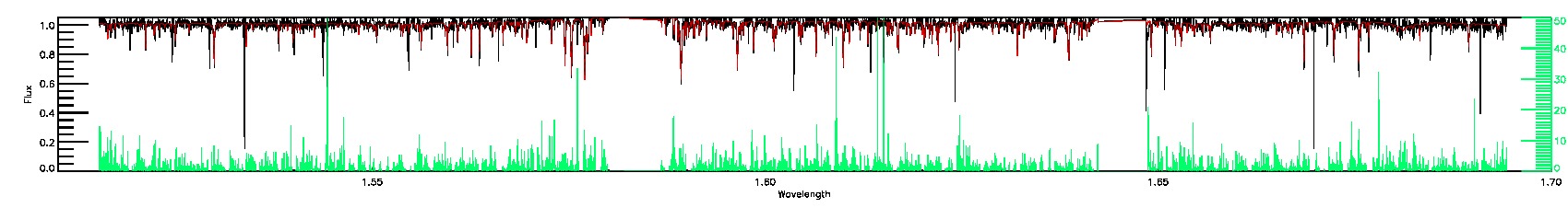
120-61-O
| 5227. | +/- | 12. |
| 5245. | +/- | 105. |
| 4.27 | +/- | 0. |
| 4.41 | +/- | 0. |
| 0.40 | +/- | 0. |
| 0.40 | +/- | -NaN |
| -0.32 | +/- | 0. |
| -0.32 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -0.23 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 5.56 | +/- | 0. |
| 0.74 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.14 |
| -9999.00 |
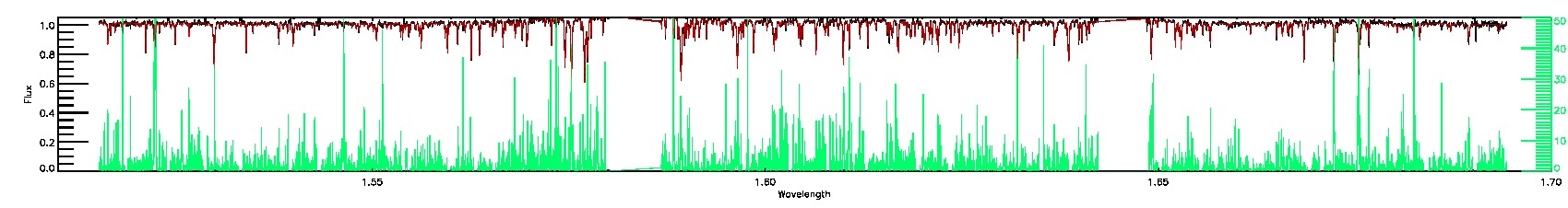
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
120-61-O
| 5176. | +/- | 89. |
| -10000. | +/- | -NaN |
| 4.26 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.03 | +/- | 0. |
| 1.03 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.34 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.34 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.07 | +/- | 0. |
| 0.70 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.31 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.98 |
| -9999.00 |

120-61-O
| 5772. | +/- | 27. |
| 5719. | +/- | 143. |
| 4.07 | +/- | 0. |
| 4.04 | +/- | 0. |
| 1.45 | +/- | 0. |
| 1.45 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -0.14 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 4.50 | +/- | 0. |
| 0.65 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -2.48 |
| -9999.00 |

STAR_WARN,SN_WARN
120-61-O
| 5248. | +/- | 23. |
| 5241. | +/- | 130. |
| 4.27 | +/- | 0. |
| 4.19 | +/- | 0. |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.22 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 2.60 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.99 |
| -9999.00 |

BRIGHT_NEIGHBOR
STAR_WARN,COLORTE_WARN,SN_WARN
120-61-O
| 3464. | +/- | 7. |
| 3567. | +/- | 82. |
| 4.29 | +/- | 0. |
| 4.77 | +/- | 0. |
| 0.94 | +/- | 0. |
| 0.94 | +/- | -NaN |
| -0.33 | +/- | 0. |
| -0.33 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 5.69 | +/- | 0. |
| 0.75 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.90 |
| -9999.00 |
SUSPECT_BROAD_LINES
120-61-O
| 3994. | +/- | 5. |
| 4087. | +/- | 88. |
| 4.39 | +/- | 0. |
| 4.68 | +/- | 0. |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.12 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 14.57 | +/- | 0. |
| 1.16 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.06 |
| -9999.00 |

120-61-O
| 5679. | +/- | 28. |
| 5636. | +/- | 144. |
| 4.45 | +/- | 0. |
| 4.43 | +/- | 0. |
| 0.62 | +/- | 0. |
| 0.62 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.13 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.62 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.09 |
| -9999.00 |

BRIGHT_NEIGHBOR
120-61-O
| 4755. | +/- | 9. |
| 4872. | +/- | 106. |
| 2.42 | +/- | 0. |
| 2.36 | +/- | 0. |
| 1.02 | +/- | 0. |
| 1.02 | +/- | -NaN |
| -1.35 | +/- | 0. |
| -1.35 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.17 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.30 | +/- | 0. |
| 6.52 | +/- | 0. |
| 0.81 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -2.06 |
| -9999.00 |
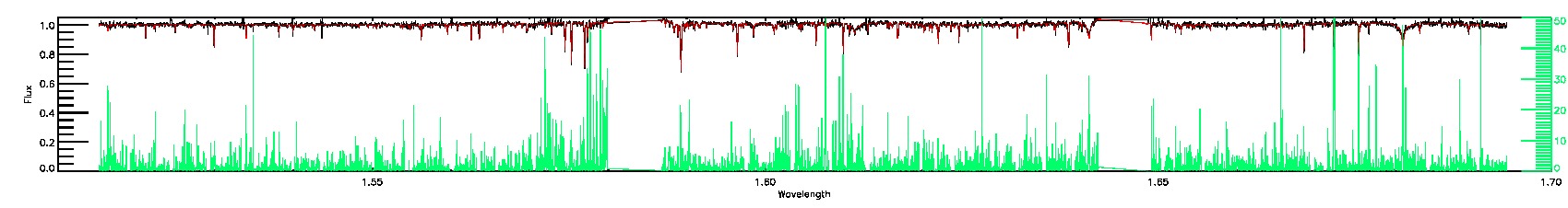
120-61-O
| 4794. | +/- | 6. |
| 4857. | +/- | 94. |
| 4.49 | +/- | 0. |
| 4.67 | +/- | 0. |
| 1.15 | +/- | 0. |
| 1.15 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -0.20 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 1.74 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.23 |
| -9999.00 |

120-61-O
| 6017. | +/- | 35. |
| 5934. | +/- | 161. |
| 4.56 | +/- | 0. |
| 4.46 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.00 | +/- | 0. |
| 6.92 | +/- | 0. |
| 0.84 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.08 |
| -9999.00 |
120-61-O
| 3747. | +/- | 1. |
| 3881. | +/- | 71. |
| 0.59 | +/- | 0. |
| 0.58 | +/- | 0. |
| 2.50 | +/- | 0. |
| 2.50 | +/- | -NaN |
| -1.06 | +/- | 0. |
| -1.06 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.11 | +/- | 0. |
| 5.51 | +/- | 0. |
| 0.74 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -1.89 |
| -9999.00 |
120-61-O
| 6601. | +/- | 23. |
| 6455. | +/- | 170. |
| 4.28 | +/- | 0. |
| 4.12 | +/- | 0. |
| 1.46 | +/- | 0. |
| 1.46 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -0.21 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.09 | +/- | 0. |
| 9.17 | +/- | 0. |
| 0.96 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -0.26 |
| -9999.00 |
120-61-O
| 5796. | +/- | 28. |
| 5736. | +/- | 151. |
| 4.20 | +/- | 0. |
| 4.11 | +/- | 0. |
| 0.89 | +/- | 0. |
| 0.89 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.18 | +/- | 0. |
| -0.18 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.71 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.01 |
| -9999.00 |
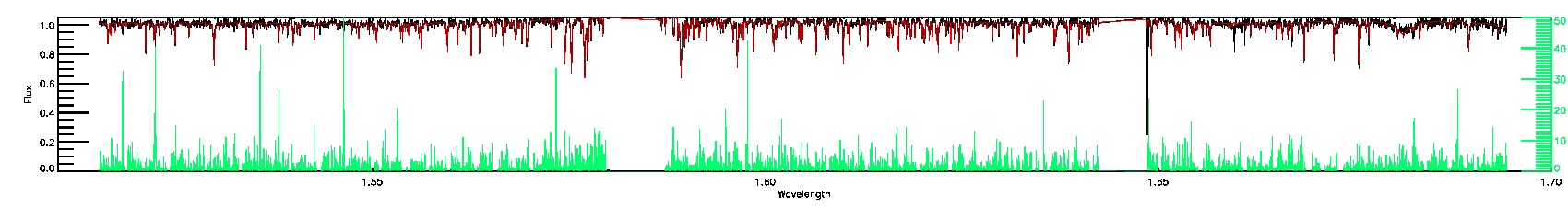
120-61-O
| 5468. | +/- | 9. |
| 5442. | +/- | 104. |
| 4.68 | +/- | 0. |
| 4.63 | +/- | 0. |
| 1.62 | +/- | 0. |
| 1.62 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 5.33 | +/- | 0. |
| 0.73 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.25 |
| -9999.00 |

120-61-O
| 4686. | +/- | 8. |
| 4772. | +/- | 102. |
| 3.90 | +/- | 0. |
| 4.13 | +/- | 0. |
| 0.40 | +/- | 0. |
| 0.40 | +/- | -NaN |
| -0.46 | +/- | 0. |
| -0.45 | +/- | 0. |
| -0.30 | +/- | 0. |
| -0.30 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.14 | +/- | 0. |
| 6.09 | +/- | 0. |
| 0.78 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.22 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.53 |
| -9999.00 |
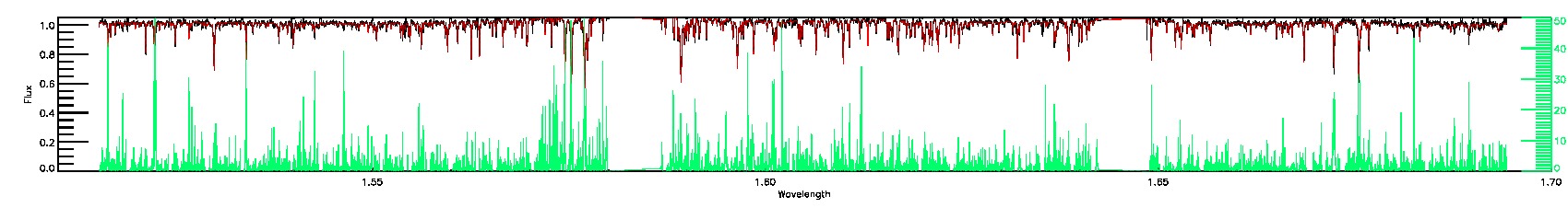
STAR_WARN,SN_WARN
120-61-O
| 5511. | +/- | 30. |
| 5479. | +/- | 141. |
| 4.07 | +/- | 0. |
| 4.00 | +/- | 0. |
| 0.98 | +/- | 0. |
| 0.98 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.84 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.42 |
| -9999.00 |

BRIGHT_NEIGHBOR
120-61-O
| 5788. | +/- | 31. |
| 5737. | +/- | 152. |
| 4.49 | +/- | 0. |
| 4.48 | +/- | 0. |
| 0.85 | +/- | 0. |
| 0.85 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.47 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.48 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
120-61-O
| 3361. | +/- | 6. |
| 3467. | +/- | 77. |
| 4.56 | +/- | 0. |
| 5.10 | +/- | 0. |
| 2.00 | +/- | 0. |
| 2.00 | +/- | -NaN |
| -0.42 | +/- | 0. |
| -0.41 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.21 | +/- | 0. |
| -0.21 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 7.14 | +/- | 0. |
| 0.85 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.42 |
| -9999.00 |
120-61-O
| 5957. | +/- | 30. |
| 5892. | +/- | 150. |
| 4.42 | +/- | 0. |
| 4.44 | +/- | 0. |
| 0.97 | +/- | 0. |
| 0.97 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -0.35 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.94 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.16 |
| -9999.00 |
120-61-O
| 5451. | +/- | 11. |
| 5418. | +/- | 120. |
| 4.33 | +/- | 0. |
| 4.20 | +/- | 0. |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 1.80 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.04 |
| -9999.00 |

120-61-O
| 4938. | +/- | 6. |
| 4974. | +/- | 89. |
| 3.71 | +/- | 0. |
| 3.65 | +/- | 0. |
| 0.99 | +/- | 0. |
| 0.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.91 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.02 |
| -9999.00 |

120-61-O
| 5700. | +/- | 12. |
| 5649. | +/- | 112. |
| 4.49 | +/- | 0. |
| 4.40 | +/- | 0. |
| 0.85 | +/- | 0. |
| 0.85 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 10.73 | +/- | 0. |
| 1.03 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |

120-61-O
| 5307. | +/- | 8. |
| 5303. | +/- | 105. |
| 4.48 | +/- | 0. |
| 4.49 | +/- | 0. |
| 0.43 | +/- | 0. |
| 0.43 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 1.56 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.13 |
| -9999.00 |

120-61-O
| 5384. | +/- | 11. |
| 5374. | +/- | 111. |
| 4.62 | +/- | 0. |
| 4.64 | +/- | 0. |
| 1.12 | +/- | 0. |
| 1.12 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.00 | +/- | 0. |
| 1.89 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.30 |
| -9999.00 |

120-61-O
| 5366. | +/- | 11. |
| 5347. | +/- | 114. |
| 4.05 | +/- | 0. |
| 3.97 | +/- | 0. |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.16 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.74 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.33 |
| -9999.00 |

120-61-O
| 5962. | +/- | 28. |
| 5893. | +/- | 153. |
| 4.41 | +/- | 0. |
| 4.40 | +/- | 0. |
| 0.79 | +/- | 0. |
| 0.79 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -0.29 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.07 | +/- | 0. |
| 2.45 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.87 |
| -9999.00 |

120-61-O
| 6010. | +/- | 27. |
| 5923. | +/- | 153. |
| 4.53 | +/- | 0. |
| 4.38 | +/- | 0. |
| 1.02 | +/- | 0. |
| 1.02 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.04 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.22 |
| -9999.00 |
120-61-O
| 5739. | +/- | 11. |
| 5677. | +/- | 112. |
| 4.24 | +/- | 0. |
| 4.09 | +/- | 0. |
| 0.94 | +/- | 0. |
| 0.94 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.16 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 6.36 | +/- | 0. |
| 0.80 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.30 |
| -9999.00 |

120-61-O
| 4915. | +/- | 11. |
| 4951. | +/- | 111. |
| 4.58 | +/- | 0. |
| 4.60 | +/- | 0. |
| 1.55 | +/- | 0. |
| 1.55 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 1.80 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.16 |
| -9999.00 |

120-61-O
| 5408. | +/- | 8. |
| 5382. | +/- | 101. |
| 4.39 | +/- | 0. |
| 4.28 | +/- | 0. |
| 0.53 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.01 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.19 |
| -9999.00 |

STAR_WARN,SN_WARN
120-61-O
| 4630. | +/- | 11. |
| 4704. | +/- | 109. |
| 4.50 | +/- | 0. |
| 4.62 | +/- | 0. |
| 0.43 | +/- | 0. |
| 0.43 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 4.72 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.17 |
| -9999.00 |

120-61-O
| 4327. | +/- | 7. |
| 4418. | +/- | 99. |
| 4.41 | +/- | 0. |
| 4.61 | +/- | 0. |
| 0.74 | +/- | 0. |
| 0.74 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.56 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.05 |
| -9999.00 |

STAR_WARN,SN_WARN
120-61-O
| 4431. | +/- | 9. |
| 4531. | +/- | 108. |
| 4.09 | +/- | 0. |
| 4.36 | +/- | 0. |
| 0.71 | +/- | 0. |
| 0.71 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -0.26 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 6.29 | +/- | 0. |
| 0.80 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.11 |
| -9999.00 |

STAR_BAD
STAR_WARN,SN_WARN
120-61-O
| 4098. | +/- | 10. |
| -10000. | +/- | -NaN |
| 4.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.40 | +/- | 0. |
| 0.40 | +/- | -NaN |
| -0.42 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 13.21 | +/- | 0. |
| 1.12 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.29 |
| -9999.00 |

120-61-O
| 5760. | +/- | 16. |
| 5692. | +/- | 130. |
| 4.48 | +/- | 0. |
| 4.30 | +/- | 0. |
| 0.72 | +/- | 0. |
| 0.72 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.23 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.86 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.52 |
| -9999.00 |

120-61-O
| 4714. | +/- | 8. |
| 4788. | +/- | 106. |
| 4.52 | +/- | 0. |
| 4.73 | +/- | 0. |
| 0.67 | +/- | 0. |
| 0.67 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -0.25 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.66 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.43 |
| -9999.00 |

120-61-O
| 5816. | +/- | 24. |
| 5759. | +/- | 147. |
| 4.34 | +/- | 0. |
| 4.31 | +/- | 0. |
| 1.32 | +/- | 0. |
| 1.32 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.16 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.08 | +/- | 0. |
| 1.79 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| 1.00 |
| -9999.00 |
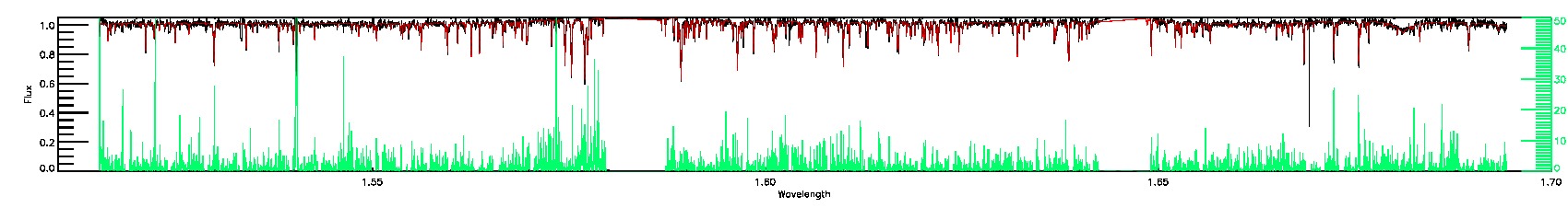
STAR_WARN,SN_WARN
120-61-O
| 6032. | +/- | 54. |
| 5950. | +/- | 172. |
| 4.24 | +/- | 0. |
| 4.17 | +/- | 0. |
| 1.46 | +/- | 0. |
| 1.46 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -0.17 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.45 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 4.27 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| -2.35 |
| -9999.00 |
120-61-O
| 4246. | +/- | 3. |
| 4360. | +/- | 89. |
| 4.27 | +/- | 0. |
| 4.71 | +/- | 0. |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| -0.59 | +/- | 0. |
| -0.59 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.13 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.14 | +/- | 0. |
| 5.13 | +/- | 0. |
| 0.71 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.63 |
| -9999.00 |

BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_WARN,SN_WARN
120-61-O
| 3850. | +/- | 9. |
| 3933. | +/- | 87. |
| 4.54 | +/- | 0. |
| 4.75 | +/- | 0. |
| 0.92 | +/- | 0. |
| 0.92 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.14 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.07 | +/- | 0. |
| 37.96 | +/- | 0. |
| 1.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.35 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_WARN,COLORTE_WARN,SN_WARN
120-61-O
| 3538. | +/- | 11. |
| 3640. | +/- | 87. |
| 4.20 | +/- | 0. |
| 4.66 | +/- | 0. |
| 0.77 | +/- | 0. |
| 0.77 | +/- | -NaN |
| -0.32 | +/- | 0. |
| -0.32 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -0.18 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 13.42 | +/- | 0. |
| 1.13 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.67 |
| -9999.00 |
120-61-O
| 5458. | +/- | 11. |
| 5433. | +/- | 111. |
| 4.42 | +/- | 0. |
| 4.37 | +/- | 0. |
| 0.71 | +/- | 0. |
| 0.71 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 4.44 | +/- | 0. |
| 0.65 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.25 |
| -9999.00 |
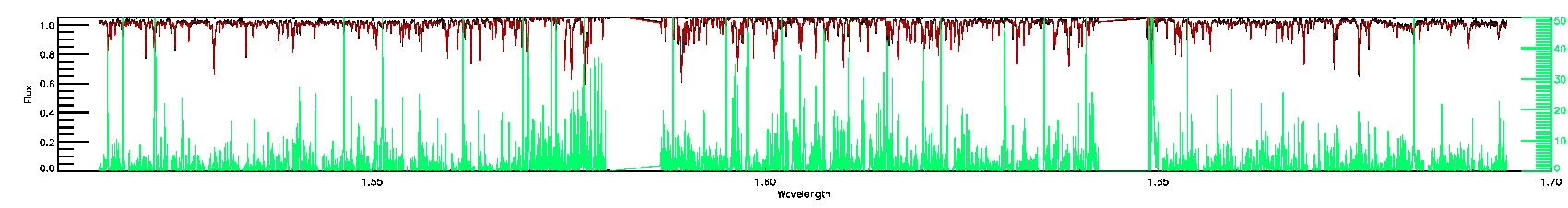
120-61-O
| 5524. | +/- | 35. |
| 5506. | +/- | 145. |
| 4.39 | +/- | 0. |
| 4.46 | +/- | 0. |
| 1.10 | +/- | 0. |
| 1.10 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -0.29 | +/- | 0. |
| -0.17 | +/- | 0. |
| -0.17 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 2.53 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.02 |
| -9999.00 |

120-61-O
| 6295. | +/- | 25. |
| 6219. | +/- | 151. |
| 4.41 | +/- | 0. |
| 4.64 | +/- | 0. |
| 0.94 | +/- | 0. |
| 0.94 | +/- | -NaN |
| -1.03 | +/- | 0. |
| -1.03 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.22 | +/- | 0. |
| 2.55 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -1.98 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -1.21 |
| -9999.00 |
120-61-O
| 5425. | +/- | 38. |
| 5438. | +/- | 150. |
| 3.92 | +/- | 0. |
| 4.15 | +/- | 0. |
| 1.08 | +/- | 0. |
| 1.08 | +/- | -NaN |
| -0.75 | +/- | 0. |
| -0.75 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.19 | +/- | 0. |
| 4.59 | +/- | 0. |
| 0.66 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.33 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| -2.33 |
| -9999.00 |
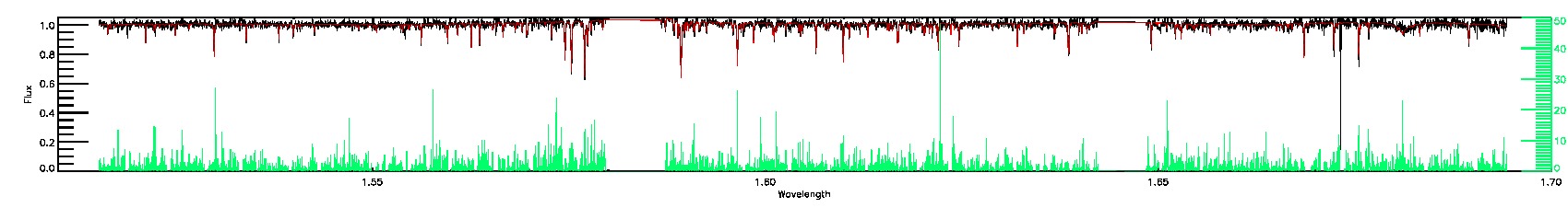
STAR_WARN,SN_WARN
120-61-O
| 4897. | +/- | 15. |
| 4923. | +/- | 115. |
| 4.43 | +/- | 0. |
| 4.35 | +/- | 0. |
| 0.80 | +/- | 0. |
| 0.80 | +/- | -NaN |
| 0.38 | +/- | 0. |
| 0.38 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.14 | +/- | 0. |
| 1.57 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.20 |
| -9999.00 |

STAR_WARN,SN_WARN
120-61-O
| 4906. | +/- | 24. |
| 4950. | +/- | 124. |
| 4.37 | +/- | 0. |
| 4.46 | +/- | 0. |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 3.64 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.62 |
| -9999.00 |
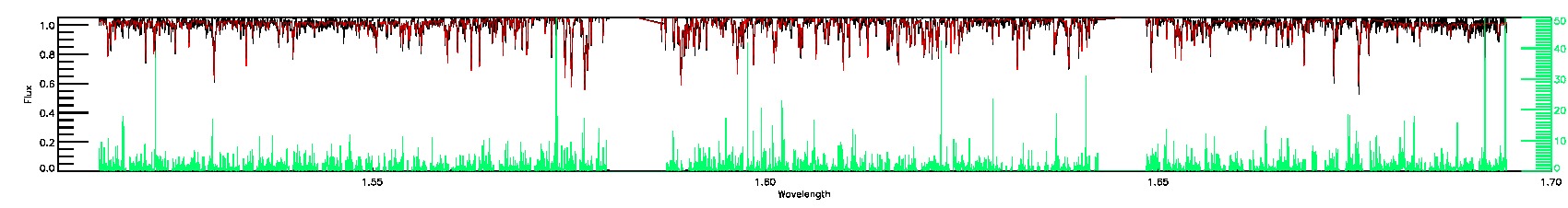
120-61-O
| 4757. | +/- | 5. |
| 4806. | +/- | 91. |
| 3.66 | +/- | 0. |
| 3.55 | +/- | 0. |
| 0.54 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.60 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| 0.31 |
| -9999.00 |

STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
120-61-O
| 4345. | +/- | 36. |
| -10000. | +/- | -NaN |
| 4.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.56 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.24 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.98 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.08 |
| -9999.00 |

120-61-O
| 5009. | +/- | 11. |
| 5026. | +/- | 113. |
| 4.64 | +/- | 0. |
| 4.56 | +/- | 0. |
| 0.87 | +/- | 0. |
| 0.87 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.31 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.07 | +/- | 0. |
| 2.25 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.25 |
| -9999.00 |
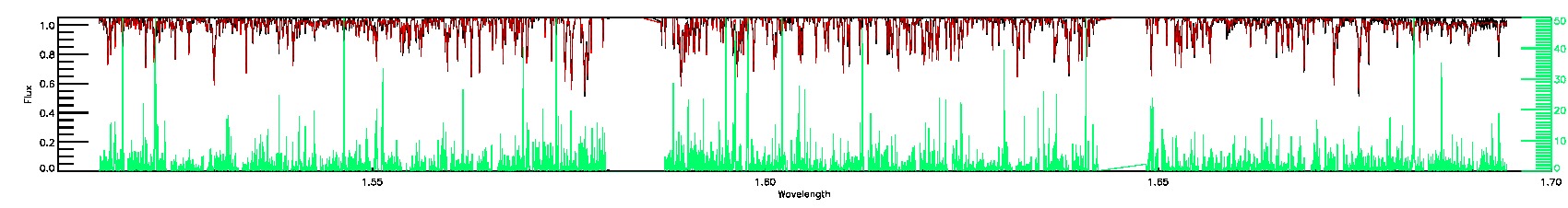
120-61-O
| 5456. | +/- | 18. |
| 5424. | +/- | 133. |
| 4.28 | +/- | 0. |
| 4.16 | +/- | 0. |
| 0.40 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 1.69 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| 0.52 |
| -9999.00 |

120-61-O
| 4882. | +/- | 5. |
| 4925. | +/- | 88. |
| 3.33 | +/- | 0. |
| 3.15 | +/- | 0. |
| 1.18 | +/- | 0. |
| 1.18 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.92 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.78 |
| -9999.00 |

120-61-O
| 6263. | +/- | 15. |
| 6149. | +/- | 133. |
| 4.64 | +/- | 0. |
| 4.48 | +/- | 0. |
| 1.11 | +/- | 0. |
| 1.11 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.75 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.28 |
| -9999.00 |
120-61-O
| 5782. | +/- | 9. |
| 5716. | +/- | 115. |
| 4.39 | +/- | 0. |
| 4.24 | +/- | 0. |
| 1.00 | +/- | 0. |
| 1.00 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.14 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 3.08 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.32 |
| -9999.00 |

120-61-O
| 5860. | +/- | 12. |
| 5797. | +/- | 120. |
| 4.45 | +/- | 0. |
| 4.41 | +/- | 0. |
| 0.82 | +/- | 0. |
| 0.82 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -0.26 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.25 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.17 |
| -9999.00 |
120-61-O
| 6279. | +/- | 16. |
| 6161. | +/- | 134. |
| 4.11 | +/- | 0. |
| 3.92 | +/- | 0. |
| 1.79 | +/- | 0. |
| 1.79 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.02 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 9.45 | +/- | 0. |
| 0.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.29 |
| -9999.00 |
120-61-O
| 5960. | +/- | 26. |
| 5883. | +/- | 149. |
| 4.58 | +/- | 0. |
| 4.49 | +/- | 0. |
| 0.85 | +/- | 0. |
| 0.85 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.13 | +/- | 0. |
| -0.13 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -0.26 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.32 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.36 |
| -9999.00 |
120-61-O
| 5842. | +/- | 39. |
| 5790. | +/- | 159. |
| 4.43 | +/- | 0. |
| 4.47 | +/- | 0. |
| 0.48 | +/- | 0. |
| 0.48 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -0.36 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.07 | +/- | 0. |
| 1.61 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -2.19 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_WARN,SN_WARN
120-61-O
| 5966. | +/- | 140. |
| 5915. | +/- | 186. |
| 4.05 | +/- | 0. |
| 4.22 | +/- | 0. |
| 0.45 | +/- | 0. |
| 0.45 | +/- | -NaN |
| -0.74 | +/- | 0. |
| -0.73 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 20.44 | +/- | 0. |
| 1.31 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.16 |
| -9999.00 |
STAR_WARN,SN_WARN
120-61-O
| 5306. | +/- | 35. |
| 5298. | +/- | 140. |
| 4.58 | +/- | 0. |
| 4.56 | +/- | 0. |
| 1.22 | +/- | 0. |
| 1.22 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.06 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 1.57 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -9999.00 |
| -9999.00 |

STAR_WARN,SN_WARN
120-61-O
| 4682. | +/- | 17. |
| 4743. | +/- | 114. |
| 4.59 | +/- | 0. |
| 4.65 | +/- | 0. |
| 0.56 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.05 | +/- | 0. |
| 1.54 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.31 |
| -9999.00 |

120-61-O
| 5715. | +/- | 39. |
| 5680. | +/- | 151. |
| 4.15 | +/- | 0. |
| 4.24 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.43 | +/- | -NaN |
| -0.42 | +/- | 0. |
| -0.41 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -0.44 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.06 | +/- | 0. |
| 3.77 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| -0.57 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
120-61-O
| 3320. | +/- | 5. |
| 3417. | +/- | 75. |
| 4.54 | +/- | 0. |
| 4.99 | +/- | 0. |
| 1.85 | +/- | 0. |
| 1.85 | +/- | -NaN |
| -0.21 | +/- | 0. |
| -0.21 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.06 | +/- | 0. |
| 6.63 | +/- | 0. |
| 0.82 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.23 |
| -9999.00 |
SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
120-61-O
| 5652. | +/- | 35. |
| 5603. | +/- | 136. |
| 4.78 | +/- | 0. |
| 4.68 | +/- | 0. |
| 0.63 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -0.35 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.09 | +/- | 0. |
| 71.70 | +/- | 0. |
| 1.86 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.16 |
| -9999.00 |
STAR_WARN,SN_WARN
120-61-O
| 4936. | +/- | 25. |
| 4977. | +/- | 126. |
| 4.51 | +/- | 0. |
| 4.60 | +/- | 0. |
| 0.68 | +/- | 0. |
| 0.68 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.02 | +/- | 0. |
| 1.83 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -2.44 |
| -9999.00 |

SUSPECT_BROAD_LINES
120-61-O
| 5969. | +/- | 15. |
| 5884. | +/- | 122. |
| 4.78 | +/- | 0. |
| 4.62 | +/- | 0. |
| 0.66 | +/- | 0. |
| 0.66 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.07 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.45 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.00 | +/- | 0. |
| 15.37 | +/- | 0. |
| 1.19 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.52 |
| -9999.00 |
120-61-O
| 4932. | +/- | 23. |
| 5027. | +/- | 133. |
| 2.69 | +/- | 0. |
| 2.65 | +/- | 0. |
| 0.47 | +/- | 0. |
| 0.47 | +/- | -NaN |
| -1.32 | +/- | 0. |
| -1.32 | +/- | 0. |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -0.40 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.29 | +/- | 0. |
| 6.41 | +/- | 0. |
| 0.81 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.31 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -1.93 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.66 |
| -9999.00 |

120-61-O
| 5046. | +/- | 12. |
| 5066. | +/- | 112. |
| 4.47 | +/- | 0. |
| 4.46 | +/- | 0. |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.05 | +/- | 0. |
| 7.78 | +/- | 0. |
| 0.89 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.19 |
| -9999.00 |
120-61-O
| 4892. | +/- | 12. |
| 4952. | +/- | 113. |
| 4.52 | +/- | 0. |
| 4.75 | +/- | 0. |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| -0.39 | +/- | 0. |
| -0.38 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -0.17 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.07 | +/- | 0. |
| 1.72 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.33 |
| -9999.00 |

STAR_WARN,SN_WARN
120-61-O
| 5486. | +/- | 50. |
| 5474. | +/- | 151. |
| 4.40 | +/- | 0. |
| 4.49 | +/- | 0. |
| 1.39 | +/- | 0. |
| 1.39 | +/- | -NaN |
| -0.32 | +/- | 0. |
| -0.32 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.55 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -0.80 |
| -9999.00 |

STAR_WARN,COLORTE_WARN,SN_WARN
120-61-O
| 3735. | +/- | 11. |
| 3846. | +/- | 94. |
| 4.40 | +/- | 0. |
| 4.90 | +/- | 0. |
| 0.70 | +/- | 0. |
| 0.70 | +/- | -NaN |
| -0.53 | +/- | 0. |
| -0.53 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.07 | +/- | 0. |
| 4.29 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -0.08 |
| -9999.00 |
120-61-O
| 5673. | +/- | 19. |
| 5625. | +/- | 134. |
| 4.57 | +/- | 0. |
| 4.50 | +/- | 0. |
| 0.58 | +/- | 0. |
| 0.58 | +/- | -NaN |
| -0.00 | +/- | 0. |
| 0.00 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 5.08 | +/- | 0. |
| 0.71 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -0.53 |
| -9999.00 |

120-61-O
| 5702. | +/- | 24. |
| 5667. | +/- | 135. |
| 4.11 | +/- | 0. |
| 4.18 | +/- | 0. |
| 0.95 | +/- | 0. |
| 0.95 | +/- | -NaN |
| -0.38 | +/- | 0. |
| -0.38 | +/- | 0. |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.69 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.27 |
| -9999.00 |

STAR_BAD
STAR_WARN,COLORTE_WARN
120-61-O
| 3547. | +/- | 8. |
| -10000. | +/- | -NaN |
| 3.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.15 | +/- | 0. |
| 1.15 | +/- | -NaN |
| -1.52 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.45 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.20 | +/- | 0. |
| 0.86 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.15 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| -1.45 |
| -9999.00 |
120-61-O
| 4496. | +/- | 4. |
| 4586. | +/- | 92. |
| 4.42 | +/- | 0. |
| 4.58 | +/- | 0. |
| 1.17 | +/- | 0. |
| 1.17 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 12.74 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.15 |
| -9999.00 |

120-61-O
| 6315. | +/- | 16. |
| 6192. | +/- | 134. |
| 4.65 | +/- | 0. |
| 4.44 | +/- | 0. |
| 1.15 | +/- | 0. |
| 1.15 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.08 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 5.49 | +/- | 0. |
| 0.74 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.53 |
| -9999.00 |
120-61-O
| 4927. | +/- | 9. |
| 4979. | +/- | 102. |
| 3.59 | +/- | 0. |
| 3.53 | +/- | 0. |
| 1.18 | +/- | 0. |
| 1.18 | +/- | -NaN |
| -0.30 | +/- | 0. |
| -0.30 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.13 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.53 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.51 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
120-61-O
| 6414. | +/- | 24. |
| 6287. | +/- | 156. |
| 4.67 | +/- | 0. |
| 4.52 | +/- | 0. |
| 0.96 | +/- | 0. |
| 0.96 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -0.16 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 4.33 | +/- | 0. |
| 0.64 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.28 |
| -9999.00 |
SUSPECT_BROAD_LINES
120-61-O
| 6280. | +/- | 15. |
| 6157. | +/- | 132. |
| 4.09 | +/- | 0. |
| 3.86 | +/- | 0. |
| 1.86 | +/- | 0. |
| 1.86 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.10 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 26.83 | +/- | 0. |
| 1.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.19 |
| -9999.00 |
120-61-O
| 5487. | +/- | 29. |
| 5474. | +/- | 138. |
| 4.24 | +/- | 0. |
| 4.33 | +/- | 0. |
| 0.81 | +/- | 0. |
| 0.81 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -0.30 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 6.71 | +/- | 0. |
| 0.83 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.36 |
| -9999.00 |
120-61-O
| 5356. | +/- | 15. |
| 5329. | +/- | 126. |
| 4.44 | +/- | 0. |
| 4.27 | +/- | 0. |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| 0.38 | +/- | 0. |
| 0.38 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.74 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.20 |
| -9999.00 |

120-61-O
| 5346. | +/- | 10. |
| 5341. | +/- | 105. |
| 3.91 | +/- | 0. |
| 3.91 | +/- | 0. |
| 0.99 | +/- | 0. |
| 0.99 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.13 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 4.57 | +/- | 0. |
| 0.66 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.22 |
| -9999.00 |

120-61-O
| 5949. | +/- | 17. |
| 5871. | +/- | 138. |
| 4.34 | +/- | 0. |
| 4.23 | +/- | 0. |
| 1.10 | +/- | 0. |
| 1.10 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 9.35 | +/- | 0. |
| 0.97 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.15 |
| -9999.00 |

120-61-O
| 5148. | +/- | 17. |
| 5166. | +/- | 126. |
| 4.56 | +/- | 0. |
| 4.64 | +/- | 0. |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 1.57 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| 0.14 |
| -9999.00 |

120-61-O
| 4927. | +/- | 16. |
| 4966. | +/- | 118. |
| 4.55 | +/- | 0. |
| 4.62 | +/- | 0. |
| 1.03 | +/- | 0. |
| 1.03 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.01 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 1.96 | +/- | 0. |
| 0.29 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.34 |
| -9999.00 |

STAR_WARN,SN_WARN
120-61-O
| 4892. | +/- | 18. |
| 4950. | +/- | 123. |
| 3.67 | +/- | 0. |
| 3.68 | +/- | 0. |
| 0.51 | +/- | 0. |
| 0.51 | +/- | -NaN |
| -0.34 | +/- | 0. |
| -0.34 | +/- | 0. |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| -0.21 | +/- | 0. |
| -0.21 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.21 | +/- | 0. |
| 3.62 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.24 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.55 |
| -9999.00 |

SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES,BAD_RV_COMBINATION
120-61-O
| 5049. | +/- | 21. |
| 5067. | +/- | 119. |
| 4.24 | +/- | 0. |
| 4.22 | +/- | 0. |
| 4.20 | +/- | 0. |
| 4.20 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.15 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 58.13 | +/- | 0. |
| 1.76 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.23 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.07 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_WARN,SN_WARN
120-61-O
| 4668. | +/- | 25. |
| 4795. | +/- | 133. |
| 2.12 | +/- | 0. |
| 2.06 | +/- | 0. |
| 1.87 | +/- | 0. |
| 1.87 | +/- | -NaN |
| -1.37 | +/- | 0. |
| -1.37 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.25 | +/- | 0. |
| 6.59 | +/- | 0. |
| 0.82 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.91 |
| -9999.00 |

120-61-O
| 5840. | +/- | 26. |
| 5772. | +/- | 152. |
| 4.33 | +/- | 0. |
| 4.21 | +/- | 0. |
| 1.16 | +/- | 0. |
| 1.16 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 1.85 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.04 |
| -9999.00 |

STAR_WARN,SN_WARN
120-61-O
| 5653. | +/- | 55. |
| 5615. | +/- | 157. |
| 4.31 | +/- | 0. |
| 4.32 | +/- | 0. |
| 0.63 | +/- | 0. |
| 0.63 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -0.18 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -0.18 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 6.59 | +/- | 0. |
| 0.82 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |

120-61-O
| 5683. | +/- | 31. |
| 5638. | +/- | 147. |
| 4.51 | +/- | 0. |
| 4.46 | +/- | 0. |
| 1.04 | +/- | 0. |
| 1.04 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.52 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.57 |
| -9999.00 |
STAR_WARN,SN_WARN
120-61-O
| 5863. | +/- | 33. |
| 5793. | +/- | 159. |
| 4.58 | +/- | 0. |
| 4.47 | +/- | 0. |
| 0.83 | +/- | 0. |
| 0.83 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 1.94 | +/- | 0. |
| 0.29 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -0.56 |
| -9999.00 |

120-61-O
| 4664. | +/- | 4. |
| 4737. | +/- | 83. |
| 2.84 | +/- | 0. |
| 2.48 | +/- | 0. |
| 0.86 | +/- | 0. |
| 0.86 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | 0. |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.09 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.02 |
| -9999.00 |

STAR_WARN,SN_WARN
120-61-O
| 4174. | +/- | 12. |
| 4267. | +/- | 102. |
| 4.34 | +/- | 0. |
| 4.59 | +/- | 0. |
| 0.49 | +/- | 0. |
| 0.49 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -0.12 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 5.76 | +/- | 0. |
| 0.76 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.10 |
| -9999.00 |
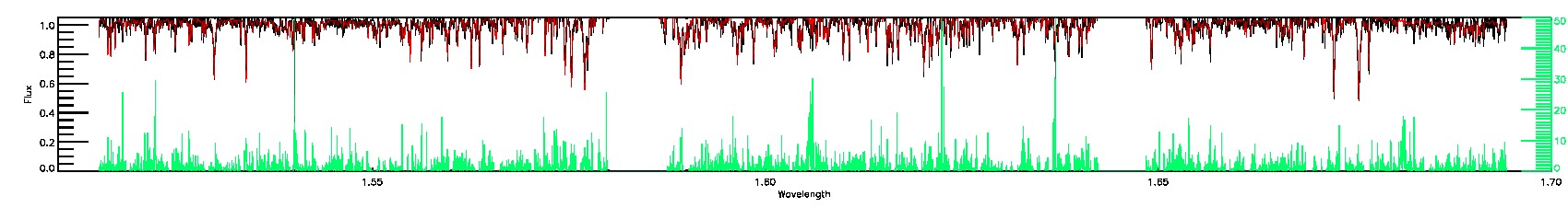
120-61-O
| 4877. | +/- | 6. |
| 4920. | +/- | 91. |
| 4.48 | +/- | 0. |
| 4.54 | +/- | 0. |
| 1.42 | +/- | 0. |
| 1.42 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 1.59 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.05 |
| -9999.00 |

BRIGHT_NEIGHBOR
120-61-O
| 5758. | +/- | 25. |
| 5701. | +/- | 150. |
| 4.43 | +/- | 0. |
| 4.35 | +/- | 0. |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.02 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.08 | +/- | 0. |
| 1.61 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 0.12 |
| -9999.00 |
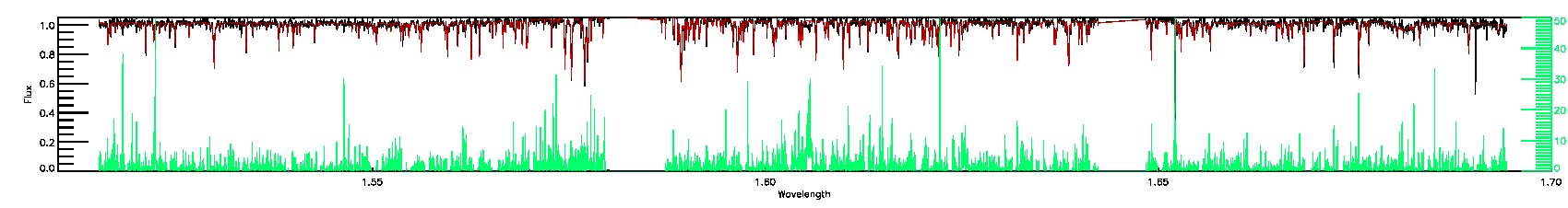
STAR_WARN,SN_WARN
120-61-O
| 4393. | +/- | 9. |
| 4479. | +/- | 106. |
| 4.41 | +/- | 0. |
| 4.55 | +/- | 0. |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.07 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.43 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -0.26 |
| -9999.00 |

