STAR_WARN,COLORTE_WARN
280-33-O
| 5240. | +/- | 7. |
| 5238. | +/- | 97. |
| 4.66 | +/- | 0. |
| 4.62 | +/- | 0. |
| 1.64 | +/- | 0. |
| 1.64 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 4.86 | +/- | 0. |
| 0.69 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.14 |
| -9999.00 |

PERSIST_JUMP_NEG
STAR_BAD,CHI2_BAD
STAR_WARN,CHI2_WARN
280-33-O
| 3160. | +/- | 1. |
| -10000. | +/- | -NaN |
| 4.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.98 | +/- | 0. |
| 3.98 | +/- | -NaN |
| 0.52 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.75 | +/- | 0. |
| 0.83 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.50 |
| -9999.00 |
STAR_BAD,CHI2_BAD
STAR_WARN,CHI2_WARN
280-33-O
| 3081. | +/- | 1. |
| -10000. | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.44 | +/- | 0. |
| 3.44 | +/- | -NaN |
| -0.99 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.29 | +/- | 0. |
| 0.72 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.67 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5211. | +/- | 39. |
| -10000. | +/- | -NaN |
| 1.31 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.07 | +/- | 0. |
| 4.07 | +/- | -NaN |
| -0.60 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.26 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.20 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.48 |
| -9999.00 |

280-33-O
| 5874. | +/- | 34. |
| 5806. | +/- | 151. |
| 4.22 | +/- | 0. |
| 4.14 | +/- | 0. |
| 1.18 | +/- | 0. |
| 1.18 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.11 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.20 |
| -9999.00 |

STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN
280-33-O
| 5228. | +/- | 9. |
| -10000. | +/- | -NaN |
| 1.66 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.79 | +/- | 0. |
| 0.79 | +/- | -NaN |
| -2.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| -0.71 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.70 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.48 |
| -9999.00 |

280-33-O
| 3835. | +/- | 1. |
| 3939. | +/- | 67. |
| 0.38 | +/- | 0. |
| 0.28 | +/- | 0. |
| 2.79 | +/- | 0. |
| 2.79 | +/- | -NaN |
| -0.37 | +/- | 0. |
| -0.37 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.22 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.05 | +/- | 0. |
| 3.67 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.23 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.24 |
| -9999.00 |
280-33-O
| 6090. | +/- | 14. |
| 5990. | +/- | 125. |
| 4.20 | +/- | 0. |
| 4.02 | +/- | 0. |
| 1.33 | +/- | 0. |
| 1.33 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.44 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 11.61 | +/- | 0. |
| 1.06 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.05 |
| -9999.00 |
280-33-O
| 5707. | +/- | 11. |
| 5654. | +/- | 112. |
| 3.88 | +/- | 0. |
| 3.77 | +/- | 0. |
| 1.48 | +/- | 0. |
| 1.48 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.03 | +/- | 0. |
| 5.24 | +/- | 0. |
| 0.72 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.04 |
| -9999.00 |

280-33-O
| 5353. | +/- | 8. |
| 5342. | +/- | 101. |
| 3.96 | +/- | 0. |
| 3.93 | +/- | 0. |
| 1.26 | +/- | 0. |
| 1.26 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.94 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.07 |
| -9999.00 |
SUSPECT_RV_COMBINATION
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN
280-33-O
| 6962. | +/- | 4. |
| -10000. | +/- | -NaN |
| 2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.72 | +/- | 0. |
| 0.72 | +/- | -NaN |
| -0.92 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.21 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 17.78 | +/- | 0. |
| 1.25 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.50 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_WARN,COLORTE_WARN
280-33-O
| 4455. | +/- | 3. |
| 4547. | +/- | 85. |
| 2.68 | +/- | 0. |
| 2.47 | +/- | 0. |
| 1.11 | +/- | 0. |
| 1.11 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.08 | +/- | 0. |
| 3.07 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.29 |
| -9999.00 |
280-33-O
| 5742. | +/- | 14. |
| 5703. | +/- | 119. |
| 4.32 | +/- | 0. |
| 4.40 | +/- | 0. |
| 1.09 | +/- | 0. |
| 1.09 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -0.40 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.14 | +/- | 0. |
| 1.68 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.36 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
280-33-O
| 5671. | +/- | 53. |
| -10000. | +/- | -NaN |
| 1.79 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.79 | +/- | 0. |
| 4.79 | +/- | -NaN |
| -0.74 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.22 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.56 | +/- | 0. |
| 0.66 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.37 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES
STAR_BAD,ROTATION_BAD
STAR_WARN,ROTATION_WARN
280-33-O
| 5708. | +/- | 5. |
| -10000. | +/- | -NaN |
| 0.93 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.84 | +/- | 0. |
| 0.84 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| -0.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.73 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.20 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.48 |
| -9999.00 |

STAR_BAD,CHI2_BAD
STAR_WARN,CHI2_WARN
280-33-O
| 3155. | +/- | 1. |
| -10000. | +/- | -NaN |
| 3.99 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.67 | +/- | 0. |
| 3.67 | +/- | -NaN |
| 0.52 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.51 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.72 | +/- | 0. |
| 0.83 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.51 |
| -9999.00 |
280-33-O
| 5909. | +/- | 9. |
| 5825. | +/- | 117. |
| 4.54 | +/- | 0. |
| 4.33 | +/- | 0. |
| 0.64 | +/- | 0. |
| 0.64 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 5.77 | +/- | 0. |
| 0.76 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.50 |
| -9999.00 |
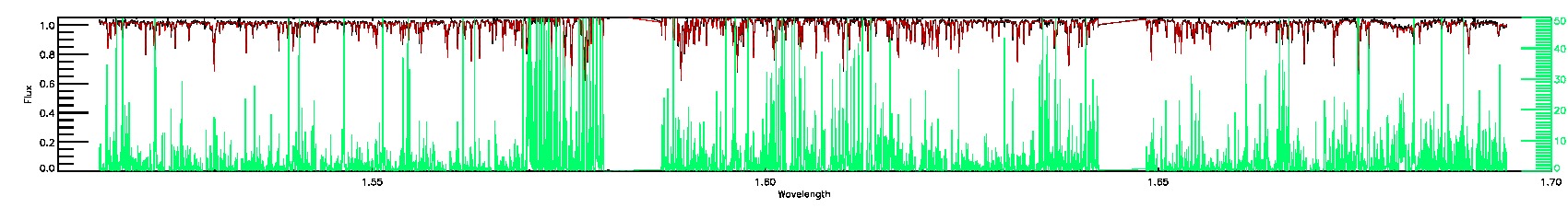
STAR_WARN,COLORTE_WARN
280-33-O
| 3794. | +/- | 1. |
| 3929. | +/- | 72. |
| 0.66 | +/- | 0. |
| 0.65 | +/- | 0. |
| 2.22 | +/- | 0. |
| 2.22 | +/- | -NaN |
| -1.09 | +/- | 0. |
| -1.09 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.20 | +/- | 0. |
| 5.60 | +/- | 0. |
| 0.75 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -1.56 |
| -9999.00 |
280-33-O
| 5738. | +/- | 15. |
| 5700. | +/- | 119. |
| 4.41 | +/- | 0. |
| 4.48 | +/- | 0. |
| 1.10 | +/- | 0. |
| 1.10 | +/- | -NaN |
| -0.41 | +/- | 0. |
| -0.40 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.15 | +/- | 0. |
| 3.46 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.62 |
| -9999.00 |

280-33-O
| 3523. | +/- | 1. |
| 3633. | +/- | 63. |
| -0.10 | +/- | 0. |
| -0.19 | +/- | 0. |
| 2.81 | +/- | 0. |
| 2.81 | +/- | -NaN |
| -0.52 | +/- | 0. |
| -0.52 | +/- | 0. |
| -0.35 | +/- | 0. |
| -0.35 | +/- | -NaN |
| 0.40 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 4.01 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.37 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -1.46 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,ROTATION_BAD
STAR_WARN,ROTATION_WARN
280-33-O
| 3255. | +/- | 1. |
| -10000. | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.70 | +/- | 0. |
| 4.70 | +/- | -NaN |
| -1.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.65 | +/- | 0. |
| 0.82 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -1.37 |
| -9999.00 |
280-33-O
| 4876. | +/- | 5. |
| 4914. | +/- | 86. |
| 4.48 | +/- | 0. |
| 4.48 | +/- | 0. |
| 0.97 | +/- | 0. |
| 0.97 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 3.37 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.20 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5868. | +/- | 21. |
| -10000. | +/- | -NaN |
| 1.62 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.66 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.19 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 1.27 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.71 |
| -9999.00 |
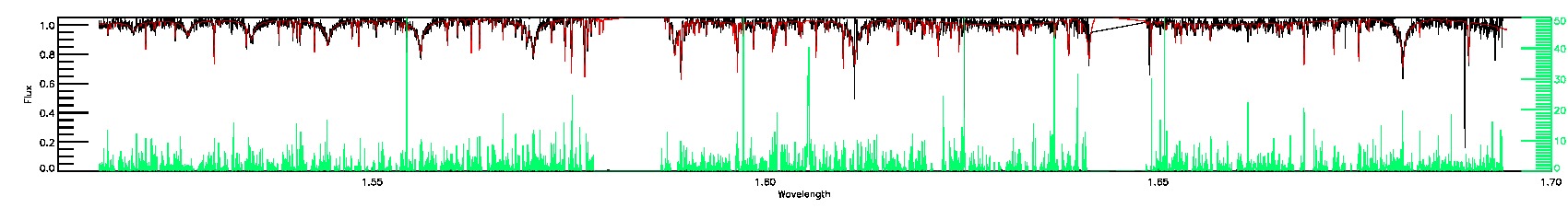
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5472. | +/- | 25. |
| -10000. | +/- | -NaN |
| 1.64 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.96 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.15 |
| -9999.00 |
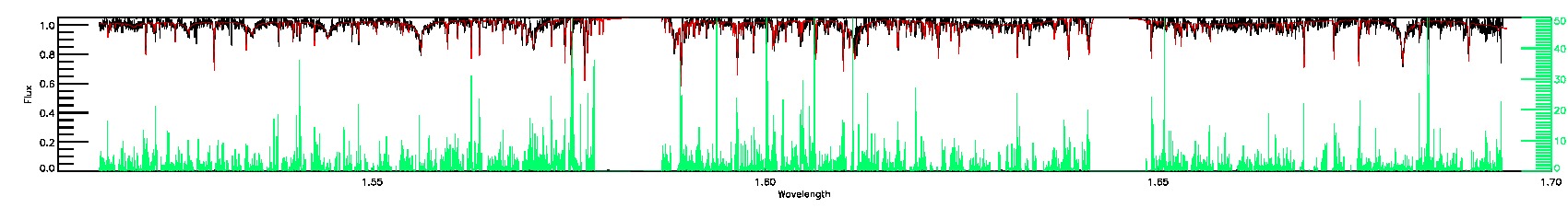
280-33-O
| 5493. | +/- | 10. |
| 5473. | +/- | 108. |
| 4.02 | +/- | 0. |
| 4.05 | +/- | 0. |
| 1.22 | +/- | 0. |
| 1.22 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -0.15 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.08 | +/- | 0. |
| 3.24 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.57 |
| -9999.00 |
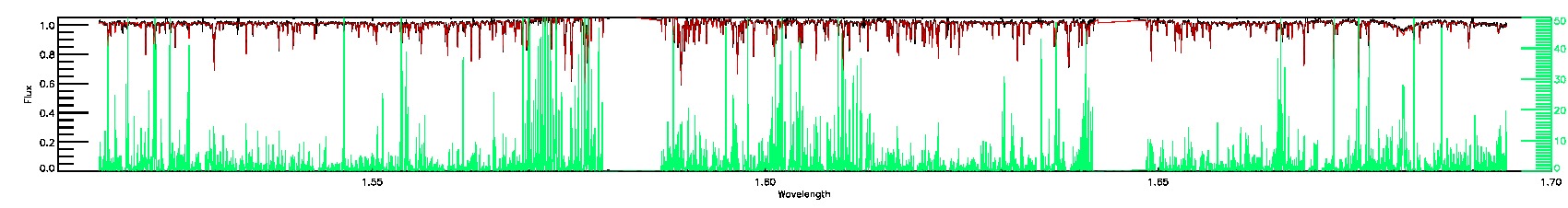
280-33-O
| 3389. | +/- | 1. |
| 3501. | +/- | 60. |
| 0.18 | +/- | 0. |
| 0.11 | +/- | 0. |
| 2.87 | +/- | 0. |
| 2.87 | +/- | -NaN |
| -0.56 | +/- | 0. |
| -0.56 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.01 | +/- | 0. |
| 4.10 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| -0.96 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,ROTATION_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5613. | +/- | 34. |
| -10000. | +/- | -NaN |
| 1.69 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.66 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| 0.31 |
| -9999.00 |
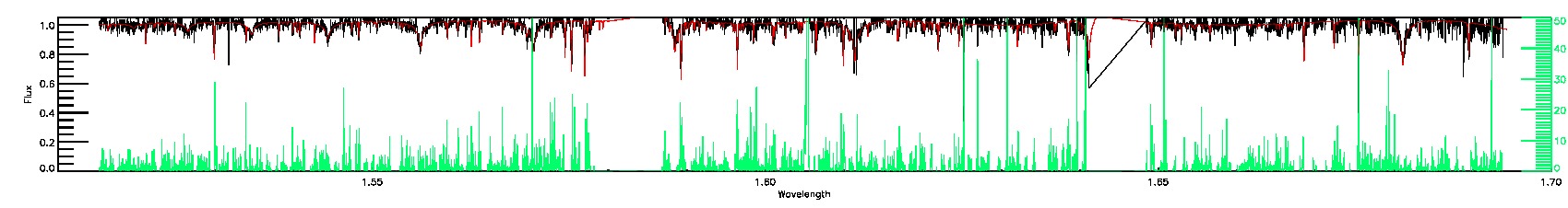
280-33-O
| 3875. | +/- | 1. |
| 3979. | +/- | 68. |
| 0.41 | +/- | 0. |
| 0.31 | +/- | 0. |
| 3.35 | +/- | 0. |
| 3.35 | +/- | -NaN |
| -0.37 | +/- | 0. |
| -0.37 | +/- | 0. |
| -0.25 | +/- | 0. |
| -0.25 | +/- | -NaN |
| 0.57 | +/- | 0. |
| 0.57 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.04 | +/- | 0. |
| 3.67 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.27 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.04 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5412. | +/- | 25. |
| -10000. | +/- | -NaN |
| 1.73 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.75 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.53 |
| -9999.00 |
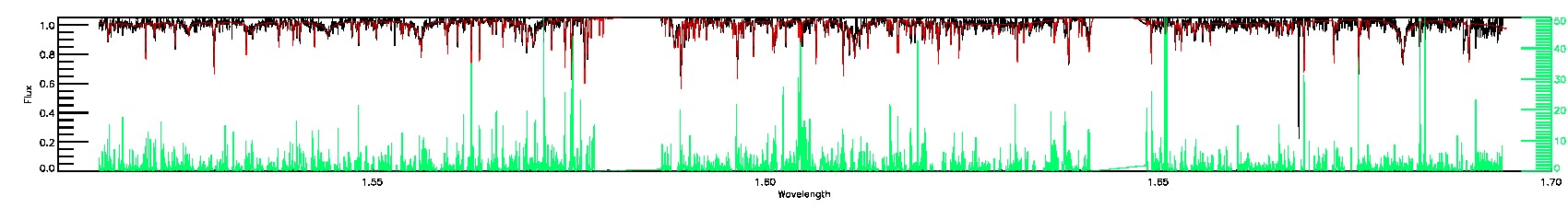
BRIGHT_NEIGHBOR
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN,SN_WARN
280-33-O
| 5297. | +/- | 15. |
| -10000. | +/- | -NaN |
| 1.30 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.49 | +/- | 0. |
| 4.49 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.49 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.14 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| 0.22 |
| -9999.00 |
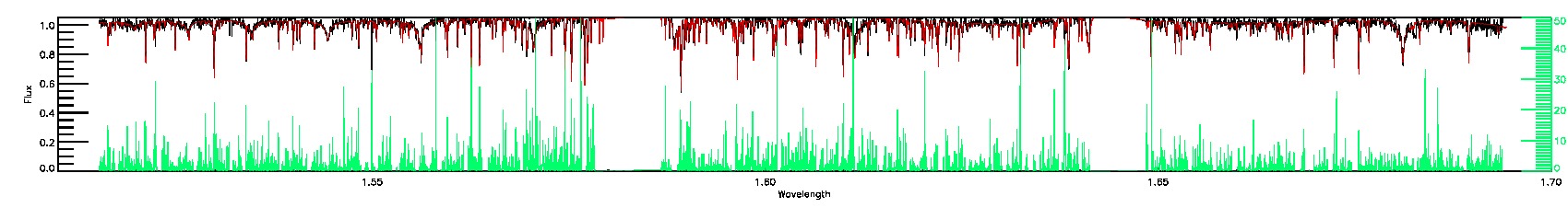
STAR_BAD,CHI2_BAD
STAR_WARN,CHI2_WARN
280-33-O
| 3084. | +/- | 1. |
| -10000. | +/- | -NaN |
| -0.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.87 | +/- | 0. |
| 3.87 | +/- | -NaN |
| -1.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.41 | +/- | 0. |
| 0.73 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.78 |
| -9999.00 |
STAR_BAD,CHI2_BAD
STAR_WARN,CHI2_WARN
280-33-O
| 3054. | +/- | 1. |
| -10000. | +/- | -NaN |
| 0.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.63 | +/- | 0. |
| 3.63 | +/- | -NaN |
| -1.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.41 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.47 | +/- | 0. |
| 0.74 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.41 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -1.24 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
280-33-O
| 5298. | +/- | 7. |
| 5291. | +/- | 99. |
| 4.52 | +/- | 0. |
| 4.49 | +/- | 0. |
| 1.26 | +/- | 0. |
| 1.26 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 1.76 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.00 |
| -9999.00 |

BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5795. | +/- | 22. |
| -10000. | +/- | -NaN |
| 1.64 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.93 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.36 |
| -9999.00 |

280-33-O
| 4379. | +/- | 2. |
| 4493. | +/- | 81. |
| 2.00 | +/- | 0. |
| 1.85 | +/- | 0. |
| 1.72 | +/- | 0. |
| 1.72 | +/- | -NaN |
| -0.60 | +/- | 0. |
| -0.60 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.26 | +/- | 0. |
| 4.20 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| -0.51 |
| -9999.00 |
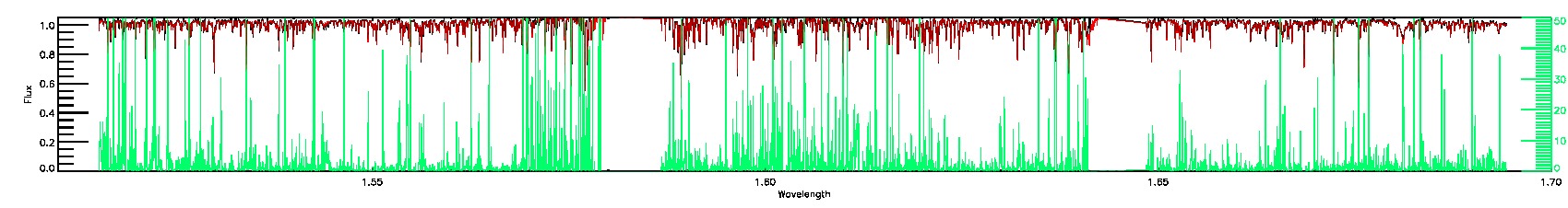
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5438. | +/- | 24. |
| -10000. | +/- | -NaN |
| 1.62 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.46 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.87 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.50 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| 0.50 |
| -9999.00 |

STAR_BAD
STAR_WARN,CHI2_WARN
280-33-O
| 3228. | +/- | 1. |
| -10000. | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.10 | +/- | 0. |
| 3.10 | +/- | -NaN |
| -0.93 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.08 | +/- | 0. |
| 0.71 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.51 |
| -9999.00 |
280-33-O
| 5264. | +/- | 7. |
| 5261. | +/- | 98. |
| 4.64 | +/- | 0. |
| 4.61 | +/- | 0. |
| 1.71 | +/- | 0. |
| 1.71 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 3.02 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.16 |
| -9999.00 |

BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,COLORTE_BAD,SN_BAD
STAR_WARN,COLORTE_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5375. | +/- | 52. |
| -10000. | +/- | -NaN |
| 1.55 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.76 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.61 | +/- | 0. |
| 0.66 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.23 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.44 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,ROTATION_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN
280-33-O
| 3000. | +/- | 0. |
| -10000. | +/- | -NaN |
| 0.91 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.64 | +/- | 0. |
| 0.64 | +/- | -NaN |
| -2.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.21 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.50 | +/- | 0. |
| 1.02 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| -2.20 |
| -9999.00 |
PERSIST_JUMP_NEG,SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,CHI2_WARN,COLORTE_WARN,ROTATION_WARN
280-33-O
| 3000. | +/- | 0. |
| -10000. | +/- | -NaN |
| 1.81 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -1.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.08 | +/- | 0. |
| 0.85 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -1.54 |
| -9999.00 |
STAR_BAD
STAR_WARN,COLORTE_WARN,SN_WARN
280-33-O
| 5477. | +/- | 23. |
| -10000. | +/- | -NaN |
| 1.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.74 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.18 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.10 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5555. | +/- | 26. |
| -10000. | +/- | -NaN |
| 1.80 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.53 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.02 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.20 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.87 |
| -9999.00 |

280-33-O
| 3808. | +/- | 1. |
| 3915. | +/- | 67. |
| -0.04 | +/- | 0. |
| -0.13 | +/- | 0. |
| 3.40 | +/- | 0. |
| 3.40 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -0.44 | +/- | 0. |
| -0.48 | +/- | 0. |
| -0.48 | +/- | -NaN |
| 0.56 | +/- | 0. |
| 0.56 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.06 | +/- | 0. |
| 3.83 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.50 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.11 |
| -9999.00 |
280-33-O
| 5433. | +/- | 9. |
| 5408. | +/- | 102. |
| 3.88 | +/- | 0. |
| 3.78 | +/- | 0. |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.04 | +/- | 0. |
| 2.76 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.22 |
| -9999.00 |

BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,ROTATION_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5758. | +/- | 25. |
| -10000. | +/- | -NaN |
| 1.73 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.81 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.35 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.75 | +/- | 0. |
| 0.68 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| 0.01 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5719. | +/- | 22. |
| -10000. | +/- | -NaN |
| 1.80 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.78 | +/- | 0. |
| 4.78 | +/- | -NaN |
| -0.55 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.09 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.20 |
| -9999.00 |
280-33-O
| 4387. | +/- | 2. |
| 4475. | +/- | 76. |
| 2.16 | +/- | 0. |
| 1.92 | +/- | 0. |
| 1.69 | +/- | 0. |
| 1.69 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.04 | +/- | 0. |
| 2.94 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.22 |
| -9999.00 |

STAR_BAD,CHI2_BAD
STAR_WARN,CHI2_WARN
280-33-O
| 3002. | +/- | 0. |
| -10000. | +/- | -NaN |
| 1.22 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.24 | +/- | 0. |
| 4.24 | +/- | -NaN |
| -1.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.56 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.54 | +/- | 0. |
| 0.74 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.55 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -0.87 |
| -9999.00 |
STAR_WARN,SN_WARN
280-33-O
| 5876. | +/- | 17. |
| 5822. | +/- | 167. |
| 1.56 | +/- | 0. |
| 1.41 | +/- | 0. |
| 4.39 | +/- | 0. |
| 4.39 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -0.40 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.04 | +/- | 1. |
| 0.04 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.09 | +/- | 0. |
| 3.74 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.40 |
| -9999.00 |

280-33-O
| 6279. | +/- | 16. |
| 6171. | +/- | 137. |
| 4.17 | +/- | 0. |
| 4.08 | +/- | 0. |
| 1.64 | +/- | 0. |
| 1.64 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -0.22 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.07 | +/- | 0. |
| 7.17 | +/- | 0. |
| 0.86 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.17 |
| -9999.00 |
280-33-O
| 3731. | +/- | 1. |
| 3841. | +/- | 66. |
| 0.01 | +/- | 0. |
| -0.08 | +/- | 0. |
| 3.55 | +/- | 0. |
| 3.55 | +/- | -NaN |
| -0.51 | +/- | 0. |
| -0.51 | +/- | 0. |
| -0.20 | +/- | 0. |
| -0.20 | +/- | -NaN |
| 0.37 | +/- | 0. |
| 0.37 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.06 | +/- | 0. |
| 3.99 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.21 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -1.15 |
| -9999.00 |
280-33-O
| 5968. | +/- | 16. |
| 5895. | +/- | 124. |
| 4.18 | +/- | 0. |
| 4.13 | +/- | 0. |
| 1.30 | +/- | 0. |
| 1.30 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -0.20 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.32 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |

STAR_BAD,CHI2_BAD
STAR_WARN,CHI2_WARN
280-33-O
| 3000. | +/- | 0. |
| -10000. | +/- | -NaN |
| 1.75 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.13 | +/- | 0. |
| 4.13 | +/- | -NaN |
| -0.53 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.03 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.70 |
| -9999.00 |
PERSIST_JUMP_NEG
STAR_BAD,CHI2_BAD
STAR_WARN,CHI2_WARN
280-33-O
| 3000. | +/- | 0. |
| -10000. | +/- | -NaN |
| 0.72 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.37 | +/- | 0. |
| 4.37 | +/- | -NaN |
| -1.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.84 | +/- | 0. |
| 0.77 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.39 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -1.13 |
| -9999.00 |
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN,SN_WARN
280-33-O
| 5606. | +/- | 18. |
| -10000. | +/- | -NaN |
| 1.79 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.81 | +/- | 0. |
| 3.81 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.41 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.85 |
| -9999.00 |
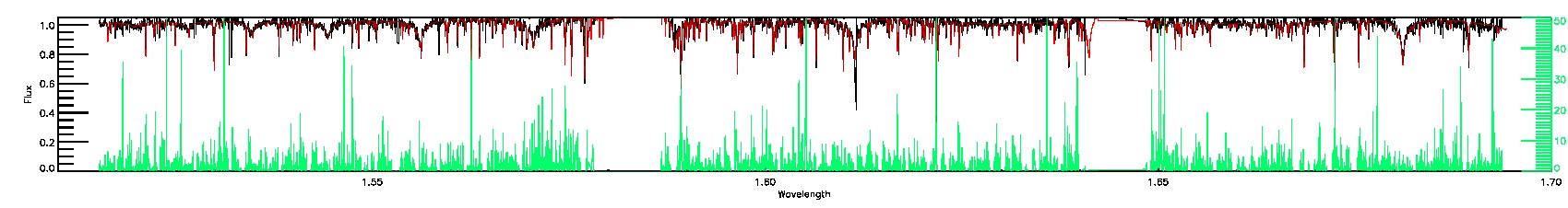
SUSPECT_BROAD_LINES
STAR_BAD,COLORTE_BAD,ROTATION_BAD
STAR_WARN,COLORTE_WARN,ROTATION_WARN
280-33-O
| 5354. | +/- | 5. |
| -10000. | +/- | -NaN |
| 0.80 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.74 | +/- | 0. |
| 1.74 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.68 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.76 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.43 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -2.10 |
| -9999.00 |

STAR_BAD,CHI2_BAD
STAR_WARN,CHI2_WARN,COLORTE_WARN
280-33-O
| 3001. | +/- | 0. |
| -10000. | +/- | -NaN |
| 1.79 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.81 | +/- | 0. |
| 3.81 | +/- | -NaN |
| -0.52 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.01 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.72 |
| -9999.00 |
280-33-O
| 5903. | +/- | 10. |
| 5824. | +/- | 118. |
| 4.43 | +/- | 0. |
| 4.26 | +/- | 0. |
| 1.48 | +/- | 0. |
| 1.48 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | 0. |
| -0.14 | +/- | 0. |
| -0.14 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.47 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.03 |
| -9999.00 |

STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN,SN_WARN
280-33-O
| 5547. | +/- | 24. |
| -10000. | +/- | -NaN |
| 1.79 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.77 | +/- | 0. |
| 4.77 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.66 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| 0.29 |
| -9999.00 |

280-33-O
| 3858. | +/- | 1. |
| 3962. | +/- | 68. |
| 0.51 | +/- | 0. |
| 0.41 | +/- | 0. |
| 2.65 | +/- | 0. |
| 2.65 | +/- | -NaN |
| -0.39 | +/- | 0. |
| -0.39 | +/- | 0. |
| -0.20 | +/- | 0. |
| -0.20 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.04 | +/- | 0. |
| 3.71 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.21 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.98 |
| -9999.00 |
STAR_BAD,CHI2_BAD
STAR_WARN,CHI2_WARN
280-33-O
| 3000. | +/- | 0. |
| -10000. | +/- | -NaN |
| 0.54 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.24 | +/- | 0. |
| 3.24 | +/- | -NaN |
| -0.84 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.22 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.32 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.85 | +/- | 0. |
| 0.69 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.21 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.70 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD
STAR_WARN,SN_WARN
280-33-O
| 5226. | +/- | 32. |
| -10000. | +/- | -NaN |
| 1.69 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.79 | +/- | 0. |
| 4.79 | +/- | -NaN |
| -0.46 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.88 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -2.41 |
| -9999.00 |
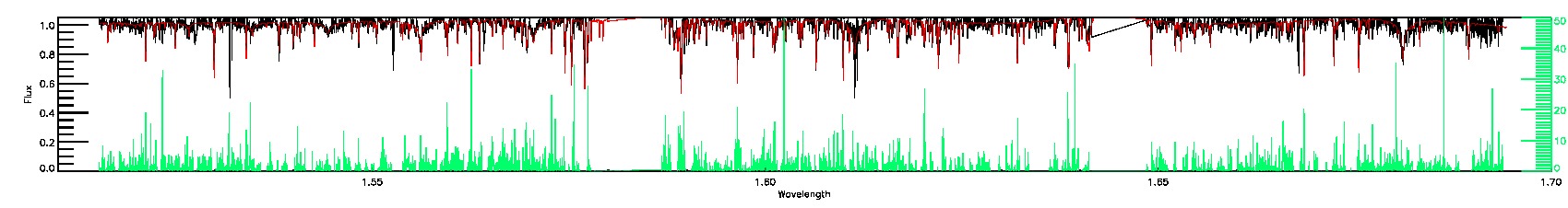
280-33-O
| 4607. | +/- | 3. |
| 4689. | +/- | 82. |
| 2.66 | +/- | 0. |
| 2.36 | +/- | 0. |
| 1.32 | +/- | 0. |
| 1.32 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.13 | +/- | 0. |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.15 | +/- | 0. |
| 3.19 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.24 |
| -9999.00 |
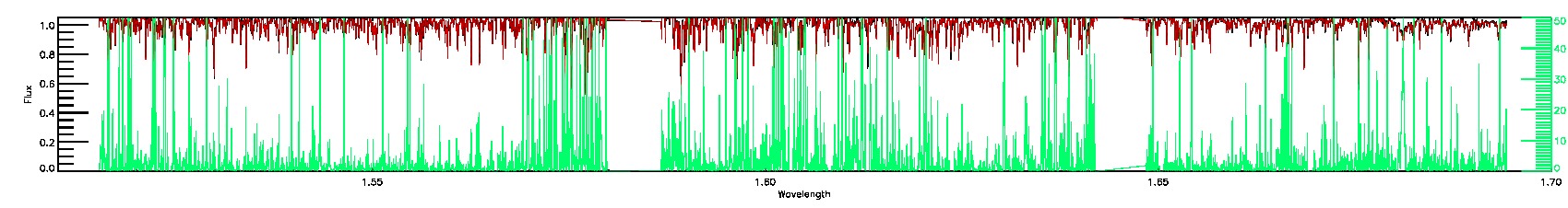
280-33-O
| 3693. | +/- | 1. |
| 3799. | +/- | 65. |
| -0.13 | +/- | 0. |
| -0.22 | +/- | 0. |
| 3.40 | +/- | 0. |
| 3.40 | +/- | -NaN |
| -0.43 | +/- | 0. |
| -0.43 | +/- | 0. |
| -0.24 | +/- | 0. |
| -0.24 | +/- | -NaN |
| 0.37 | +/- | 0. |
| 0.37 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.06 | +/- | 0. |
| 3.80 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.26 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.87 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN
280-33-O
| 5713. | +/- | 5. |
| -10000. | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.46 | +/- | 0. |
| 0.46 | +/- | -NaN |
| -2.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| -0.39 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.25 | +/- | 0. |
| 1.09 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.25 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| -2.27 |
| -9999.00 |
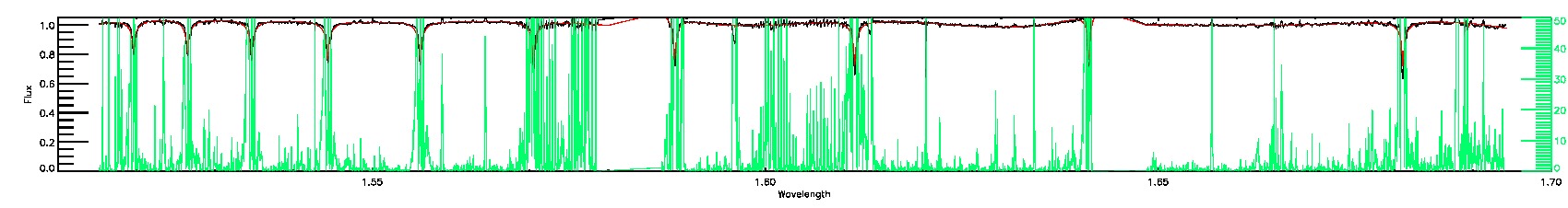
STAR_WARN,COLORTE_WARN
280-33-O
| 4054. | +/- | 2. |
| 4159. | +/- | 88. |
| 0.63 | +/- | 0. |
| 0.53 | +/- | 0. |
| 3.09 | +/- | 0. |
| 3.09 | +/- | -NaN |
| -0.38 | +/- | 0. |
| -0.38 | +/- | 0. |
| -0.20 | +/- | 0. |
| -0.20 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.06 | +/- | 0. |
| 3.70 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.22 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -1.33 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,COLORTE_BAD,ROTATION_BAD
STAR_WARN,COLORTE_WARN,ROTATION_WARN
280-33-O
| 5463. | +/- | 11. |
| -10000. | +/- | -NaN |
| 1.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.33 | +/- | 0. |
| 4.33 | +/- | -NaN |
| -0.66 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.36 | +/- | 0. |
| 0.64 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.02 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
280-33-O
| 5984. | +/- | 13. |
| 5900. | +/- | 122. |
| 4.14 | +/- | 0. |
| 4.00 | +/- | 0. |
| 1.56 | +/- | 0. |
| 1.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.00 | +/- | 0. |
| 5.58 | +/- | 0. |
| 0.75 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.04 |
| -9999.00 |
STAR_BAD,CHI2_BAD
STAR_WARN,CHI2_WARN
280-33-O
| 3007. | +/- | 0. |
| -10000. | +/- | -NaN |
| 1.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.51 | +/- | 0. |
| 3.51 | +/- | -NaN |
| -1.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.45 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.95 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.56 | +/- | 0. |
| 0.82 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.44 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -1.36 |
| -9999.00 |
280-33-O
| 4148. | +/- | 2. |
| 4273. | +/- | 80. |
| 1.50 | +/- | 0. |
| 1.42 | +/- | 0. |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -0.85 | +/- | 0. |
| -0.85 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.06 | +/- | 0. |
| 4.86 | +/- | 0. |
| 0.69 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -1.50 |
| -9999.00 |
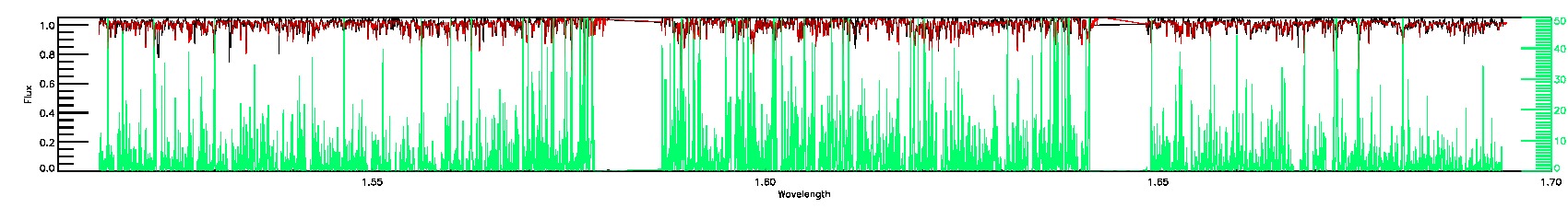
STAR_BAD
STAR_WARN,COLORTE_WARN
280-33-O
| 6644. | +/- | 4. |
| -10000. | +/- | -NaN |
| 2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.41 | +/- | 0. |
| 4.41 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 17.36 | +/- | 0. |
| 1.24 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.45 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -1.44 |
| -9999.00 |
280-33-O
| 3792. | +/- | 1. |
| 3899. | +/- | 67. |
| 0.23 | +/- | 0. |
| 0.13 | +/- | 0. |
| 3.64 | +/- | 0. |
| 3.64 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -0.44 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.22 | +/- | -NaN |
| 0.52 | +/- | 0. |
| 0.52 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.06 | +/- | 0. |
| 3.83 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.24 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -1.37 |
| -9999.00 |
280-33-O
| 5726. | +/- | 11. |
| 5673. | +/- | 113. |
| 4.49 | +/- | 0. |
| 4.42 | +/- | 0. |
| 1.25 | +/- | 0. |
| 1.25 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.04 | +/- | 0. |
| 3.54 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.07 |
| -9999.00 |

BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5591. | +/- | 27. |
| 5570. | +/- | 157. |
| 1.98 | +/- | 0. |
| 1.80 | +/- | 0. |
| 4.31 | +/- | 0. |
| 4.31 | +/- | -NaN |
| -0.41 | +/- | 0. |
| -0.41 | +/- | 0. |
| -0.16 | +/- | 0. |
| -0.16 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.76 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.77 |
| -9999.00 |
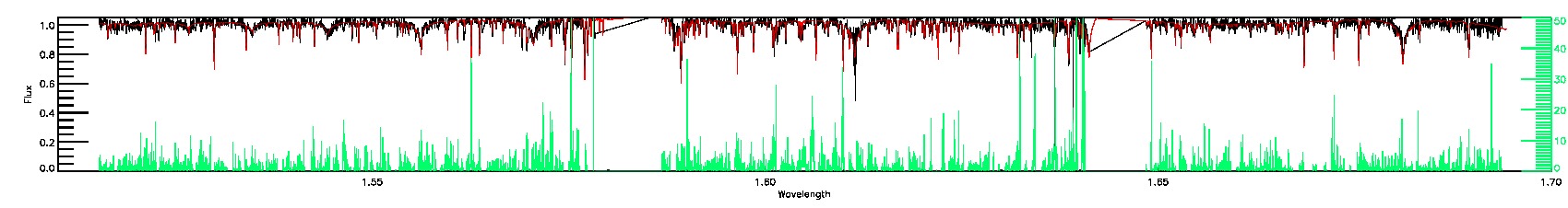
280-33-O
| 3999. | +/- | 1. |
| 4104. | +/- | 71. |
| 0.54 | +/- | 0. |
| 0.44 | +/- | 0. |
| 3.28 | +/- | 0. |
| 3.28 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -0.40 | +/- | 0. |
| -0.39 | +/- | 0. |
| -0.39 | +/- | -NaN |
| 0.56 | +/- | 0. |
| 0.56 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.05 | +/- | 0. |
| 3.73 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.40 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.07 |
| -9999.00 |
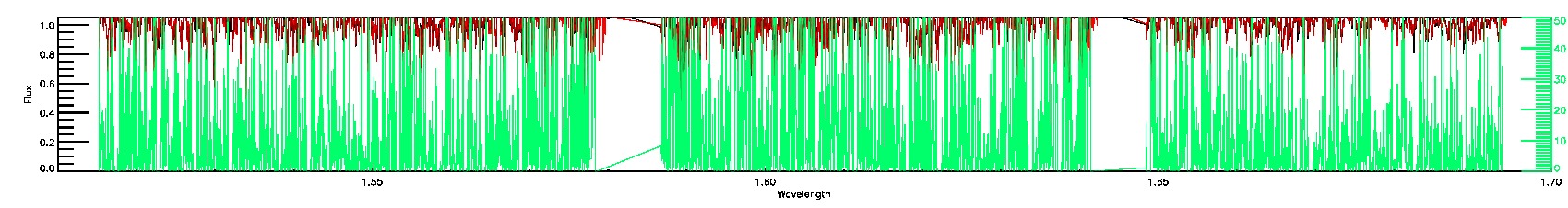
280-33-O
| 4202. | +/- | 1. |
| 4305. | +/- | 75. |
| 1.89 | +/- | 0. |
| 1.71 | +/- | 0. |
| 1.59 | +/- | 0. |
| 1.59 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -0.36 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.09 | +/- | 0. |
| 3.64 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.42 |
| -9999.00 |
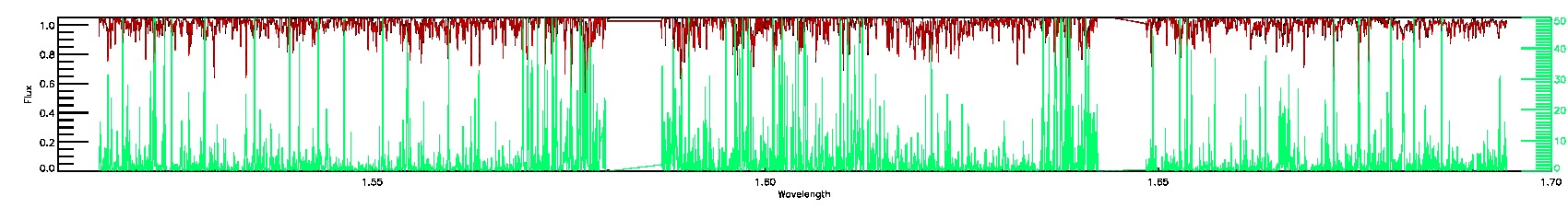
STAR_WARN,COLORTE_WARN
280-33-O
| 7836. | +/- | 12. |
| 7634. | +/- | 212. |
| 4.49 | +/- | 0. |
| 4.07 | +/- | 0. |
| 0.65 | +/- | 0. |
| 0.65 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 78.51 | +/- | 0. |
| 1.89 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| -0.38 |
| -9999.00 |
280-33-O
| 4972. | +/- | 6. |
| 5005. | +/- | 90. |
| 3.06 | +/- | 0. |
| 2.68 | +/- | 0. |
| 1.36 | +/- | 0. |
| 1.36 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.96 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.17 |
| -9999.00 |

280-33-O
| 3638. | +/- | 1. |
| 3746. | +/- | 64. |
| 0.13 | +/- | 0. |
| 0.04 | +/- | 0. |
| 2.67 | +/- | 0. |
| 2.67 | +/- | -NaN |
| -0.46 | +/- | 0. |
| -0.46 | +/- | 0. |
| -0.30 | +/- | 0. |
| -0.30 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.05 | +/- | 0. |
| 3.87 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.31 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.99 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
280-33-O
| 6440. | +/- | 16. |
| 6300. | +/- | 139. |
| 4.70 | +/- | 0. |
| 4.45 | +/- | 0. |
| 1.14 | +/- | 0. |
| 1.14 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 6.98 | +/- | 0. |
| 0.84 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.10 |
| -9999.00 |
280-33-O
| 5322. | +/- | 11. |
| 5332. | +/- | 105. |
| 3.77 | +/- | 0. |
| 3.81 | +/- | 0. |
| 1.08 | +/- | 0. |
| 1.08 | +/- | -NaN |
| -0.41 | +/- | 0. |
| -0.40 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.75 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.33 |
| -9999.00 |

280-33-O
| 3611. | +/- | 1. |
| 3717. | +/- | 63. |
| 0.21 | +/- | 0. |
| 0.11 | +/- | 0. |
| 2.66 | +/- | 0. |
| 2.66 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -0.40 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.22 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.03 | +/- | 0. |
| 3.74 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.25 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| -1.28 |
| -9999.00 |
PERSIST_JUMP_NEG
STAR_BAD
STAR_WARN,CHI2_WARN
280-33-O
| 3000. | +/- | 0. |
| -10000. | +/- | -NaN |
| 1.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.06 | +/- | 0. |
| 4.06 | +/- | -NaN |
| -1.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.70 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.83 | +/- | 0. |
| 0.77 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.69 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -1.04 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,COLORTE_BAD,SN_BAD
STAR_WARN,CHI2_WARN,COLORTE_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5328. | +/- | 55. |
| -10000. | +/- | -NaN |
| 1.42 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.79 | +/- | 0. |
| 4.79 | +/- | -NaN |
| -0.63 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.29 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.35 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.75 |
| -9999.00 |
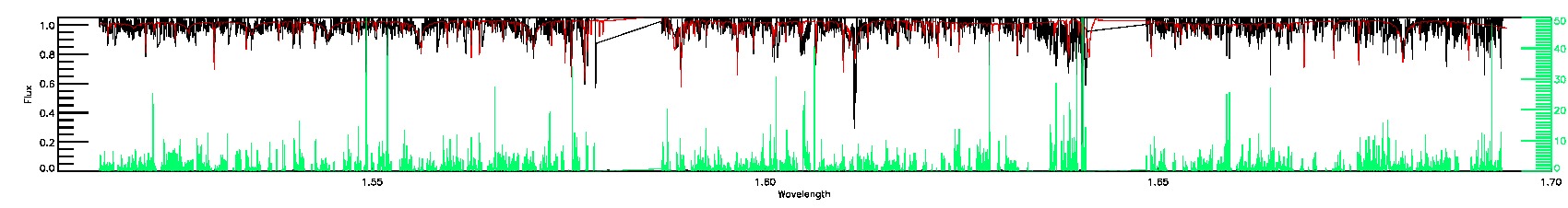
280-33-O
| 3836. | +/- | 1. |
| 3942. | +/- | 68. |
| 0.32 | +/- | 0. |
| 0.22 | +/- | 0. |
| 3.13 | +/- | 0. |
| 3.13 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -0.40 | +/- | 0. |
| -0.23 | +/- | 0. |
| -0.23 | +/- | -NaN |
| 0.47 | +/- | 0. |
| 0.47 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.04 | +/- | 0. |
| 3.74 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.24 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.27 |
| -9999.00 |
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN
280-33-O
| 5968. | +/- | 11. |
| -10000. | +/- | -NaN |
| 4.59 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.19 | +/- | 0. |
| 1.19 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.62 | +/- | 0. |
| 0.66 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.09 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
280-33-O
| 3451. | +/- | 1. |
| 3576. | +/- | 69. |
| 0.07 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.50 | +/- | 0. |
| 2.50 | +/- | -NaN |
| -0.86 | +/- | 0. |
| -0.86 | +/- | 0. |
| -0.21 | +/- | 0. |
| -0.21 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 4.88 | +/- | 0. |
| 0.69 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.22 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -1.78 |
| -9999.00 |
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN
280-33-O
| 4845. | +/- | 5. |
| -10000. | +/- | -NaN |
| 1.30 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.83 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.09 |
| -9999.00 |
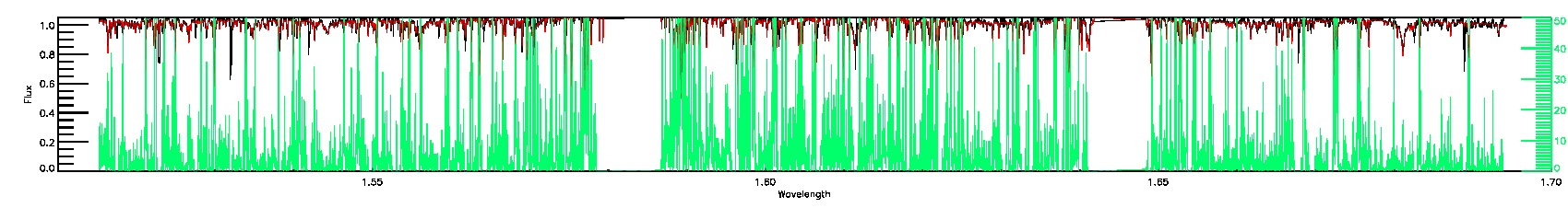
STAR_BAD
STAR_WARN,COLORTE_WARN
280-33-O
| 4991. | +/- | 3. |
| -10000. | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.28 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.22 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.33 |
| -9999.00 |

280-33-O
| 5952. | +/- | 11. |
| 5873. | +/- | 121. |
| 4.15 | +/- | 0. |
| 4.03 | +/- | 0. |
| 1.80 | +/- | 0. |
| 1.80 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 5.39 | +/- | 0. |
| 0.73 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |

280-33-O
| 3350. | +/- | 1. |
| 3458. | +/- | 59. |
| 0.34 | +/- | 0. |
| 0.25 | +/- | 0. |
| 3.50 | +/- | 0. |
| 3.50 | +/- | -NaN |
| -0.46 | +/- | 0. |
| -0.46 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.04 | +/- | 0. |
| 3.87 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.84 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
280-33-O
| 3568. | +/- | 1. |
| 3676. | +/- | 63. |
| 0.10 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.57 | +/- | 0. |
| 2.57 | +/- | -NaN |
| -0.45 | +/- | 0. |
| -0.45 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.22 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.04 | +/- | 0. |
| 3.85 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.24 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -1.35 |
| -9999.00 |
280-33-O
| 5365. | +/- | 11. |
| 5365. | +/- | 105. |
| 4.56 | +/- | 0. |
| 4.66 | +/- | 0. |
| 1.48 | +/- | 0. |
| 1.48 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -0.28 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.18 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.45 |
| -9999.00 |

280-33-O
| 3802. | +/- | 1. |
| 3910. | +/- | 68. |
| 0.26 | +/- | 0. |
| 0.17 | +/- | 0. |
| 3.19 | +/- | 0. |
| 3.19 | +/- | -NaN |
| -0.47 | +/- | 0. |
| -0.47 | +/- | 0. |
| -0.21 | +/- | 0. |
| -0.21 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.05 | +/- | 0. |
| 3.88 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.23 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -1.40 |
| -9999.00 |
STAR_BAD,CHI2_BAD
STAR_WARN,CHI2_WARN
280-33-O
| 3221. | +/- | 0. |
| -10000. | +/- | -NaN |
| -0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.36 | +/- | 0. |
| 3.36 | +/- | -NaN |
| -0.53 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.04 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.16 |
| -9999.00 |
STAR_BAD
STAR_WARN,COLORTE_WARN
280-33-O
| 3394. | +/- | 1. |
| -10000. | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.25 | +/- | 0. |
| 3.25 | +/- | -NaN |
| -0.53 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.03 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.25 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| -0.97 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
280-33-O
| 5107. | +/- | 44. |
| -10000. | +/- | -NaN |
| 1.63 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.79 | +/- | 0. |
| 4.79 | +/- | -NaN |
| -0.39 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.72 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| 0.87 |
| -9999.00 |

280-33-O
| 5875. | +/- | 22. |
| 5815. | +/- | 132. |
| 4.28 | +/- | 0. |
| 4.28 | +/- | 0. |
| 1.49 | +/- | 0. |
| 1.49 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -0.26 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.15 | +/- | 0. |
| 15.42 | +/- | 0. |
| 1.19 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| -0.96 |
| -9999.00 |
280-33-O
| 5551. | +/- | 9. |
| 5513. | +/- | 106. |
| 4.07 | +/- | 0. |
| 3.97 | +/- | 0. |
| 0.98 | +/- | 0. |
| 0.98 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.78 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.16 |
| -9999.00 |

PERSIST_JUMP_NEG
STAR_BAD,CHI2_BAD
STAR_WARN,CHI2_WARN,COLORTE_WARN,SN_WARN
280-33-O
| 3006. | +/- | 2. |
| -10000. | +/- | -NaN |
| 1.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.05 | +/- | 0. |
| 4.05 | +/- | -NaN |
| -0.69 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.42 | +/- | 0. |
| 0.65 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -0.87 |
| -9999.00 |
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN
280-33-O
| 6351. | +/- | 19. |
| -10000. | +/- | -NaN |
| 4.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.13 | +/- | 0. |
| 1.13 | +/- | -NaN |
| -0.30 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.21 | +/- | 0. |
| 0.72 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.03 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN,ROTATION_WARN
280-33-O
| 5663. | +/- | 10. |
| -10000. | +/- | -NaN |
| 1.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.82 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.75 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.19 |
| -9999.00 |

SUSPECT_BROAD_LINES
280-33-O
| 6709. | +/- | 16. |
| 6549. | +/- | 155. |
| 4.47 | +/- | 0. |
| 4.28 | +/- | 0. |
| 0.45 | +/- | 0. |
| 0.45 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -0.19 | +/- | 0. |
| -0.16 | +/- | 0. |
| -0.16 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 21.76 | +/- | 0. |
| 1.34 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.72 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD
STAR_WARN,SN_WARN
280-33-O
| 5703. | +/- | 28. |
| -10000. | +/- | -NaN |
| 1.73 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.60 | +/- | 0. |
| 4.60 | +/- | -NaN |
| -0.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.20 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.90 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.24 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.10 |
| -9999.00 |

280-33-O
| 5059. | +/- | 8. |
| 5098. | +/- | 97. |
| 3.51 | +/- | 0. |
| 3.40 | +/- | 0. |
| 0.55 | +/- | 0. |
| 0.55 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -0.36 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.65 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.06 |
| -9999.00 |

280-33-O
| 3450. | +/- | 1. |
| 3561. | +/- | 61. |
| -0.23 | +/- | 0. |
| -0.32 | +/- | 0. |
| 3.04 | +/- | 0. |
| 3.04 | +/- | -NaN |
| -0.52 | +/- | 0. |
| -0.52 | +/- | 0. |
| -0.23 | +/- | 0. |
| -0.23 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.05 | +/- | 0. |
| 4.00 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.25 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -1.51 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5641. | +/- | 20. |
| -10000. | +/- | -NaN |
| 1.67 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.92 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.18 |
| -9999.00 |
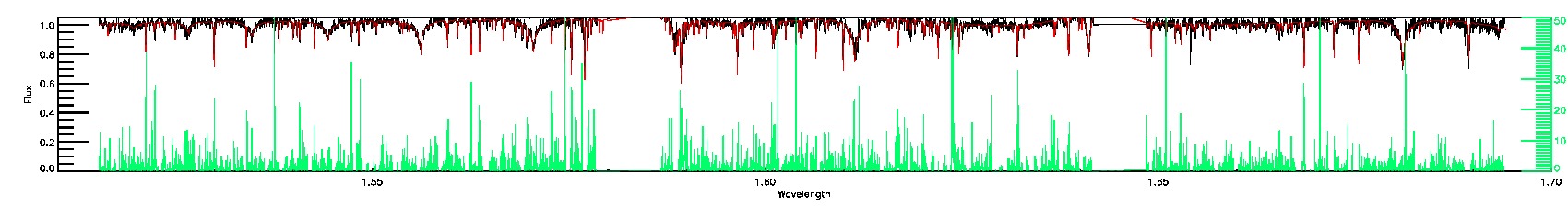
280-33-O
| 5131. | +/- | 6. |
| 5142. | +/- | 94. |
| 4.54 | +/- | 0. |
| 4.52 | +/- | 0. |
| 1.28 | +/- | 0. |
| 1.28 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 1.61 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.15 |
| -9999.00 |

STAR_BAD,CHI2_BAD
STAR_WARN,CHI2_WARN
280-33-O
| 3001. | +/- | 0. |
| -10000. | +/- | -NaN |
| 0.62 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.30 | +/- | 0. |
| 3.30 | +/- | -NaN |
| -0.76 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.30 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.45 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.62 | +/- | 0. |
| 0.66 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.29 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.82 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,COLORTE_BAD,ROTATION_BAD
STAR_WARN,COLORTE_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5784. | +/- | 30. |
| -10000. | +/- | -NaN |
| 2.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.62 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.24 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -2.36 |
| -9999.00 |

280-33-O
| 6480. | +/- | 14. |
| 6331. | +/- | 139. |
| 4.08 | +/- | 0. |
| 3.78 | +/- | 0. |
| 2.40 | +/- | 0. |
| 2.40 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | 0. |
| -0.10 | +/- | 0. |
| -0.10 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.06 | +/- | 0. |
| 7.06 | +/- | 0. |
| 0.85 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.26 |
| -9999.00 |
280-33-O
| 6018. | +/- | 13. |
| 5929. | +/- | 123. |
| 4.47 | +/- | 0. |
| 4.32 | +/- | 0. |
| 0.86 | +/- | 0. |
| 0.86 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 3.49 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.10 |
| -9999.00 |
SUSPECT_RV_COMBINATION,SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,COLORTE_WARN
280-33-O
| 7446. | +/- | 12. |
| -10000. | +/- | -NaN |
| 4.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.50 | +/- | 0. |
| 4.50 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 72.53 | +/- | 0. |
| 1.86 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.14 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.15 |
| -9999.00 |
STAR_BAD
STAR_WARN,CHI2_WARN
280-33-O
| 3000. | +/- | 0. |
| -10000. | +/- | -NaN |
| 0.56 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.71 | +/- | 0. |
| 3.71 | +/- | -NaN |
| -1.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.80 | +/- | 0. |
| 0.76 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.43 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.81 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,ROTATION_BAD
STAR_WARN,COLORTE_WARN,ROTATION_WARN
280-33-O
| 5909. | +/- | 7. |
| -10000. | +/- | -NaN |
| 1.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.58 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.14 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.28 |
| -9999.00 |
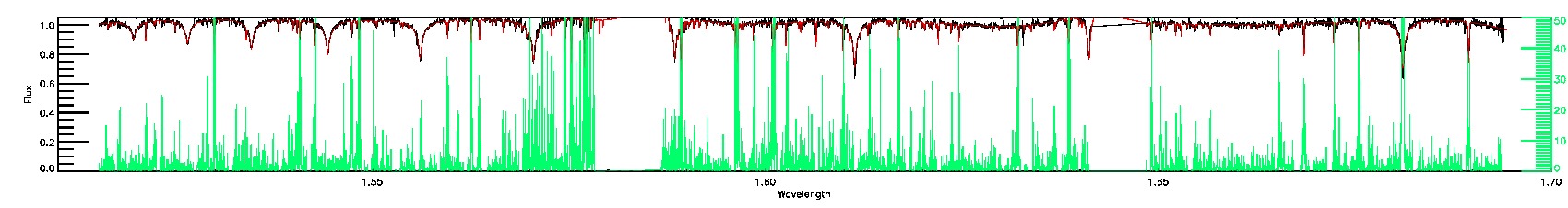
280-33-O
| 3966. | +/- | 2. |
| 4072. | +/- | 89. |
| 0.69 | +/- | 0. |
| 0.60 | +/- | 0. |
| 3.04 | +/- | 0. |
| 3.04 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -0.40 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.04 | +/- | 0. |
| 3.74 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -1.36 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD
STAR_WARN,SN_WARN
280-33-O
| 5440. | +/- | 26. |
| -10000. | +/- | -NaN |
| 1.86 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.78 | +/- | 0. |
| 4.78 | +/- | -NaN |
| -0.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.70 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.50 |
| -9999.00 |

STAR_BAD,CHI2_BAD
STAR_WARN,CHI2_WARN
280-33-O
| 3000. | +/- | 0. |
| -10000. | +/- | -NaN |
| 2.99 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.86 | +/- | 0. |
| 3.86 | +/- | -NaN |
| -0.97 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.23 | +/- | 0. |
| 0.72 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.73 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.91 |
| -9999.00 |
280-33-O
| 6257. | +/- | 20. |
| 6154. | +/- | 137. |
| 4.38 | +/- | 0. |
| 4.31 | +/- | 0. |
| 1.20 | +/- | 0. |
| 1.20 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -0.28 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 6.08 | +/- | 0. |
| 0.78 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.26 |
| -9999.00 |
280-33-O
| 6084. | +/- | 15. |
| 5991. | +/- | 127. |
| 4.58 | +/- | 0. |
| 4.45 | +/- | 0. |
| 0.77 | +/- | 0. |
| 0.77 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.19 | +/- | 0. |
| -0.19 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.24 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| 5.43 | +/- | 0. |
| 0.73 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.09 |
| -9999.00 |
280-33-O
| 3508. | +/- | 1. |
| 3615. | +/- | 62. |
| 0.07 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.82 | +/- | 0. |
| 2.82 | +/- | -NaN |
| -0.43 | +/- | 0. |
| -0.43 | +/- | 0. |
| -0.23 | +/- | 0. |
| -0.23 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.08 | +/- | 0. |
| 3.81 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.25 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| -1.12 |
| -9999.00 |
280-33-O
| 4024. | +/- | 1. |
| 4129. | +/- | 78. |
| 0.68 | +/- | 0. |
| 0.58 | +/- | 0. |
| 2.77 | +/- | 0. |
| 2.77 | +/- | -NaN |
| -0.39 | +/- | 0. |
| -0.39 | +/- | 0. |
| -0.21 | +/- | 0. |
| -0.21 | +/- | -NaN |
| 0.43 | +/- | 0. |
| 0.43 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.04 | +/- | 0. |
| 3.72 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.23 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.53 |
| -9999.00 |
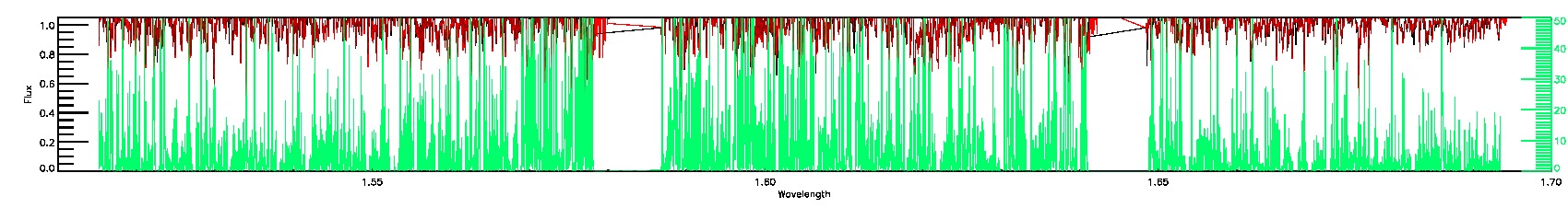
280-33-O
| 6245. | +/- | 18. |
| 6139. | +/- | 135. |
| 4.54 | +/- | 0. |
| 4.43 | +/- | 0. |
| 0.75 | +/- | 0. |
| 0.75 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -0.19 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.01 | +/- | 0. |
| 4.93 | +/- | 0. |
| 0.69 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -0.27 |
| -9999.00 |
280-33-O
| 4182. | +/- | 1. |
| 4294. | +/- | 76. |
| 1.97 | +/- | 0. |
| 1.81 | +/- | 0. |
| 1.53 | +/- | 0. |
| 1.53 | +/- | -NaN |
| -0.53 | +/- | 0. |
| -0.53 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.22 | +/- | 0. |
| 4.04 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.74 |
| -9999.00 |
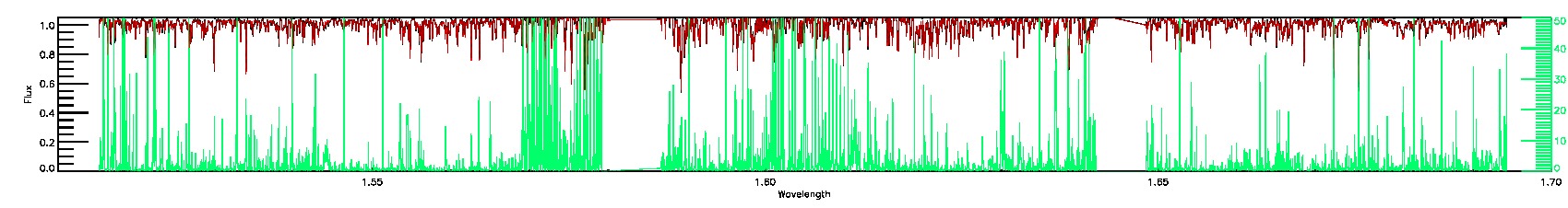
280-33-O
| 5455. | +/- | 8. |
| 5433. | +/- | 105. |
| 4.58 | +/- | 0. |
| 4.55 | +/- | 0. |
| 1.58 | +/- | 0. |
| 1.58 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 2.73 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
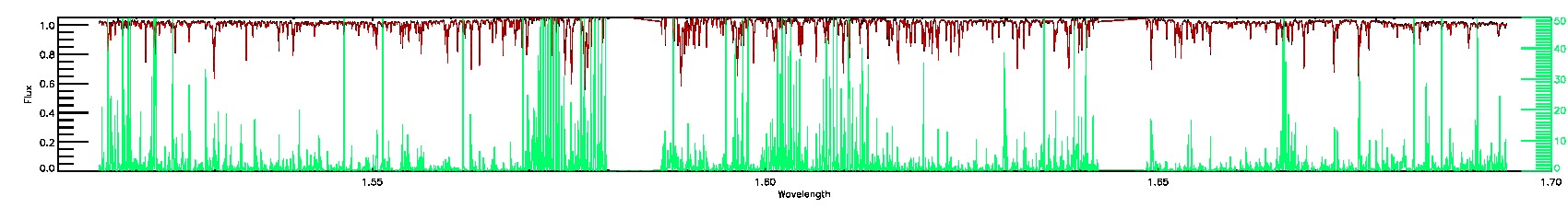
SUSPECT_BROAD_LINES
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN,SN_WARN
280-33-O
| 6791. | +/- | 14. |
| -10000. | +/- | -NaN |
| 2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.43 | +/- | 0. |
| 4.43 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 13.26 | +/- | 0. |
| 1.12 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -2.39 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
280-33-O
| 6390. | +/- | 19. |
| 6264. | +/- | 141. |
| 4.62 | +/- | 0. |
| 4.45 | +/- | 0. |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -0.09 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 3.47 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.42 |
| -9999.00 |
280-33-O
| 5928. | +/- | 12. |
| 5859. | +/- | 123. |
| 4.04 | +/- | 0. |
| 4.01 | +/- | 0. |
| 1.51 | +/- | 0. |
| 1.51 | +/- | -NaN |
| -0.21 | +/- | 0. |
| -0.21 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.04 | +/- | 0. |
| 3.70 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.18 |
| -9999.00 |

STAR_BAD
280-33-O
| 3422. | +/- | 1. |
| -10000. | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.94 | +/- | 0. |
| 2.94 | +/- | -NaN |
| -0.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.81 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.62 |
| -9999.00 |
280-33-O
| 6104. | +/- | 19. |
| 6028. | +/- | 134. |
| 4.46 | +/- | 0. |
| 4.51 | +/- | 0. |
| 1.01 | +/- | 0. |
| 1.01 | +/- | -NaN |
| -0.51 | +/- | 0. |
| -0.51 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.04 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.05 | +/- | 0. |
| 1.87 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.77 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD
STAR_WARN,SN_WARN
280-33-O
| 5248. | +/- | 23. |
| -10000. | +/- | -NaN |
| 1.61 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.91 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.43 |
| -9999.00 |
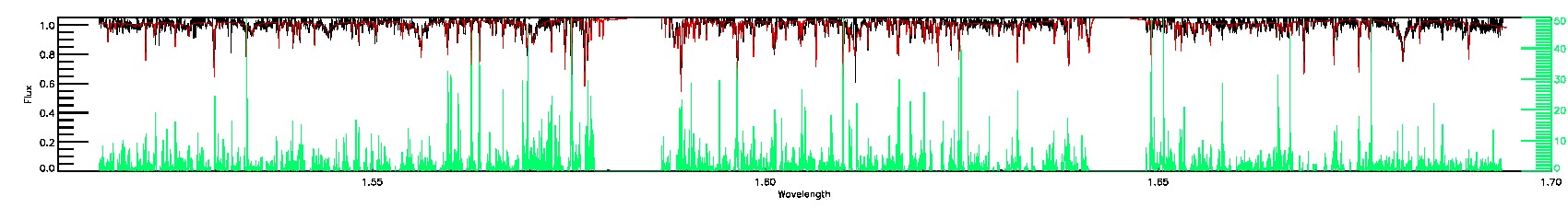
SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5648. | +/- | 18. |
| -10000. | +/- | -NaN |
| 1.55 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.79 | +/- | 0. |
| 4.79 | +/- | -NaN |
| -0.57 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.14 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.14 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.33 |
| -9999.00 |

STAR_BAD,CHI2_BAD
STAR_WARN,CHI2_WARN
280-33-O
| 3000. | +/- | 0. |
| -10000. | +/- | -NaN |
| 0.66 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.49 | +/- | 0. |
| 2.49 | +/- | -NaN |
| -1.60 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.39 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.55 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.43 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -1.59 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
280-33-O
| 3552. | +/- | 1. |
| 3661. | +/- | 63. |
| -0.02 | +/- | 0. |
| -0.10 | +/- | 0. |
| 2.81 | +/- | 0. |
| 2.81 | +/- | -NaN |
| -0.48 | +/- | 0. |
| -0.48 | +/- | 0. |
| -0.23 | +/- | 0. |
| -0.23 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.04 | +/- | 0. |
| 3.92 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.24 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -1.14 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN,ROTATION_WARN
280-33-O
| 5621. | +/- | 4. |
| -10000. | +/- | -NaN |
| 0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.32 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.44 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,ROTATION_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN
280-33-O
| 3071. | +/- | 1. |
| -10000. | +/- | -NaN |
| 2.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -1.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.63 | +/- | 0. |
| 0.75 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -1.14 |
| -9999.00 |
280-33-O
| 4810. | +/- | 5. |
| 4862. | +/- | 86. |
| 3.27 | +/- | 0. |
| 3.09 | +/- | 0. |
| 0.94 | +/- | 0. |
| 0.94 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.29 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.93 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.06 |
| -9999.00 |

BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN
280-33-O
| 5382. | +/- | 8. |
| -10000. | +/- | -NaN |
| 0.79 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.79 | +/- | 0. |
| 4.79 | +/- | -NaN |
| -0.41 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.30 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.75 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.45 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.18 |
| -9999.00 |

STAR_BAD,CHI2_BAD
STAR_WARN,CHI2_WARN
280-33-O
| 3032. | +/- | 1. |
| -10000. | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.38 | +/- | 0. |
| 3.38 | +/- | -NaN |
| -1.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.58 | +/- | 0. |
| 0.75 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.83 |
| -9999.00 |
280-33-O
| 5872. | +/- | 10. |
| 5802. | +/- | 119. |
| 3.70 | +/- | 0. |
| 3.57 | +/- | 0. |
| 1.29 | +/- | 0. |
| 1.29 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | 0. |
| -0.86 | +/- | 0. |
| -0.86 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 3.02 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.89 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.10 |
| -9999.00 |

STAR_WARN,SN_WARN
280-33-O
| 3792. | +/- | 2. |
| 3900. | +/- | 91. |
| 0.05 | +/- | 0. |
| -0.04 | +/- | 0. |
| 3.15 | +/- | 0. |
| 3.15 | +/- | -NaN |
| -0.47 | +/- | 0. |
| -0.47 | +/- | 0. |
| -0.24 | +/- | 0. |
| -0.24 | +/- | -NaN |
| 0.60 | +/- | 0. |
| 0.60 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.03 | +/- | 0. |
| 3.88 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.26 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.34 |
| -9999.00 |
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN
280-33-O
| 5793. | +/- | 8. |
| -10000. | +/- | -NaN |
| 4.42 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.86 | +/- | 0. |
| 0.86 | +/- | -NaN |
| 0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.14 | +/- | 0. |
| 0.33 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.05 |
| -9999.00 |
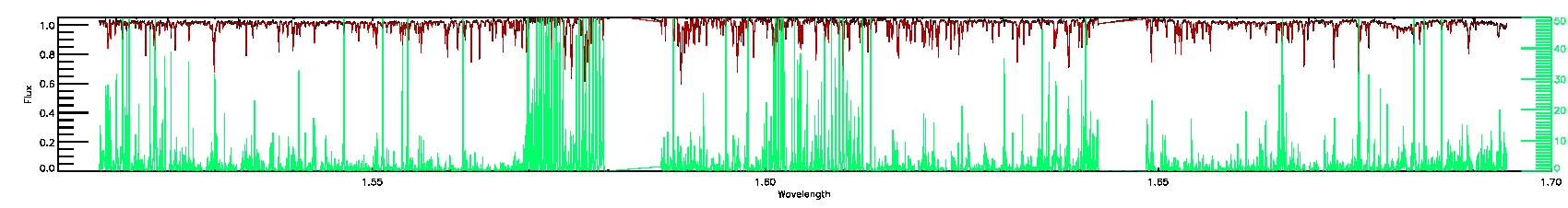
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5386. | +/- | 22. |
| -10000. | +/- | -NaN |
| 1.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.77 | +/- | 0. |
| 4.77 | +/- | -NaN |
| -0.42 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.77 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.09 |
| -9999.00 |

280-33-O
| 3402. | +/- | 1. |
| 3511. | +/- | 60. |
| -0.34 | +/- | 0. |
| -0.43 | +/- | 0. |
| 2.65 | +/- | 0. |
| 2.65 | +/- | -NaN |
| -0.48 | +/- | 0. |
| -0.48 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.04 | +/- | 0. |
| 3.92 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.62 |
| -9999.00 |
STAR_BAD,CHI2_BAD
STAR_WARN,CHI2_WARN,COLORTE_WARN
280-33-O
| 3068. | +/- | 1. |
| -10000. | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.42 | +/- | 0. |
| 3.42 | +/- | -NaN |
| -1.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.41 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.37 | +/- | 0. |
| 0.73 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -0.95 |
| -9999.00 |
280-33-O
| 5497. | +/- | 10. |
| 5477. | +/- | 108. |
| 4.48 | +/- | 0. |
| 4.51 | +/- | 0. |
| 0.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -0.16 | +/- | 0. |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.14 | +/- | 0. |
| 1.55 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.12 |
| -9999.00 |

280-33-O
| 3488. | +/- | 1. |
| 3590. | +/- | 61. |
| 0.15 | +/- | 0. |
| 0.04 | +/- | 0. |
| 2.59 | +/- | 0. |
| 2.59 | +/- | -NaN |
| -0.33 | +/- | 0. |
| -0.33 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.22 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.07 | +/- | 0. |
| 3.59 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.24 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -0.84 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN
280-33-O
| 4234. | +/- | 2. |
| -10000. | +/- | -NaN |
| 0.22 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.46 | +/- | 0. |
| 4.46 | +/- | -NaN |
| -0.53 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.60 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.03 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.38 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.09 |
| -9999.00 |
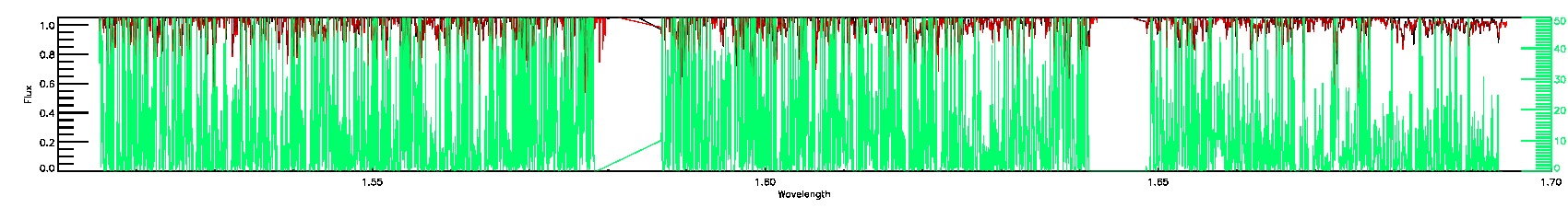
BRIGHT_NEIGHBOR
STAR_WARN,SN_WARN
280-33-O
| 5795. | +/- | 22. |
| 5750. | +/- | 168. |
| 1.63 | +/- | 0. |
| 1.47 | +/- | 0. |
| 4.04 | +/- | 0. |
| 4.04 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -0.40 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.07 | +/- | 1. |
| 0.07 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.11 | +/- | 0. |
| 3.73 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.65 |
| -9999.00 |

PERSIST_JUMP_NEG
STAR_BAD,CHI2_BAD
STAR_WARN,CHI2_WARN
280-33-O
| 3026. | +/- | 1. |
| -10000. | +/- | -NaN |
| 1.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.01 | +/- | 0. |
| 4.01 | +/- | -NaN |
| -0.97 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.39 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.22 | +/- | 0. |
| 0.72 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.38 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.94 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,COLORTE_WARN
280-33-O
| 6563. | +/- | 13. |
| -10000. | +/- | -NaN |
| 3.88 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.74 | +/- | 0. |
| 1.74 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 19.50 | +/- | 0. |
| 1.29 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.50 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -2.10 |
| -9999.00 |
BRIGHT_NEIGHBOR,LOW_SNR,SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,COLORTE_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,COLORTE_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5840. | +/- | 75. |
| -10000. | +/- | -NaN |
| 2.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.79 | +/- | 0. |
| 4.79 | +/- | -NaN |
| -0.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.35 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.21 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.66 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.51 |
| -9999.00 |

PERSIST_JUMP_NEG,SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN
280-33-O
| 3000. | +/- | 0. |
| -10000. | +/- | -NaN |
| 1.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -2.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.92 | +/- | 0. |
| 1.04 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.40 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.04 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_WARN,ROTATION_WARN
280-33-O
| 3732. | +/- | 1. |
| 3846. | +/- | 67. |
| -0.24 | +/- | 0. |
| -0.32 | +/- | 0. |
| 4.13 | +/- | 0. |
| 4.13 | +/- | -NaN |
| -0.60 | +/- | 0. |
| -0.60 | +/- | 0. |
| -0.25 | +/- | 0. |
| -0.25 | +/- | -NaN |
| 0.50 | +/- | 0. |
| 0.50 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.05 | +/- | 0. |
| 4.19 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.27 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.49 |
| -9999.00 |
STAR_BAD
STAR_WARN,COLORTE_WARN
280-33-O
| 3090. | +/- | 0. |
| -10000. | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.23 | +/- | 0. |
| 3.23 | +/- | -NaN |
| -0.67 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.39 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.38 | +/- | 0. |
| 0.64 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.69 |
| -9999.00 |
BAD_PIXELS,BRIGHT_NEIGHBOR,LOW_SNR,SUSPECT_RV_COMBINATION,BAD_RV_COMBINATION
STAR_BAD
STAR_WARN,SN_WARN
280-33-O
| 5970. | +/- | 26. |
| -10000. | +/- | -NaN |
| 1.59 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.79 | +/- | 0. |
| 4.79 | +/- | -NaN |
| -0.58 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.15 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.78 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
280-33-O
| 3456. | +/- | 1. |
| 3566. | +/- | 61. |
| -0.11 | +/- | 0. |
| -0.19 | +/- | 0. |
| 2.81 | +/- | 0. |
| 2.81 | +/- | -NaN |
| -0.51 | +/- | 0. |
| -0.51 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.06 | +/- | -NaN |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 3.98 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -1.16 |
| -9999.00 |
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN
280-33-O
| 7177. | +/- | 5. |
| -10000. | +/- | -NaN |
| 2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.78 | +/- | 1. |
| 0.78 | +/- | -NaN |
| -2.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 18.54 | +/- | 0. |
| 1.27 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -2.41 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN
280-33-O
| 6140. | +/- | 16. |
| -10000. | +/- | -NaN |
| 4.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.27 | +/- | 0. |
| 1.27 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.86 | +/- | 0. |
| 0.77 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.63 |
| -9999.00 |
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN,SN_WARN
280-33-O
| 5638. | +/- | 29. |
| -10000. | +/- | -NaN |
| 1.81 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.78 | +/- | 0. |
| 4.78 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.36 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.39 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.44 |
| -9999.00 |
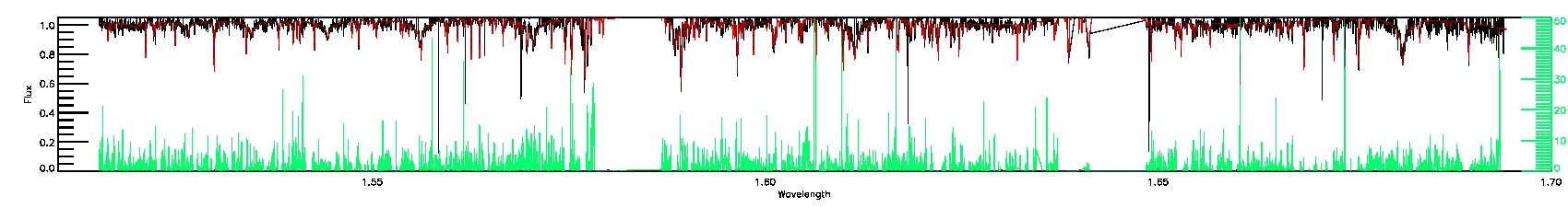
STAR_BAD,CHI2_BAD
STAR_WARN,CHI2_WARN
280-33-O
| 3000. | +/- | 0. |
| -10000. | +/- | -NaN |
| 0.70 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.56 | +/- | 0. |
| 3.56 | +/- | -NaN |
| -1.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.71 | +/- | 0. |
| 0.76 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.47 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -0.98 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5516. | +/- | 31. |
| -10000. | +/- | -NaN |
| 1.76 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.79 | +/- | 0. |
| 4.79 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.42 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.24 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.99 |
| -9999.00 |

STAR_BAD
STAR_WARN,CHI2_WARN
280-33-O
| 3599. | +/- | 2. |
| -10000. | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.01 | +/- | 0. |
| 3.01 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.26 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.19 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.09 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_WARN,SN_WARN
280-33-O
| 5785. | +/- | 22. |
| 5739. | +/- | 165. |
| 1.84 | +/- | 0. |
| 1.66 | +/- | 0. |
| 4.29 | +/- | 0. |
| 4.29 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -0.35 | +/- | 0. |
| -0.16 | +/- | 0. |
| -0.16 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.07 | +/- | 0. |
| 3.64 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.17 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.32 |
| -9999.00 |

280-33-O
| 6314. | +/- | 18. |
| 6204. | +/- | 139. |
| 4.37 | +/- | 0. |
| 4.29 | +/- | 0. |
| 0.72 | +/- | 0. |
| 0.72 | +/- | -NaN |
| -0.27 | +/- | 0. |
| -0.27 | +/- | 0. |
| -0.11 | +/- | 0. |
| -0.11 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.07 | +/- | 0. |
| 10.55 | +/- | 0. |
| 1.02 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.60 |
| -9999.00 |
280-33-O
| 6103. | +/- | 15. |
| 6008. | +/- | 127. |
| 4.56 | +/- | 0. |
| 4.42 | +/- | 0. |
| 0.50 | +/- | 0. |
| 0.50 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.00 | +/- | 0. |
| 4.06 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.12 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,COLORTE_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5594. | +/- | 64. |
| -10000. | +/- | -NaN |
| 1.72 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.37 | +/- | 0. |
| 4.37 | +/- | -NaN |
| -0.84 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.26 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.83 | +/- | 0. |
| 0.68 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.22 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.83 |
| -9999.00 |
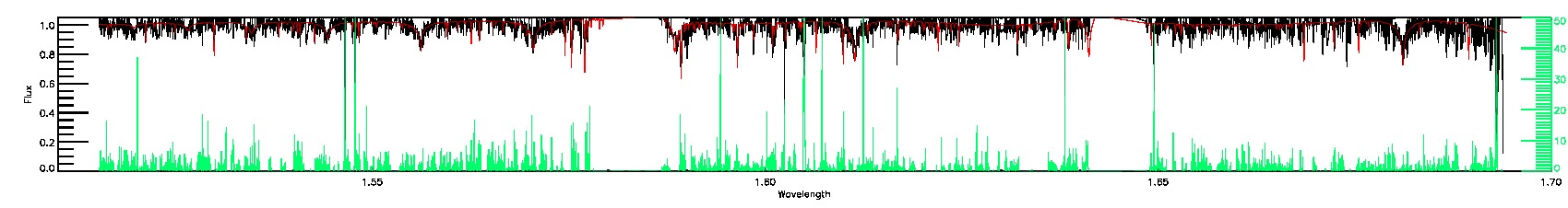
SUSPECT_BROAD_LINES
280-33-O
| 6026. | +/- | 19. |
| 5955. | +/- | 130. |
| 4.05 | +/- | 0. |
| 4.08 | +/- | 0. |
| 0.66 | +/- | 0. |
| 0.66 | +/- | -NaN |
| -0.42 | +/- | 0. |
| -0.42 | +/- | 0. |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 26.69 | +/- | 0. |
| 1.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.70 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5712. | +/- | 47. |
| -10000. | +/- | -NaN |
| 2.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.46 | +/- | 0. |
| 4.46 | +/- | -NaN |
| -0.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.67 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.19 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.24 |
| -9999.00 |

280-33-O
| 3531. | +/- | 1. |
| 3641. | +/- | 75. |
| 0.12 | +/- | 0. |
| 0.04 | +/- | 0. |
| 2.56 | +/- | 0. |
| 2.56 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -0.50 | +/- | 0. |
| -0.24 | +/- | 0. |
| -0.24 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.05 | +/- | 0. |
| 3.96 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.25 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -1.43 |
| -9999.00 |
STAR_BAD,CHI2_BAD
STAR_WARN,CHI2_WARN
280-33-O
| 3270. | +/- | 1. |
| -10000. | +/- | -NaN |
| -0.24 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.46 | +/- | 0. |
| 3.46 | +/- | -NaN |
| -0.62 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.34 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.26 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.56 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5420. | +/- | 26. |
| -10000. | +/- | -NaN |
| 1.66 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.41 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.75 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -2.49 |
| -9999.00 |

STAR_BAD
STAR_WARN,SN_WARN
280-33-O
| 5470. | +/- | 35. |
| -10000. | +/- | -NaN |
| 1.85 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.57 | +/- | 0. |
| 4.57 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.24 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.41 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.68 |
| -9999.00 |
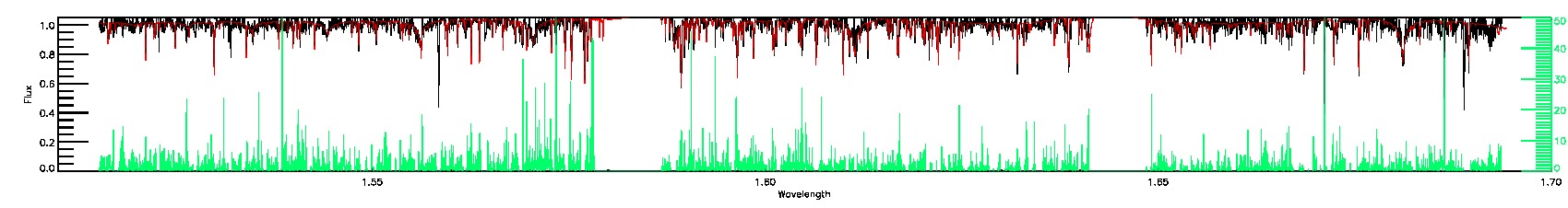
SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,SN_WARN
280-33-O
| 6831. | +/- | 17. |
| -10000. | +/- | -NaN |
| 2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.32 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 14.28 | +/- | 0. |
| 1.15 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.27 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.31 |
| -9999.00 |
280-33-O
| 3480. | +/- | 1. |
| 3583. | +/- | 61. |
| -0.17 | +/- | 0. |
| -0.27 | +/- | 0. |
| 2.83 | +/- | 0. |
| 2.83 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -0.35 | +/- | 0. |
| -0.23 | +/- | 0. |
| -0.23 | +/- | -NaN |
| 0.50 | +/- | 0. |
| 0.50 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.10 | +/- | 0. |
| 3.64 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.26 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| -0.75 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,COLORTE_BAD,ROTATION_BAD
STAR_WARN,COLORTE_WARN,ROTATION_WARN
280-33-O
| 6000. | +/- | 6. |
| -10000. | +/- | -NaN |
| 0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.24 | +/- | 0. |
| 1.24 | +/- | -NaN |
| -1.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.32 | +/- | 0. |
| 0.73 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
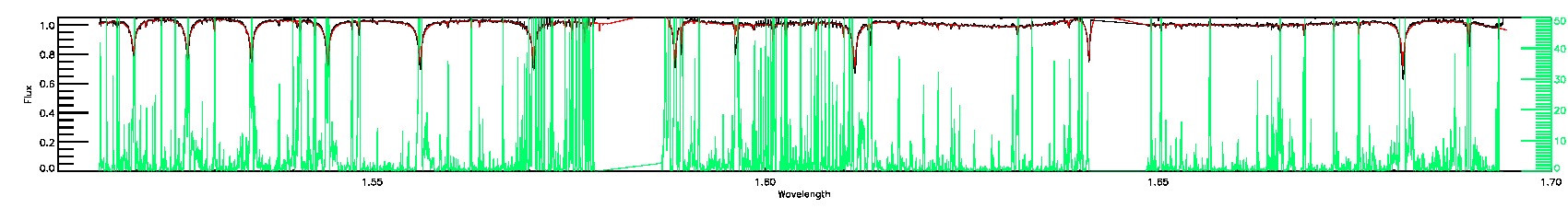
SUSPECT_BROAD_LINES
STAR_WARN,ROTATION_WARN
280-33-O
| 3839. | +/- | 1. |
| 3947. | +/- | 68. |
| 0.24 | +/- | 0. |
| 0.15 | +/- | 0. |
| 3.54 | +/- | 0. |
| 3.54 | +/- | -NaN |
| -0.45 | +/- | 0. |
| -0.45 | +/- | 0. |
| -0.18 | +/- | 0. |
| -0.18 | +/- | -NaN |
| 0.58 | +/- | 0. |
| 0.58 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.06 | +/- | 0. |
| 3.86 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.21 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -1.13 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,COLORTE_WARN,ROTATION_WARN
280-33-O
| 3773. | +/- | 1. |
| -10000. | +/- | -NaN |
| -0.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.46 | +/- | 0. |
| 4.46 | +/- | -NaN |
| -0.59 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.60 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.17 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.31 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.01 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
280-33-O
| 3336. | +/- | 1. |
| 3448. | +/- | 59. |
| 0.34 | +/- | 0. |
| 0.25 | +/- | 0. |
| 3.30 | +/- | 0. |
| 3.30 | +/- | -NaN |
| -0.53 | +/- | 0. |
| -0.53 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.04 | +/- | 0. |
| 4.03 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -0.89 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5342. | +/- | 36. |
| -10000. | +/- | -NaN |
| 1.70 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.55 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.07 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.19 |
| -9999.00 |
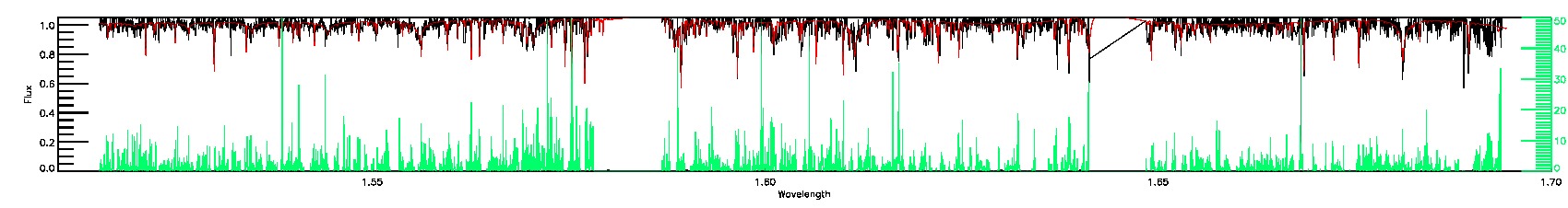
SUSPECT_BROAD_LINES
STAR_BAD,COLORTE_BAD,ROTATION_BAD
STAR_WARN,COLORTE_WARN,ROTATION_WARN
280-33-O
| 5742. | +/- | 5. |
| -10000. | +/- | -NaN |
| 0.87 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.90 | +/- | 0. |
| 0.90 | +/- | -NaN |
| -2.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.61 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.38 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -2.47 |
| -9999.00 |

BRIGHT_NEIGHBOR
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
280-33-O
| 3566. | +/- | 11. |
| -10000. | +/- | -NaN |
| -0.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.80 | +/- | 0. |
| 1.80 | +/- | -NaN |
| -0.86 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.89 | +/- | 0. |
| 0.69 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -1.59 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5642. | +/- | 40. |
| -10000. | +/- | -NaN |
| 1.65 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.79 | +/- | 0. |
| 4.79 | +/- | -NaN |
| -0.45 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.85 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.21 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| 0.37 |
| -9999.00 |

LOW_SNR
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
280-33-O
| 3602. | +/- | 15. |
| -10000. | +/- | -NaN |
| 1.70 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.78 | +/- | 0. |
| 1.78 | +/- | -NaN |
| 0.22 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.60 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.29 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| 0.21 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
280-33-O
| 5898. | +/- | 20. |
| 5839. | +/- | 124. |
| 4.19 | +/- | 0. |
| 4.22 | +/- | 0. |
| 0.90 | +/- | 0. |
| 0.90 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -0.36 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.07 | +/- | 0. |
| 3.65 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.19 |
| -9999.00 |

STAR_BAD,CHI2_BAD
STAR_WARN,CHI2_WARN
280-33-O
| 3018. | +/- | 1. |
| -10000. | +/- | -NaN |
| 1.87 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.33 | +/- | 0. |
| 4.33 | +/- | -NaN |
| -0.46 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.87 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.10 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5524. | +/- | 45. |
| -10000. | +/- | -NaN |
| 2.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.78 | +/- | 0. |
| 4.78 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.74 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.43 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN,ROTATION_WARN
280-33-O
| 5020. | +/- | 9. |
| -10000. | +/- | -NaN |
| 1.82 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.67 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| -2.36 |
| -9999.00 |
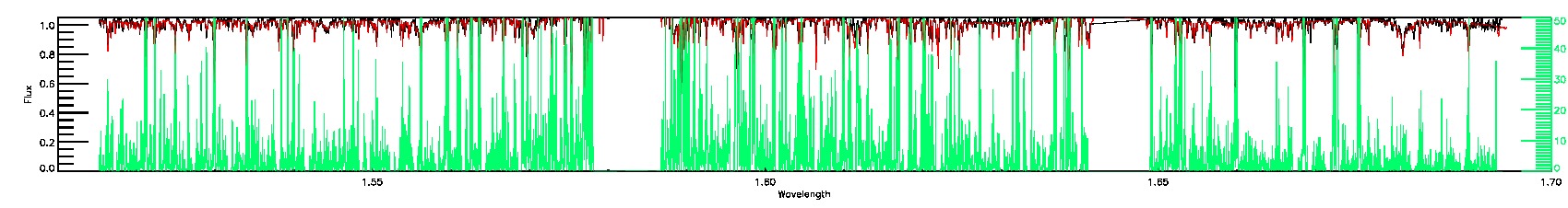
280-33-O
| 5247. | +/- | 8. |
| 5242. | +/- | 96. |
| 3.98 | +/- | 0. |
| 3.91 | +/- | 0. |
| 0.87 | +/- | 0. |
| 0.87 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.17 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.69 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.09 |
| -9999.00 |
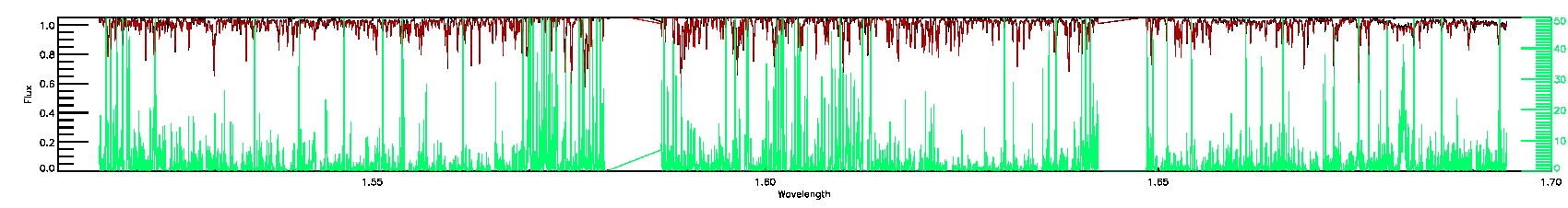
STAR_BAD
STAR_WARN,CHI2_WARN
280-33-O
| 3402. | +/- | 1. |
| -10000. | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.16 | +/- | 0. |
| 3.16 | +/- | -NaN |
| -0.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.69 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.06 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5576. | +/- | 38. |
| -10000. | +/- | -NaN |
| 2.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.65 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.98 |
| -9999.00 |

VERY_BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5770. | +/- | 29. |
| -10000. | +/- | -NaN |
| 1.65 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.79 | +/- | 0. |
| 4.79 | +/- | -NaN |
| -0.58 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.16 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.50 |
| -9999.00 |

VERY_BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5427. | +/- | 34. |
| -10000. | +/- | -NaN |
| 1.71 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.42 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.78 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.18 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| 0.61 |
| -9999.00 |

280-33-O
| 5079. | +/- | 9. |
| 5130. | +/- | 101. |
| 2.83 | +/- | 0. |
| 2.47 | +/- | 0. |
| 1.40 | +/- | 0. |
| 1.40 | +/- | -NaN |
| -0.70 | +/- | 0. |
| -0.70 | +/- | 0. |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.14 | +/- | 0. |
| 4.45 | +/- | 0. |
| 0.65 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| -1.14 |
| -9999.00 |
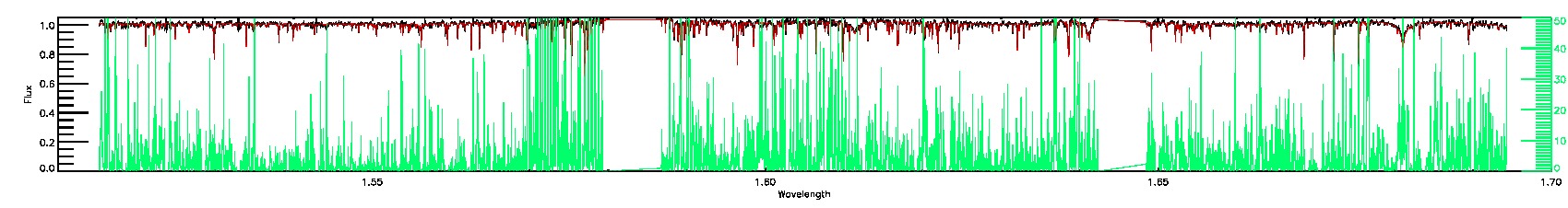
280-33-O
| 3688. | +/- | 1. |
| 3795. | +/- | 65. |
| 0.18 | +/- | 0. |
| 0.09 | +/- | 0. |
| 2.83 | +/- | 0. |
| 2.83 | +/- | -NaN |
| -0.42 | +/- | 0. |
| -0.42 | +/- | 0. |
| -0.21 | +/- | 0. |
| -0.21 | +/- | -NaN |
| 0.44 | +/- | 0. |
| 0.44 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.05 | +/- | 0. |
| 3.78 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.22 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -1.36 |
| -9999.00 |
VERY_BRIGHT_NEIGHBOR
STAR_WARN,SN_WARN
280-33-O
| 5712. | +/- | 25. |
| 5676. | +/- | 163. |
| 1.86 | +/- | 0. |
| 1.68 | +/- | 0. |
| 4.36 | +/- | 0. |
| 4.36 | +/- | -NaN |
| -0.38 | +/- | 0. |
| -0.38 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 3.69 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -1.98 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.74 |
| -9999.00 |

STAR_WARN,SN_WARN
280-33-O
| 5676. | +/- | 27. |
| 5636. | +/- | 161. |
| 1.78 | +/- | 0. |
| 1.58 | +/- | 0. |
| 4.17 | +/- | 0. |
| 4.17 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -0.20 | +/- | 0. |
| -0.27 | +/- | 0. |
| -0.27 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.06 | +/- | 0. |
| 3.32 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.23 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| 0.66 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN
280-33-O
| 3750. | +/- | 1. |
| -10000. | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.69 | +/- | 0. |
| 3.69 | +/- | -NaN |
| -0.53 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.54 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.04 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.30 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.53 |
| -9999.00 |
280-33-O
| 4710. | +/- | 4. |
| 4767. | +/- | 82. |
| 3.18 | +/- | 0. |
| 2.98 | +/- | 0. |
| 1.09 | +/- | 0. |
| 1.09 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.70 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.15 |
| -9999.00 |
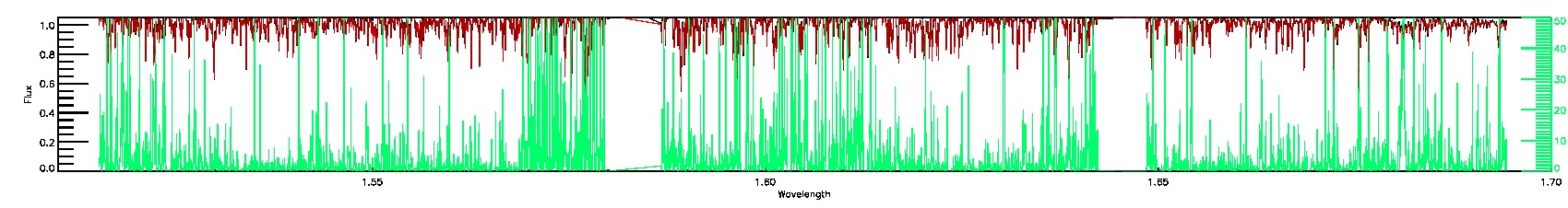
BAD_PIXELS
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN
280-33-O
| 5720. | +/- | 14. |
| -10000. | +/- | -NaN |
| 4.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.95 | +/- | 0. |
| 0.95 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.41 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.08 |
| -9999.00 |
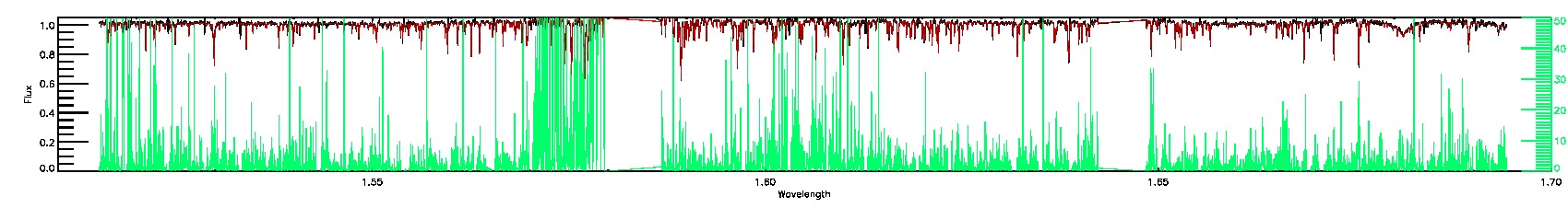
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN,ROTATION_WARN
280-33-O
| 5014. | +/- | 12. |
| -10000. | +/- | -NaN |
| 1.54 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.88 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
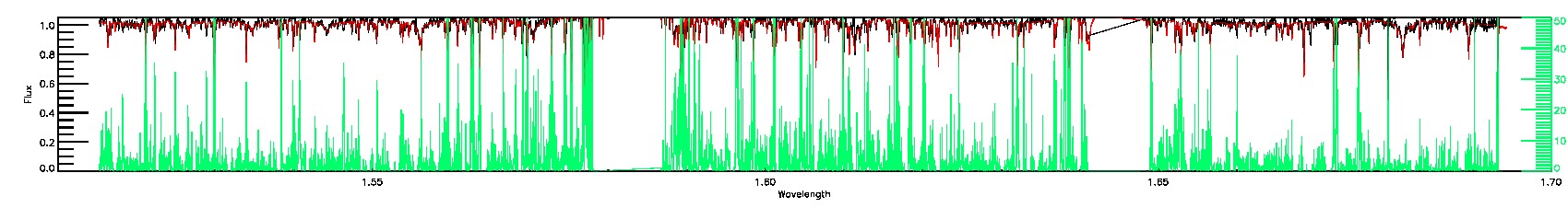
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN
280-33-O
| 7097. | +/- | 5. |
| -10000. | +/- | -NaN |
| 2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| -1.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -9999.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -9999.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.11 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN,SN_WARN
280-33-O
| 5422. | +/- | 26. |
| -10000. | +/- | -NaN |
| 1.76 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.27 | +/- | 0. |
| 4.27 | +/- | -NaN |
| -0.39 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.72 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.51 |
| -9999.00 |

280-33-O
| 3739. | +/- | 1. |
| 3848. | +/- | 77. |
| 0.05 | +/- | 0. |
| -0.03 | +/- | 0. |
| 3.19 | +/- | 0. |
| 3.19 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -0.50 | +/- | 0. |
| -0.24 | +/- | 0. |
| -0.24 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.07 | +/- | 0. |
| 3.96 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.26 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -1.16 |
| -9999.00 |
280-33-O
| 3756. | +/- | 1. |
| 3858. | +/- | 67. |
| 0.00 | +/- | 0. |
| -0.11 | +/- | 0. |
| 2.90 | +/- | 0. |
| 2.90 | +/- | -NaN |
| -0.30 | +/- | 0. |
| -0.30 | +/- | 0. |
| -0.44 | +/- | 0. |
| -0.44 | +/- | -NaN |
| 0.67 | +/- | 0. |
| 0.67 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.06 | +/- | 0. |
| 3.53 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.46 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -1.28 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
280-33-O
| 5806. | +/- | 11. |
| 5744. | +/- | 116. |
| 4.35 | +/- | 0. |
| 4.26 | +/- | 0. |
| 1.20 | +/- | 0. |
| 1.20 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.14 | +/- | 0. |
| -0.14 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.67 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -0.02 |
| -9999.00 |

280-33-O
| 3755. | +/- | 1. |
| 3864. | +/- | 67. |
| 0.23 | +/- | 0. |
| 0.14 | +/- | 0. |
| 2.61 | +/- | 0. |
| 2.61 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -0.49 | +/- | 0. |
| -0.33 | +/- | 0. |
| -0.33 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.06 | +/- | 0. |
| 3.93 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.35 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -1.44 |
| -9999.00 |
280-33-O
| 4814. | +/- | 4. |
| 4875. | +/- | 88. |
| 2.72 | +/- | 0. |
| 2.40 | +/- | 0. |
| 1.57 | +/- | 0. |
| 1.57 | +/- | -NaN |
| -0.21 | +/- | 0. |
| -0.21 | +/- | 0. |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.34 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -1.11 |
| -9999.00 |

280-33-O
| 3644. | +/- | 1. |
| 3756. | +/- | 65. |
| -0.24 | +/- | 0. |
| -0.32 | +/- | 0. |
| 3.39 | +/- | 0. |
| 3.39 | +/- | -NaN |
| -0.55 | +/- | 0. |
| -0.55 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.22 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.06 | +/- | 0. |
| 4.07 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.24 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -1.54 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5389. | +/- | 34. |
| -10000. | +/- | -NaN |
| 1.92 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.79 | +/- | 0. |
| 4.79 | +/- | -NaN |
| -0.51 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.26 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.99 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.38 |
| -9999.00 |

SUSPECT_RV_COMBINATION
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN
280-33-O
| 7266. | +/- | 6. |
| -10000. | +/- | -NaN |
| 2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.69 | +/- | 1. |
| 0.69 | +/- | -NaN |
| -2.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.55 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 13.60 | +/- | 0. |
| 1.13 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.43 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -2.19 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD
STAR_WARN,SN_WARN
280-33-O
| 5328. | +/- | 28. |
| -10000. | +/- | -NaN |
| 1.90 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.42 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.78 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.75 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD
280-33-O
| 4740. | +/- | 10. |
| -10000. | +/- | -NaN |
| 3.62 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.17 | +/- | 0. |
| 1.17 | +/- | -NaN |
| -0.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.98 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.23 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.28 |
| -9999.00 |
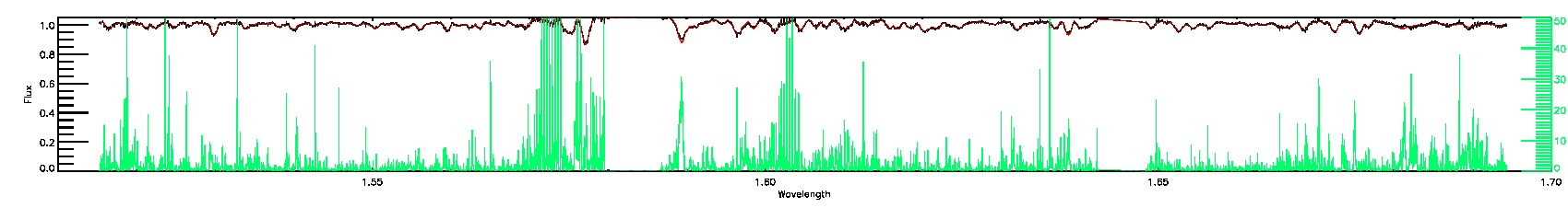
SUSPECT_BROAD_LINES
STAR_WARN,ROTATION_WARN
280-33-O
| 3672. | +/- | 1. |
| 3785. | +/- | 66. |
| -0.29 | +/- | 0. |
| -0.37 | +/- | 0. |
| 4.21 | +/- | 0. |
| 4.21 | +/- | -NaN |
| -0.57 | +/- | 0. |
| -0.57 | +/- | 0. |
| -0.25 | +/- | 0. |
| -0.25 | +/- | -NaN |
| 0.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.08 | +/- | 0. |
| 4.13 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.27 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.98 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN,SN_WARN
280-33-O
| 5197. | +/- | 14. |
| -10000. | +/- | -NaN |
| 1.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.26 | +/- | 0. |
| 4.26 | +/- | -NaN |
| -0.39 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.71 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.37 |
| -9999.00 |

280-33-O
| 4670. | +/- | 3. |
| 4730. | +/- | 81. |
| 4.56 | +/- | 0. |
| 4.59 | +/- | 0. |
| 0.84 | +/- | 0. |
| 0.84 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.04 | +/- | 0. |
| 1.85 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.18 |
| -9999.00 |

STAR_BAD
280-33-O
| 3249. | +/- | 1. |
| -10000. | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.95 | +/- | 0. |
| 2.95 | +/- | -NaN |
| -0.46 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.88 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.64 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN
280-33-O
| 3610. | +/- | 1. |
| -10000. | +/- | -NaN |
| 0.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.49 | +/- | 0. |
| 4.49 | +/- | -NaN |
| -0.61 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.24 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -1.02 |
| -9999.00 |
280-33-O
| 3803. | +/- | 1. |
| 3914. | +/- | 68. |
| -0.01 | +/- | 0. |
| -0.09 | +/- | 0. |
| 3.79 | +/- | 0. |
| 3.79 | +/- | -NaN |
| -0.54 | +/- | 0. |
| -0.54 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.22 | +/- | -NaN |
| 0.40 | +/- | 0. |
| 0.40 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.09 | +/- | 0. |
| 4.05 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.24 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.43 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN
280-33-O
| 3628. | +/- | 1. |
| -10000. | +/- | -NaN |
| 0.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.26 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.66 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -1.18 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,COLORTE_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5306. | +/- | 28. |
| -10000. | +/- | -NaN |
| 1.61 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.79 | +/- | 0. |
| 4.79 | +/- | -NaN |
| -0.51 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.00 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.61 |
| -9999.00 |
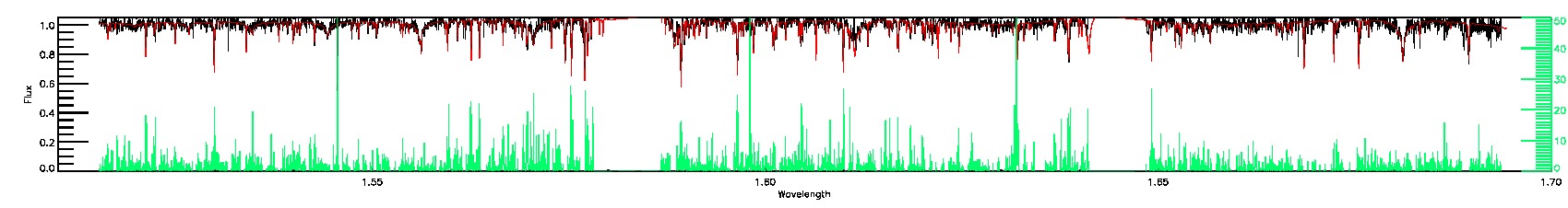
280-33-O
| 3801. | +/- | 1. |
| 3909. | +/- | 68. |
| 0.21 | +/- | 0. |
| 0.12 | +/- | 0. |
| 3.41 | +/- | 0. |
| 3.41 | +/- | -NaN |
| -0.47 | +/- | 0. |
| -0.47 | +/- | 0. |
| -0.23 | +/- | 0. |
| -0.23 | +/- | -NaN |
| 0.43 | +/- | 0. |
| 0.43 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.08 | +/- | 0. |
| 3.90 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.25 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.12 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,COLORTE_WARN,ROTATION_WARN
280-33-O
| 3720. | +/- | 1. |
| -10000. | +/- | -NaN |
| 0.67 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.54 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.06 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -0.94 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD
STAR_WARN,SN_WARN
280-33-O
| 5030. | +/- | 16. |
| -10000. | +/- | -NaN |
| 1.30 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.52 | +/- | 0. |
| 4.52 | +/- | -NaN |
| -0.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.31 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.59 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.33 |
| -9999.00 |
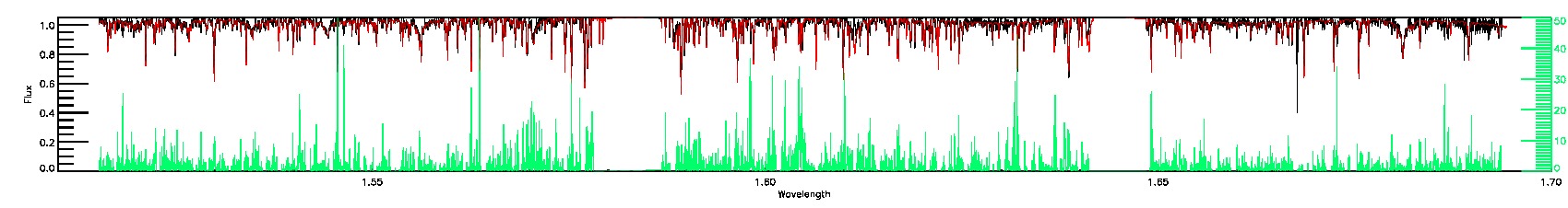
SUSPECT_BROAD_LINES
280-33-O
| 5620. | +/- | 13. |
| 5578. | +/- | 109. |
| 4.61 | +/- | 0. |
| 4.54 | +/- | 0. |
| 1.14 | +/- | 0. |
| 1.14 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.02 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| -0.21 | +/- | 0. |
| -0.21 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.06 | +/- | 0. |
| 20.43 | +/- | 0. |
| 1.31 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.07 |
| -9999.00 |

280-33-O
| 3864. | +/- | 1. |
| 3973. | +/- | 69. |
| 0.28 | +/- | 0. |
| 0.19 | +/- | 0. |
| 3.42 | +/- | 0. |
| 3.42 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -0.50 | +/- | 0. |
| -0.25 | +/- | 0. |
| -0.25 | +/- | -NaN |
| 0.37 | +/- | 0. |
| 0.37 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.08 | +/- | 0. |
| 3.95 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.26 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -1.21 |
| -9999.00 |
280-33-O
| 5674. | +/- | 11. |
| 5627. | +/- | 112. |
| 3.98 | +/- | 0. |
| 3.91 | +/- | 0. |
| 1.35 | +/- | 0. |
| 1.35 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.99 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.11 |
| -9999.00 |
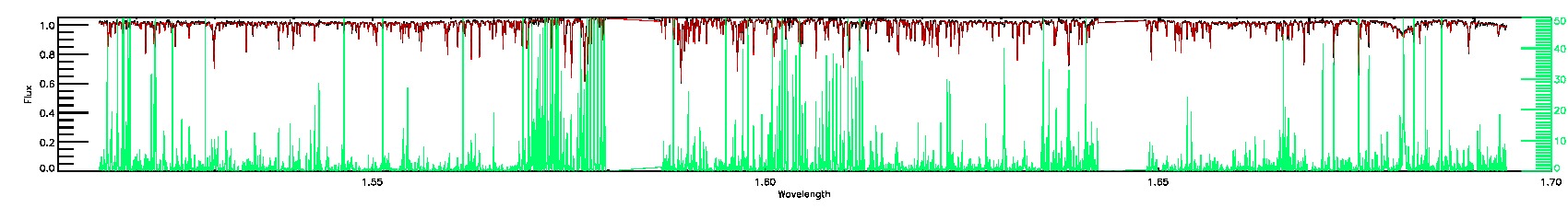
SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN
280-33-O
| 3692. | +/- | 1. |
| -10000. | +/- | -NaN |
| 0.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.57 | +/- | 0. |
| 4.57 | +/- | -NaN |
| -0.61 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.41 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.22 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -1.03 |
| -9999.00 |
280-33-O
| 3768. | +/- | 1. |
| 3877. | +/- | 67. |
| 0.17 | +/- | 0. |
| 0.08 | +/- | 0. |
| 3.01 | +/- | 0. |
| 3.01 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -0.49 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.22 | +/- | -NaN |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.03 | +/- | 0. |
| 3.94 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.24 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.38 |
| -9999.00 |
280-33-O
| 4877. | +/- | 5. |
| 4928. | +/- | 89. |
| 3.07 | +/- | 0. |
| 2.90 | +/- | 0. |
| 1.20 | +/- | 0. |
| 1.20 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.01 | +/- | 0. |
| 3.22 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.20 |
| -9999.00 |
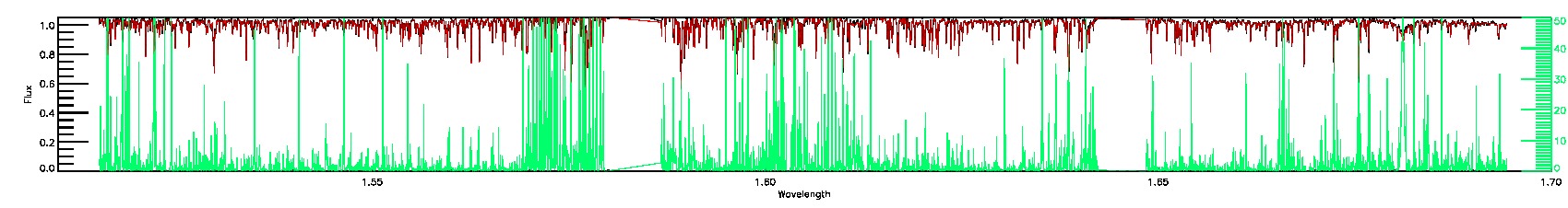
280-33-O
| 3813. | +/- | 1. |
| 3918. | +/- | 67. |
| 0.33 | +/- | 0. |
| 0.23 | +/- | 0. |
| 3.42 | +/- | 0. |
| 3.42 | +/- | -NaN |
| -0.39 | +/- | 0. |
| -0.39 | +/- | 0. |
| -0.24 | +/- | 0. |
| -0.24 | +/- | -NaN |
| 0.53 | +/- | 0. |
| 0.53 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -0.06 | +/- | 0. |
| 3.72 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.26 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -0.30 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_WARN,SN_WARN
280-33-O
| 5097. | +/- | 13. |
| 5128. | +/- | 131. |
| 0.98 | +/- | 0. |
| 0.87 | +/- | 0. |
| 4.23 | +/- | 0. |
| 4.23 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -0.28 | +/- | 0. |
| -0.37 | +/- | 0. |
| -0.37 | +/- | -NaN |
| 0.44 | +/- | 0. |
| 0.44 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.04 | +/- | 0. |
| 3.49 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.39 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| 0.67 |
| -9999.00 |
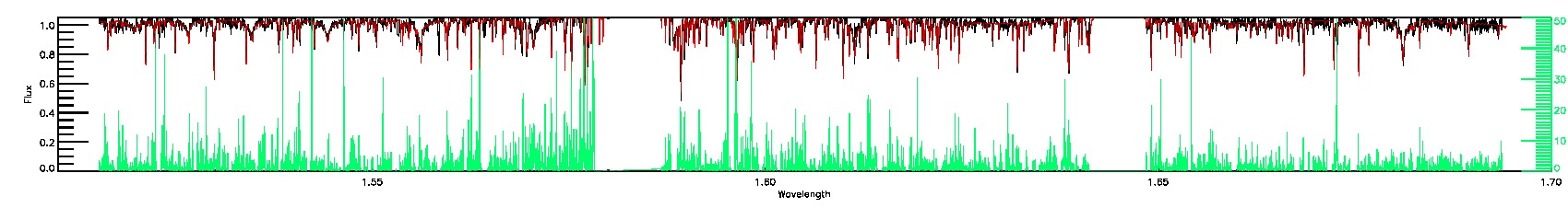
280-33-O
| 7102. | +/- | 13. |
| 6894. | +/- | 167. |
| 4.64 | +/- | 0. |
| 4.24 | +/- | 0. |
| 2.19 | +/- | 0. |
| 2.19 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | 0. |
| -0.15 | +/- | 0. |
| -0.15 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.04 | +/- | 0. |
| 12.48 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.05 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN
280-33-O
| 5256. | +/- | 8. |
| -10000. | +/- | -NaN |
| 0.72 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.79 | +/- | 0. |
| 4.79 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.94 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.06 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,COLORTE_BAD,ROTATION_BAD
STAR_WARN,COLORTE_WARN,ROTATION_WARN
280-33-O
| 5211. | +/- | 5. |
| -10000. | +/- | -NaN |
| 0.57 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.60 | +/- | 0. |
| 0.60 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.77 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.33 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -2.14 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.35 |
| -9999.00 |

STAR_WARN,SN_WARN
280-33-O
| 5684. | +/- | 23. |
| 5650. | +/- | 161. |
| 1.77 | +/- | 0. |
| 1.60 | +/- | 0. |
| 4.14 | +/- | 0. |
| 4.14 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -0.36 | +/- | 0. |
| -0.20 | +/- | 0. |
| -0.20 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.65 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.18 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.66 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN
280-33-O
| 7037. | +/- | 4. |
| -10000. | +/- | -NaN |
| 2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.33 | +/- | 19. |
| 0.33 | +/- | -NaN |
| -2.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.21 | +/- | 0. |
| 0.72 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -2.12 |
| -9999.00 |
280-33-O
| 3378. | +/- | 1. |
| 3486. | +/- | 60. |
| 0.35 | +/- | 0. |
| 0.26 | +/- | 0. |
| 3.34 | +/- | 0. |
| 3.34 | +/- | -NaN |
| -0.47 | +/- | 0. |
| -0.47 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.04 | +/- | 0. |
| 3.90 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.87 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD
STAR_WARN,SN_WARN
280-33-O
| 6853. | +/- | 19. |
| -10000. | +/- | -NaN |
| 2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.05 | +/- | 0. |
| 4.05 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.60 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.23 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.14 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,COLORTE_WARN,ROTATION_WARN
280-33-O
| 3726. | +/- | 1. |
| -10000. | +/- | -NaN |
| 0.53 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.79 | +/- | 0. |
| 4.79 | +/- | -NaN |
| -0.56 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.11 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -0.97 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5550. | +/- | 30. |
| -10000. | +/- | -NaN |
| 1.82 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.57 | +/- | 0. |
| 4.57 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.75 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.17 |
| -9999.00 |
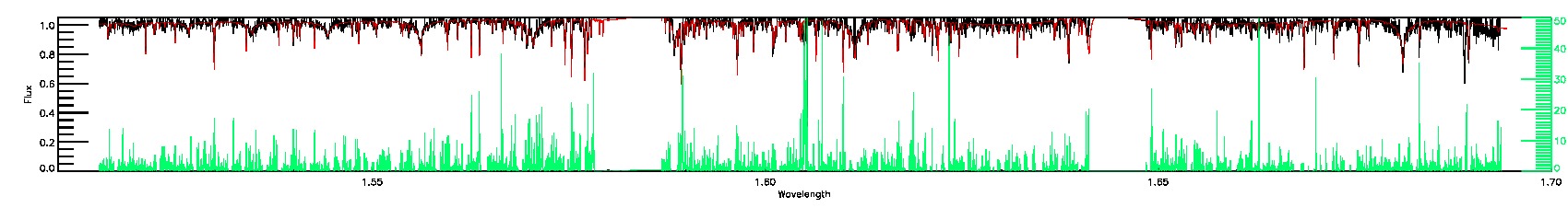
280-33-O
| 4230. | +/- | 2. |
| 4340. | +/- | 77. |
| 2.07 | +/- | 0. |
| 1.90 | +/- | 0. |
| 1.49 | +/- | 0. |
| 1.49 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -0.49 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.17 | +/- | 0. |
| 3.95 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -0.17 |
| -9999.00 |

280-33-O
| 3585. | +/- | 1. |
| 3692. | +/- | 67. |
| 0.12 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.64 | +/- | 0. |
| 2.64 | +/- | -NaN |
| -0.45 | +/- | 0. |
| -0.45 | +/- | 0. |
| -0.25 | +/- | 0. |
| -0.25 | +/- | -NaN |
| 0.37 | +/- | 0. |
| 0.37 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.08 | +/- | 0. |
| 3.84 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.27 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -1.16 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5971. | +/- | 32. |
| -10000. | +/- | -NaN |
| 1.69 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.56 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.10 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.38 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_WARN,ROTATION_WARN
280-33-O
| 3962. | +/- | 1. |
| 4066. | +/- | 70. |
| 0.19 | +/- | 0. |
| 0.09 | +/- | 0. |
| 3.71 | +/- | 0. |
| 3.71 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -0.36 | +/- | 0. |
| -0.47 | +/- | 0. |
| -0.47 | +/- | -NaN |
| 0.72 | +/- | 0. |
| 0.72 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.13 | +/- | 0. |
| 3.66 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.49 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.42 |
| -9999.00 |
STAR_BAD
STAR_WARN,COLORTE_WARN
280-33-O
| 3297. | +/- | 1. |
| -10000. | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.98 | +/- | 0. |
| 2.98 | +/- | -NaN |
| -0.57 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.32 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.14 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.25 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| -1.04 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5870. | +/- | 27. |
| -10000. | +/- | -NaN |
| 1.75 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.55 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.07 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.56 |
| -9999.00 |

STAR_BAD
STAR_WARN,SN_WARN
280-33-O
| 5272. | +/- | 38. |
| -10000. | +/- | -NaN |
| 1.62 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.78 | +/- | 0. |
| 4.78 | +/- | -NaN |
| -0.54 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.06 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.67 |
| -9999.00 |
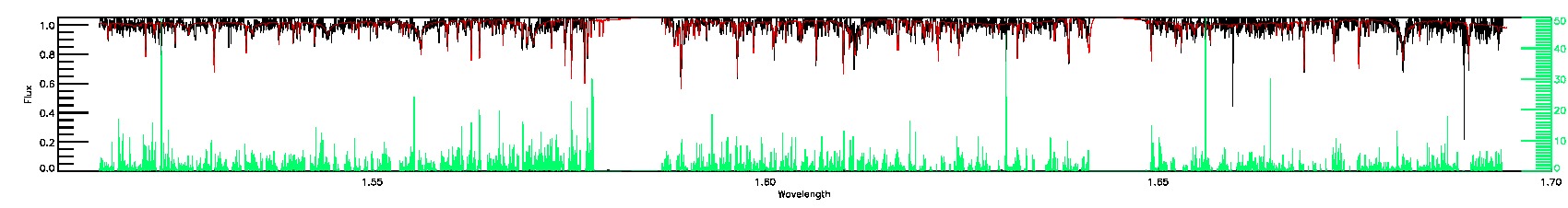
STAR_WARN,SN_WARN
280-33-O
| 3854. | +/- | 5. |
| 3967. | +/- | 99. |
| 0.19 | +/- | 0. |
| 0.12 | +/- | 0. |
| 3.33 | +/- | 0. |
| 3.33 | +/- | -NaN |
| -0.58 | +/- | 0. |
| -0.58 | +/- | 0. |
| -0.19 | +/- | 0. |
| -0.19 | +/- | -NaN |
| 0.37 | +/- | 0. |
| 0.37 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.07 | +/- | 0. |
| 4.14 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.20 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.18 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,COLORTE_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5072. | +/- | 19. |
| -10000. | +/- | -NaN |
| 1.73 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.79 | +/- | 0. |
| 4.79 | +/- | -NaN |
| -0.56 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.11 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -1.98 |
| -9999.00 |
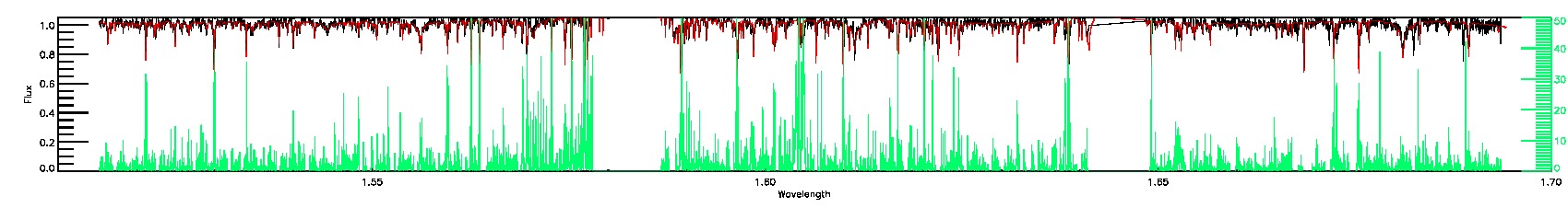
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,ROTATION_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5480. | +/- | 31. |
| -10000. | +/- | -NaN |
| 1.88 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.96 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.22 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.75 |
| -9999.00 |

STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN
280-33-O
| 5991. | +/- | 16. |
| -10000. | +/- | -NaN |
| 4.76 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.52 | +/- | 0. |
| 0.52 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.65 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.61 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_WARN,ROTATION_WARN
280-33-O
| 3781. | +/- | 1. |
| 3894. | +/- | 68. |
| -0.36 | +/- | 0. |
| -0.43 | +/- | 0. |
| 4.23 | +/- | 0. |
| 4.23 | +/- | -NaN |
| -0.57 | +/- | 0. |
| -0.57 | +/- | 0. |
| -0.24 | +/- | 0. |
| -0.24 | +/- | -NaN |
| 0.52 | +/- | 0. |
| 0.52 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.04 | +/- | 0. |
| 4.13 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.26 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.25 |
| -9999.00 |
280-33-O
| 3710. | +/- | 1. |
| 3821. | +/- | 66. |
| -0.34 | +/- | 0. |
| -0.42 | +/- | 0. |
| 3.74 | +/- | 0. |
| 3.74 | +/- | -NaN |
| -0.53 | +/- | 0. |
| -0.53 | +/- | 0. |
| -0.22 | +/- | 0. |
| -0.22 | +/- | -NaN |
| 0.47 | +/- | 0. |
| 0.47 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.05 | +/- | 0. |
| 4.04 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.24 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -1.13 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,COLORTE_WARN,ROTATION_WARN,SN_WARN
280-33-O
| 5572. | +/- | 59. |
| -10000. | +/- | -NaN |
| 1.85 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -0.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.92 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.43 |
| -9999.00 |

