STAR_WARN,SN_WARN
357-03-O
| 3618. | +/- | 3. |
| 3694. | +/- | 82. |
| 1.18 | +/- | 0. |
| 0.98 | +/- | 0. |
| 1.53 | +/- | 0. |
| 1.53 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.47 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.34 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
357-03-O
| 3331. | +/- | 1. |
| 3408. | +/- | 55. |
| 0.63 | +/- | 0. |
| 0.44 | +/- | 0. |
| 1.63 | +/- | 0. |
| 1.63 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.53 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.30 |
| -9999.00 |
357-03-O
| 3472. | +/- | 1. |
| 3540. | +/- | 65. |
| 1.27 | +/- | 0. |
| 1.05 | +/- | 0. |
| 1.55 | +/- | 0. |
| 1.55 | +/- | -NaN |
| 0.47 | +/- | 0. |
| 0.48 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.24 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.54 |
| -9999.00 |
357-03-O
| 3703. | +/- | 1. |
| 3795. | +/- | 71. |
| 1.00 | +/- | 0. |
| 0.86 | +/- | 0. |
| 1.72 | +/- | 0. |
| 1.72 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -0.09 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.06 | +/- | 0. |
| 3.12 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.01 |
| -9999.00 |
357-03-O
| 3862. | +/- | 2. |
| 3959. | +/- | 85. |
| 1.30 | +/- | 0. |
| 1.17 | +/- | 0. |
| 1.34 | +/- | 0. |
| 1.34 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -0.22 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.19 | +/- | 0. |
| 3.35 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.47 |
| -9999.00 |
357-03-O
| 3673. | +/- | 2. |
| 3740. | +/- | 76. |
| 1.53 | +/- | 0. |
| 1.27 | +/- | 0. |
| 1.30 | +/- | 0. |
| 1.30 | +/- | -NaN |
| 0.48 | +/- | 0. |
| 0.48 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.23 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.52 |
| -9999.00 |
STAR_WARN,SN_WARN
357-03-O
| 3458. | +/- | 5. |
| 3528. | +/- | 78. |
| 1.09 | +/- | 0. |
| 0.88 | +/- | 0. |
| 1.34 | +/- | 0. |
| 1.34 | +/- | -NaN |
| 0.44 | +/- | 0. |
| 0.44 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.41 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.04 | +/- | 0. |
| 2.29 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.47 |
| -9999.00 |
STAR_WARN,SN_WARN
357-03-O
| 4170. | +/- | 4. |
| 4245. | +/- | 95. |
| 2.34 | +/- | 0. |
| 2.07 | +/- | 0. |
| 1.13 | +/- | 0. |
| 1.13 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.47 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.38 |
| -9999.00 |

STAR_WARN,SN_WARN
357-03-O
| 3790. | +/- | 3. |
| 3861. | +/- | 83. |
| 1.80 | +/- | 0. |
| 1.52 | +/- | 0. |
| 1.02 | +/- | 0. |
| 1.02 | +/- | -NaN |
| 0.41 | +/- | 0. |
| 0.41 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.32 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.59 |
| -9999.00 |
357-03-O
| 3706. | +/- | 1. |
| 3790. | +/- | 75. |
| 1.37 | +/- | 0. |
| 1.19 | +/- | 0. |
| 1.59 | +/- | 0. |
| 1.59 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.20 | +/- | 0. |
| 2.81 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.20 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.02 |
| -9999.00 |
357-03-O
| 3939. | +/- | 3. |
| 4013. | +/- | 84. |
| 1.90 | +/- | 0. |
| 1.63 | +/- | 0. |
| 1.18 | +/- | 0. |
| 1.18 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.43 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 1.00 |
| -9999.00 |
357-03-O
| 3413. | +/- | 1. |
| 3505. | +/- | 69. |
| 0.47 | +/- | 0. |
| 0.33 | +/- | 0. |
| 1.79 | +/- | 0. |
| 1.79 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.08 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 3.10 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.05 |
| -9999.00 |
357-03-O
| 4602. | +/- | 9. |
| 4674. | +/- | 106. |
| 2.74 | +/- | 0. |
| 2.41 | +/- | 0. |
| 1.22 | +/- | 0. |
| 1.22 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.79 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
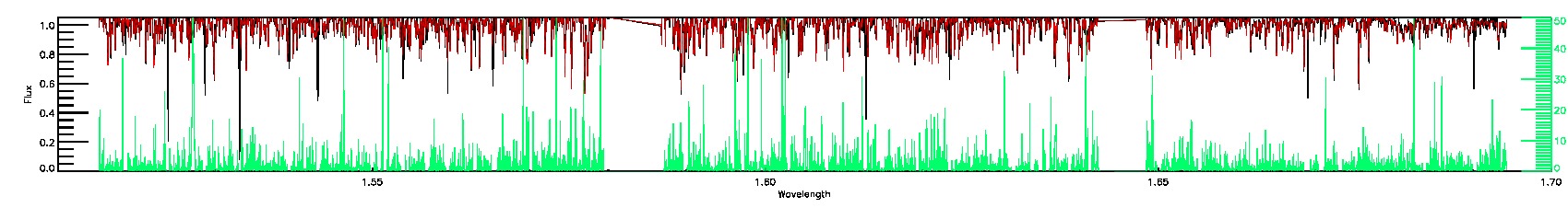
357-03-O
| 3248. | +/- | 1. |
| 3333. | +/- | 54. |
| 0.74 | +/- | 0. |
| 0.58 | +/- | 0. |
| 1.68 | +/- | 0. |
| 1.68 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.22 | +/- | 0. |
| 2.80 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.20 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.10 |
| -9999.00 |
357-03-O
| 3460. | +/- | 1. |
| 3536. | +/- | 64. |
| 0.88 | +/- | 0. |
| 0.68 | +/- | 0. |
| 1.44 | +/- | 0. |
| 1.44 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.47 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.36 |
| -9999.00 |
STAR_WARN,SN_WARN
357-03-O
| 3524. | +/- | 3. |
| 3640. | +/- | 86. |
| 0.68 | +/- | 0. |
| 0.61 | +/- | 0. |
| 2.22 | +/- | 0. |
| 2.22 | +/- | -NaN |
| -0.64 | +/- | 0. |
| -0.64 | +/- | 0. |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.27 | +/- | 0. |
| 4.31 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.91 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
357-03-O
| 4095. | +/- | 3. |
| 4186. | +/- | 88. |
| 2.09 | +/- | 0. |
| 1.86 | +/- | 0. |
| 0.70 | +/- | 0. |
| 0.70 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -0.06 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 3.06 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| -0.46 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
357-03-O
| 3213. | +/- | 1. |
| 3296. | +/- | 53. |
| 0.22 | +/- | 0. |
| 0.05 | +/- | 0. |
| 1.75 | +/- | 0. |
| 1.75 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.75 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.16 |
| -9999.00 |
357-03-O
| 3728. | +/- | 1. |
| 3836. | +/- | 72. |
| 0.89 | +/- | 0. |
| 0.80 | +/- | 0. |
| 2.02 | +/- | 0. |
| 2.02 | +/- | -NaN |
| -0.45 | +/- | 0. |
| -0.45 | +/- | 0. |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.24 | +/- | 0. |
| 3.86 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.84 |
| -9999.00 |
357-03-O
| 3943. | +/- | 2. |
| 4065. | +/- | 88. |
| 1.16 | +/- | 0. |
| 1.11 | +/- | 0. |
| 1.93 | +/- | 0. |
| 1.93 | +/- | -NaN |
| -0.79 | +/- | 0. |
| -0.79 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.28 | +/- | 0. |
| 4.70 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.46 |
| -9999.00 |
357-03-O
| 3457. | +/- | 1. |
| 3568. | +/- | 68. |
| 0.46 | +/- | 0. |
| 0.38 | +/- | 0. |
| 2.35 | +/- | 0. |
| 2.35 | +/- | -NaN |
| -0.53 | +/- | 0. |
| -0.53 | +/- | 0. |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.25 | +/- | 0. |
| 4.02 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -1.06 |
| -9999.00 |
357-03-O
| 3569. | +/- | 2. |
| 3645. | +/- | 75. |
| 1.13 | +/- | 0. |
| 0.94 | +/- | 0. |
| 1.30 | +/- | 0. |
| 1.30 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.29 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.50 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.32 |
| -9999.00 |
STAR_WARN,SN_WARN
357-03-O
| 3906. | +/- | 3. |
| 3976. | +/- | 85. |
| 1.96 | +/- | 0. |
| 1.67 | +/- | 0. |
| 1.09 | +/- | 0. |
| 1.09 | +/- | -NaN |
| 0.44 | +/- | 0. |
| 0.44 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.29 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.73 |
| -9999.00 |
357-03-O
| 3643. | +/- | 2. |
| 3731. | +/- | 75. |
| 1.17 | +/- | 0. |
| 1.02 | +/- | 0. |
| 1.68 | +/- | 0. |
| 1.68 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.20 | +/- | 0. |
| 2.92 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.74 |
| -9999.00 |
357-03-O
| 3753. | +/- | 2. |
| 3817. | +/- | 75. |
| 1.78 | +/- | 0. |
| 1.49 | +/- | 0. |
| 1.34 | +/- | 0. |
| 1.34 | +/- | -NaN |
| 0.57 | +/- | 0. |
| 0.57 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.37 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.12 | +/- | 0. |
| 0.33 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.87 |
| -9999.00 |
357-03-O
| 3655. | +/- | 1. |
| 3779. | +/- | 68. |
| 0.70 | +/- | 0. |
| 0.66 | +/- | 0. |
| 2.30 | +/- | 0. |
| 2.30 | +/- | -NaN |
| -0.84 | +/- | 0. |
| -0.84 | +/- | 0. |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.28 | +/- | 0. |
| 4.83 | +/- | 0. |
| 0.68 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -1.44 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,COLORTE_WARN,SN_WARN
357-03-O
| 3866. | +/- | 13. |
| -10000. | +/- | -NaN |
| 1.32 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| -0.58 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.14 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.93 |
| -9999.00 |
357-03-O
| 3900. | +/- | 2. |
| 3995. | +/- | 85. |
| 1.62 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.32 | +/- | 0. |
| 1.32 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.15 | +/- | 0. |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.21 | +/- | 0. |
| 3.23 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.20 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.10 |
| -9999.00 |
STAR_WARN,SN_WARN
357-03-O
| 3494. | +/- | 4. |
| 3568. | +/- | 79. |
| 1.07 | +/- | 0. |
| 0.87 | +/- | 0. |
| 1.42 | +/- | 0. |
| 1.42 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.42 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.28 |
| -9999.00 |
STAR_BAD
STAR_WARN,COLORTE_WARN
357-03-O
| 8302. | +/- | 26. |
| -10000. | +/- | -NaN |
| 3.59 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -0.59 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 93.95 | +/- | 0. |
| 1.97 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.32 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.89 |
| -9999.00 |
STAR_WARN,SN_WARN
357-03-O
| 3994. | +/- | 6. |
| 4071. | +/- | 94. |
| 1.89 | +/- | 0. |
| 1.62 | +/- | 0. |
| 1.35 | +/- | 0. |
| 1.35 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.54 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.30 |
| -9999.00 |
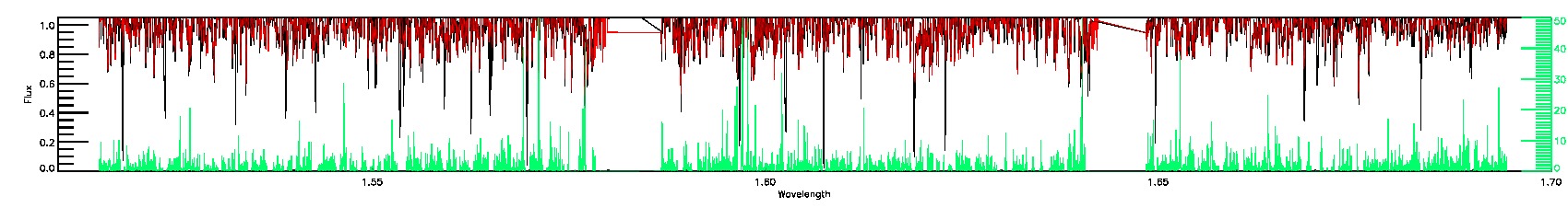
STAR_WARN,SN_WARN
357-03-O
| 4122. | +/- | 6. |
| 4234. | +/- | 105. |
| 1.57 | +/- | 0. |
| 1.44 | +/- | 0. |
| 1.44 | +/- | 0. |
| 1.44 | +/- | -NaN |
| -0.54 | +/- | 0. |
| -0.54 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.22 | +/- | 0. |
| 4.06 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 0.52 |
| -9999.00 |
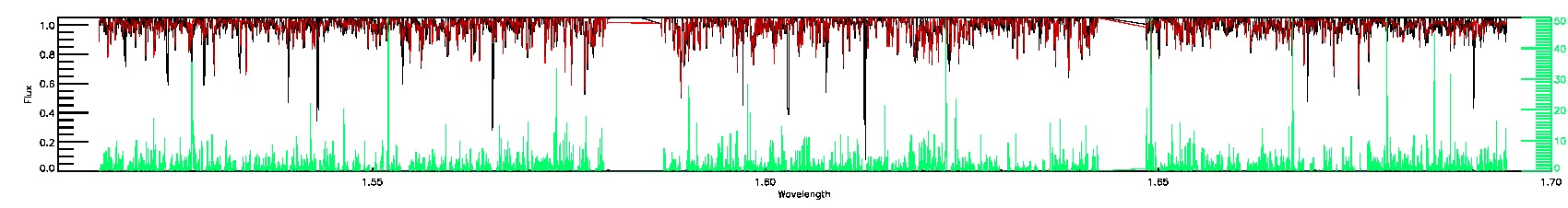
STAR_WARN,COLORTE_WARN
357-03-O
| 3192. | +/- | 1. |
| 3270. | +/- | 52. |
| 0.34 | +/- | 0. |
| 0.15 | +/- | 0. |
| 1.61 | +/- | 0. |
| 1.61 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.57 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.52 |
| -9999.00 |
STAR_WARN,SN_WARN
357-03-O
| 4038. | +/- | 3. |
| 4110. | +/- | 88. |
| 2.17 | +/- | 0. |
| 1.88 | +/- | 0. |
| 1.28 | +/- | 0. |
| 1.28 | +/- | -NaN |
| 0.40 | +/- | 0. |
| 0.40 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.35 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.90 |
| -9999.00 |

STAR_WARN,COLORTE_WARN,SN_WARN
357-03-O
| 3184. | +/- | 4. |
| 3264. | +/- | 74. |
| 0.65 | +/- | 0. |
| 0.47 | +/- | 0. |
| 1.53 | +/- | 0. |
| 1.53 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | 0. |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.21 | +/- | 0. |
| 2.66 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.22 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.95 |
| -9999.00 |
357-03-O
| 3802. | +/- | 2. |
| 3868. | +/- | 79. |
| 1.77 | +/- | 0. |
| 1.48 | +/- | 0. |
| 1.46 | +/- | 0. |
| 1.46 | +/- | -NaN |
| 0.50 | +/- | 0. |
| 0.50 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.20 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.54 |
| -9999.00 |
357-03-O
| 3378. | +/- | 1. |
| 3469. | +/- | 57. |
| 0.36 | +/- | 0. |
| 0.21 | +/- | 0. |
| 1.91 | +/- | 0. |
| 1.91 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.06 | +/- | 0. |
| 3.08 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
357-03-O
| 8574. | +/- | 33. |
| 8468. | +/- | 342. |
| 3.85 | +/- | 0. |
| 4.35 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.81 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.42 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| -1.98 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| -2.41 |
| -9999.00 |
357-03-O
| 3440. | +/- | 1. |
| 3521. | +/- | 56. |
| 0.68 | +/- | 0. |
| 0.50 | +/- | 0. |
| 1.55 | +/- | 0. |
| 1.55 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.66 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.20 |
| -9999.00 |
STAR_WARN,SN_WARN
357-03-O
| 3806. | +/- | 3. |
| 3880. | +/- | 82. |
| 1.55 | +/- | 0. |
| 1.30 | +/- | 0. |
| 1.45 | +/- | 0. |
| 1.45 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.41 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.39 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
357-03-O
| 11710. | +/- | 71. |
| -10000. | +/- | -NaN |
| 4.40 | +/- | 0. |
| 4.28 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -0.71 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 13.77 | +/- | 0. |
| 1.14 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.88 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.40 |
| -9999.00 |
357-03-O
| 3826. | +/- | 2. |
| 3896. | +/- | 75. |
| 1.65 | +/- | 0. |
| 1.38 | +/- | 0. |
| 1.52 | +/- | 0. |
| 1.52 | +/- | -NaN |
| 0.44 | +/- | 0. |
| 0.44 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.42 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.29 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.48 |
| -9999.00 |
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN
357-03-O
| 7712. | +/- | 13. |
| -10000. | +/- | -NaN |
| 4.59 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.16 | +/- | 0. |
| 1.16 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.94 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| 0.08 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
357-03-O
| 3566. | +/- | 1. |
| 3652. | +/- | 66. |
| 0.85 | +/- | 0. |
| 0.69 | +/- | 0. |
| 1.56 | +/- | 0. |
| 1.56 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.07 | +/- | 0. |
| 2.89 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
STAR_WARN,SN_WARN
357-03-O
| 3968. | +/- | 9. |
| 4101. | +/- | 104. |
| 1.41 | +/- | 0. |
| 1.38 | +/- | 0. |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| -1.05 | +/- | 0. |
| -1.05 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -0.19 | +/- | 0. |
| 5.48 | +/- | 0. |
| 0.74 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -0.88 |
| -9999.00 |
STAR_WARN,SN_WARN
357-03-O
| 3628. | +/- | 3. |
| 3698. | +/- | 80. |
| 1.31 | +/- | 0. |
| 1.09 | +/- | 0. |
| 1.33 | +/- | 0. |
| 1.33 | +/- | -NaN |
| 0.45 | +/- | 0. |
| 0.45 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.04 | +/- | 0. |
| 2.28 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.43 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 3756. | +/- | 9. |
| -10000. | +/- | -NaN |
| 1.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.22 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.10 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.69 |
| -9999.00 |
357-03-O
| 3650. | +/- | 1. |
| 3752. | +/- | 64. |
| 0.99 | +/- | 0. |
| 0.88 | +/- | 0. |
| 1.89 | +/- | 0. |
| 1.89 | +/- | -NaN |
| -0.33 | +/- | 0. |
| -0.33 | +/- | 0. |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.26 | +/- | 0. |
| 3.59 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.74 |
| -9999.00 |
357-03-O
| 3527. | +/- | 2. |
| 3596. | +/- | 72. |
| 1.25 | +/- | 0. |
| 1.04 | +/- | 0. |
| 1.33 | +/- | 0. |
| 1.33 | +/- | -NaN |
| 0.46 | +/- | 0. |
| 0.46 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.26 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.51 |
| -9999.00 |
STAR_BAD
357-03-O
| 7999. | +/- | 15. |
| -10000. | +/- | -NaN |
| 4.94 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.00 | +/- | 0. |
| 1.00 | +/- | -NaN |
| -1.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 73.16 | +/- | 0. |
| 1.86 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.75 |
| -9999.00 |
357-03-O
| 3136. | +/- | 1. |
| 3220. | +/- | 52. |
| 0.08 | +/- | 0. |
| -0.09 | +/- | 0. |
| 1.53 | +/- | 0. |
| 1.53 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.79 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.13 |
| -9999.00 |
BRIGHT_NEIGHBOR
357-03-O
| 3642. | +/- | 1. |
| 3709. | +/- | 67. |
| 1.57 | +/- | 0. |
| 1.30 | +/- | 0. |
| 1.31 | +/- | 0. |
| 1.31 | +/- | -NaN |
| 0.52 | +/- | 0. |
| 0.52 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.37 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.19 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.56 |
| -9999.00 |
357-03-O
| 3846. | +/- | 1. |
| 3913. | +/- | 72. |
| 1.85 | +/- | 0. |
| 1.55 | +/- | 0. |
| 1.57 | +/- | 0. |
| 1.57 | +/- | -NaN |
| 0.50 | +/- | 0. |
| 0.50 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.20 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.79 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
357-03-O
| 8550. | +/- | 31. |
| 8448. | +/- | 344. |
| 4.39 | +/- | 0. |
| 4.93 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.34 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.23 | +/- | 0. |
| 0.79 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.46 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -2.18 |
| -9999.00 |
357-03-O
| 3570. | +/- | 1. |
| 3638. | +/- | 60. |
| 1.25 | +/- | 0. |
| 1.03 | +/- | 0. |
| 1.40 | +/- | 0. |
| 1.40 | +/- | -NaN |
| 0.46 | +/- | 0. |
| 0.46 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.26 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.70 |
| -9999.00 |
357-03-O
| 4343. | +/- | 4. |
| 4417. | +/- | 93. |
| 2.50 | +/- | 0. |
| 2.23 | +/- | 0. |
| 1.37 | +/- | 0. |
| 1.37 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.44 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| 0.27 |
| -9999.00 |
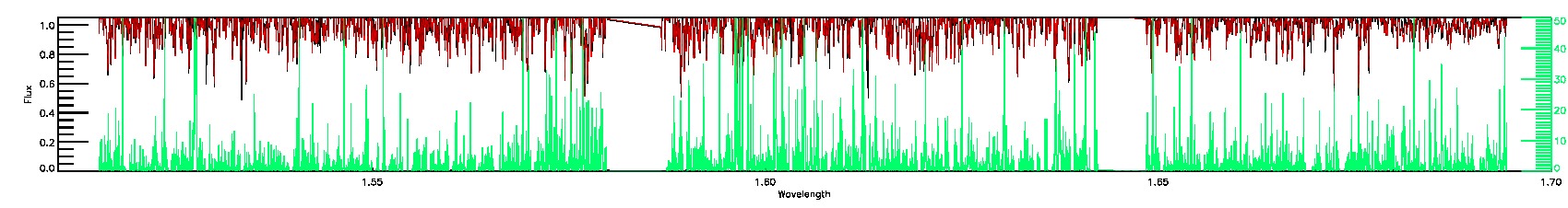
357-03-O
| 3374. | +/- | 1. |
| 3480. | +/- | 59. |
| 0.51 | +/- | 0. |
| 0.41 | +/- | 0. |
| 2.43 | +/- | 0. |
| 2.43 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -0.40 | +/- | 0. |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.26 | +/- | 0. |
| 3.75 | +/- | 0. |
| 0.57 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.20 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.64 |
| -9999.00 |
357-03-O
| 3878. | +/- | 1. |
| 3976. | +/- | 83. |
| 1.30 | +/- | 0. |
| 1.17 | +/- | 0. |
| 1.71 | +/- | 0. |
| 1.71 | +/- | -NaN |
| -0.21 | +/- | 0. |
| -0.21 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.14 | +/- | 0. |
| 3.34 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.01 |
| -9999.00 |
357-03-O
| 3540. | +/- | 1. |
| 3648. | +/- | 62. |
| 0.70 | +/- | 0. |
| 0.61 | +/- | 0. |
| 2.64 | +/- | 0. |
| 2.64 | +/- | -NaN |
| -0.47 | +/- | 0. |
| -0.47 | +/- | 0. |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.23 | +/- | 0. |
| 3.88 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -0.88 |
| -9999.00 |
BRIGHT_NEIGHBOR
357-03-O
| 3772. | +/- | 3. |
| 3843. | +/- | 80. |
| 1.60 | +/- | 0. |
| 1.34 | +/- | 0. |
| 1.24 | +/- | 0. |
| 1.24 | +/- | -NaN |
| 0.41 | +/- | 0. |
| 0.41 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.33 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.74 |
| -9999.00 |
STAR_BAD
STAR_WARN,COLORTE_WARN
357-03-O
| 7696. | +/- | 16. |
| -10000. | +/- | -NaN |
| 3.93 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.01 | +/- | 0. |
| 1.01 | +/- | -NaN |
| -1.73 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.72 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| 0.20 |
| -9999.00 |
357-03-O
| 4479. | +/- | 6. |
| 4550. | +/- | 97. |
| 2.77 | +/- | 0. |
| 2.44 | +/- | 0. |
| 1.18 | +/- | 0. |
| 1.18 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | 0. |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.35 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 0.74 |
| -9999.00 |

357-03-O
| 4269. | +/- | 3. |
| 4368. | +/- | 94. |
| 2.07 | +/- | 0. |
| 1.87 | +/- | 0. |
| 1.61 | +/- | 0. |
| 1.61 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -0.25 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.16 | +/- | 0. |
| 3.42 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.63 |
| -9999.00 |
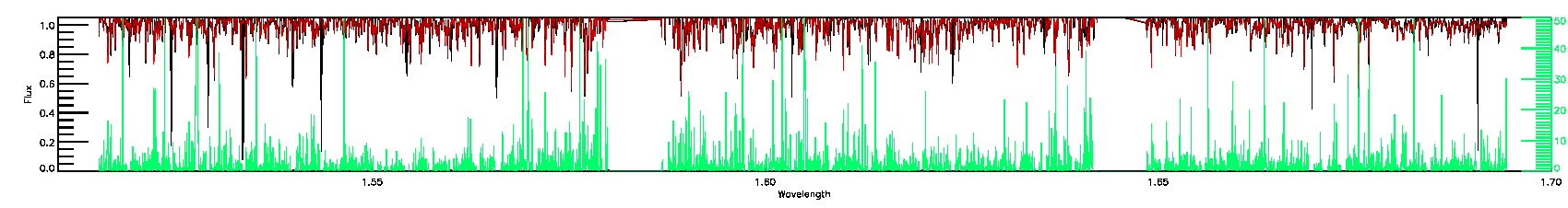
357-03-O
| 3772. | +/- | 2. |
| 3860. | +/- | 81. |
| 1.23 | +/- | 0. |
| 1.07 | +/- | 0. |
| 1.59 | +/- | 0. |
| 1.59 | +/- | -NaN |
| 0.02 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.14 | +/- | 0. |
| 2.93 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.33 |
| -9999.00 |
357-03-O
| 4596. | +/- | 12. |
| 4724. | +/- | 119. |
| 1.82 | +/- | 0. |
| 1.75 | +/- | 0. |
| 2.05 | +/- | 0. |
| 2.05 | +/- | -NaN |
| -1.18 | +/- | 0. |
| -1.18 | +/- | 0. |
| -0.18 | +/- | 0. |
| -0.18 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.30 | +/- | 0. |
| 5.91 | +/- | 0. |
| 0.77 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.22 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -1.11 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
357-03-O
| 3507. | +/- | 1. |
| 3614. | +/- | 77. |
| -0.04 | +/- | 0. |
| -0.14 | +/- | 0. |
| 2.45 | +/- | 0. |
| 2.45 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -0.43 | +/- | 0. |
| -0.01 | +/- | 0. |
| -0.01 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.81 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -0.45 |
| -9999.00 |
STAR_WARN,SN_WARN
357-03-O
| 3803. | +/- | 2. |
| 3898. | +/- | 88. |
| 1.19 | +/- | 0. |
| 1.06 | +/- | 0. |
| 1.55 | +/- | 0. |
| 1.55 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -0.16 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.08 | +/- | 0. |
| 3.25 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.29 |
| -9999.00 |
357-03-O
| 3789. | +/- | 2. |
| 3862. | +/- | 75. |
| 1.59 | +/- | 0. |
| 1.34 | +/- | 0. |
| 1.24 | +/- | 0. |
| 1.24 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.40 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 1.00 |
| -9999.00 |
357-03-O
| 3161. | +/- | 1. |
| 3240. | +/- | 51. |
| 0.27 | +/- | 0. |
| 0.08 | +/- | 0. |
| 1.71 | +/- | 0. |
| 1.71 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.58 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.26 |
| -9999.00 |
BRIGHT_NEIGHBOR
357-03-O
| 3840. | +/- | 2. |
| 3913. | +/- | 77. |
| 1.71 | +/- | 0. |
| 1.44 | +/- | 0. |
| 0.51 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.41 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.39 |
| -9999.00 |
357-03-O
| 3910. | +/- | 2. |
| 4008. | +/- | 84. |
| 1.57 | +/- | 0. |
| 1.40 | +/- | 0. |
| 1.64 | +/- | 0. |
| 1.64 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -0.22 | +/- | 0. |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.22 | +/- | 0. |
| 3.37 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
BRIGHT_NEIGHBOR
357-03-O
| 4216. | +/- | 3. |
| 4293. | +/- | 90. |
| 2.31 | +/- | 0. |
| 2.04 | +/- | 0. |
| 1.40 | +/- | 0. |
| 1.40 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.55 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.52 |
| -9999.00 |

STAR_BAD
STAR_WARN,COLORTE_WARN
357-03-O
| 8295. | +/- | 24. |
| -10000. | +/- | -NaN |
| 4.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -0.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.94 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.68 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| 0.64 |
| -9999.00 |
357-03-O
| 3842. | +/- | 2. |
| 3910. | +/- | 77. |
| 1.73 | +/- | 0. |
| 1.45 | +/- | 0. |
| 1.34 | +/- | 0. |
| 1.34 | +/- | -NaN |
| 0.46 | +/- | 0. |
| 0.46 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.26 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.74 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD
STAR_WARN,SN_WARN
357-03-O
| 6432. | +/- | 44. |
| -10000. | +/- | -NaN |
| 2.85 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.65 | +/- | 0. |
| 4.65 | +/- | -NaN |
| -1.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.23 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 11.80 | +/- | 0. |
| 1.07 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -1.69 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.92 |
| -9999.00 |
357-03-O
| 3804. | +/- | 1. |
| 3888. | +/- | 79. |
| 1.29 | +/- | 0. |
| 1.12 | +/- | 0. |
| 1.64 | +/- | 0. |
| 1.64 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.10 | +/- | 0. |
| 2.78 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.07 |
| -9999.00 |
VERY_BRIGHT_NEIGHBOR
STAR_WARN,SN_WARN
357-03-O
| 3922. | +/- | 3. |
| 4020. | +/- | 91. |
| 1.45 | +/- | 0. |
| 1.30 | +/- | 0. |
| 1.71 | +/- | 0. |
| 1.71 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -0.22 | +/- | 0. |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.20 | +/- | 0. |
| 3.37 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.60 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_WARN,COLORTE_WARN
357-03-O
| 4210. | +/- | 3. |
| 4332. | +/- | 103. |
| 1.71 | +/- | 0. |
| 1.60 | +/- | 0. |
| 1.65 | +/- | 0. |
| 1.65 | +/- | -NaN |
| -0.78 | +/- | 0. |
| -0.78 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.26 | +/- | 0. |
| 4.67 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| -1.35 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
357-03-O
| 3144. | +/- | 1. |
| 3229. | +/- | 52. |
| -0.22 | +/- | 0. |
| -0.38 | +/- | 0. |
| 1.34 | +/- | 0. |
| 1.34 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.10 | +/- | 0. |
| 2.84 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.97 |
| -9999.00 |
357-03-O
| 3884. | +/- | 2. |
| 3952. | +/- | 81. |
| 1.82 | +/- | 0. |
| 1.53 | +/- | 0. |
| 1.41 | +/- | 0. |
| 1.41 | +/- | -NaN |
| 0.46 | +/- | 0. |
| 0.46 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.25 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.74 |
| -9999.00 |
STAR_WARN,SN_WARN
357-03-O
| 4524. | +/- | 9. |
| 4611. | +/- | 106. |
| 2.82 | +/- | 0. |
| 2.61 | +/- | 0. |
| 0.79 | +/- | 0. |
| 0.79 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.10 | +/- | 0. |
| 3.04 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.77 |
| -9999.00 |
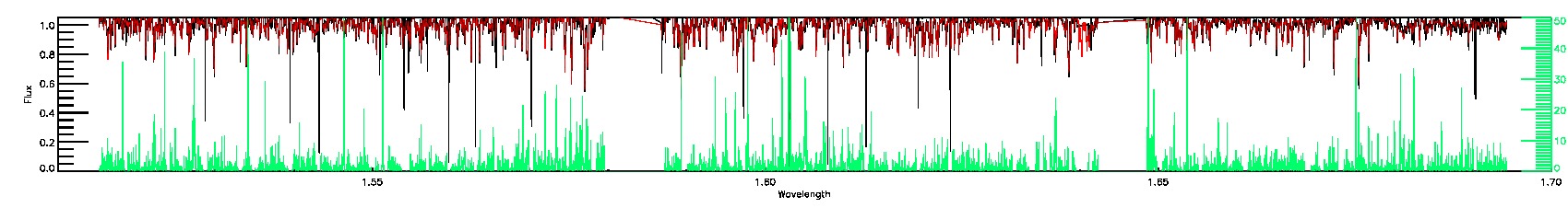
STAR_WARN,COLORTE_WARN
357-03-O
| 3192. | +/- | 1. |
| 3267. | +/- | 52. |
| 0.86 | +/- | 0. |
| 0.67 | +/- | 0. |
| 1.66 | +/- | 0. |
| 1.66 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | 0. |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.40 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.20 | +/- | 0. |
| 2.48 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.20 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.78 |
| -9999.00 |
BRIGHT_NEIGHBOR
357-03-O
| 3884. | +/- | 1. |
| 3968. | +/- | 78. |
| 1.59 | +/- | 0. |
| 1.37 | +/- | 0. |
| 1.59 | +/- | 0. |
| 1.59 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.19 | +/- | 0. |
| 2.78 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.26 |
| -9999.00 |
357-03-O
| 3889. | +/- | 2. |
| 3958. | +/- | 74. |
| 1.91 | +/- | 0. |
| 1.62 | +/- | 0. |
| 0.87 | +/- | 0. |
| 0.87 | +/- | -NaN |
| 0.45 | +/- | 0. |
| 0.45 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.27 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.73 |
| -9999.00 |
BRIGHT_NEIGHBOR
357-03-O
| 3665. | +/- | 1. |
| 3743. | +/- | 69. |
| 1.23 | +/- | 0. |
| 1.04 | +/- | 0. |
| 1.48 | +/- | 0. |
| 1.48 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.56 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.77 |
| -9999.00 |
357-03-O
| 3895. | +/- | 1. |
| 3981. | +/- | 81. |
| 1.59 | +/- | 0. |
| 1.38 | +/- | 0. |
| 1.45 | +/- | 0. |
| 1.45 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.06 | +/- | 0. |
| 2.84 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.06 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
357-03-O
| 3366. | +/- | 1. |
| 3478. | +/- | 60. |
| 0.45 | +/- | 0. |
| 0.37 | +/- | 0. |
| 2.32 | +/- | 0. |
| 2.32 | +/- | -NaN |
| -0.54 | +/- | 0. |
| -0.54 | +/- | 0. |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.28 | +/- | 0. |
| 4.07 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.21 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.75 |
| -9999.00 |
357-03-O
| 3854. | +/- | 1. |
| 3956. | +/- | 81. |
| 1.32 | +/- | 0. |
| 1.20 | +/- | 0. |
| 1.71 | +/- | 0. |
| 1.71 | +/- | -NaN |
| -0.32 | +/- | 0. |
| -0.32 | +/- | 0. |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.21 | +/- | 0. |
| 3.56 | +/- | 0. |
| 0.55 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.67 |
| -9999.00 |
357-03-O
| 3582. | +/- | 1. |
| 3657. | +/- | 65. |
| 1.16 | +/- | 0. |
| 0.97 | +/- | 0. |
| 1.47 | +/- | 0. |
| 1.47 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.25 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.05 | +/- | 0. |
| 2.46 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.63 |
| -9999.00 |
STAR_WARN,SN_WARN
357-03-O
| 3450. | +/- | 3. |
| 3524. | +/- | 78. |
| 0.92 | +/- | 0. |
| 0.72 | +/- | 0. |
| 1.37 | +/- | 0. |
| 1.37 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.41 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.43 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.40 |
| -9999.00 |
357-03-O
| 4120. | +/- | 2. |
| 4208. | +/- | 89. |
| 1.87 | +/- | 0. |
| 1.64 | +/- | 0. |
| 1.40 | +/- | 0. |
| 1.40 | +/- | -NaN |
| 0.01 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.12 | +/- | 0. |
| 2.94 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.50 |
| -9999.00 |

357-03-O
| 3688. | +/- | 1. |
| 3800. | +/- | 81. |
| 0.87 | +/- | 0. |
| 0.79 | +/- | 0. |
| 2.11 | +/- | 0. |
| 2.11 | +/- | -NaN |
| -0.56 | +/- | 0. |
| -0.56 | +/- | 0. |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.28 | +/- | 0. |
| 4.11 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -1.19 |
| -9999.00 |
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN
357-03-O
| 5755. | +/- | 42. |
| -10000. | +/- | -NaN |
| 3.95 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.99 | +/- | 0. |
| 2.99 | +/- | -NaN |
| -2.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| -0.42 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 93.91 | +/- | 0. |
| 1.97 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -2.49 |
| -9999.00 |

357-03-O
| 3757. | +/- | 2. |
| 3828. | +/- | 71. |
| 1.46 | +/- | 0. |
| 1.22 | +/- | 0. |
| 1.29 | +/- | 0. |
| 1.29 | +/- | -NaN |
| 0.40 | +/- | 0. |
| 0.40 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.34 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.99 |
| -9999.00 |
357-03-O
| 3759. | +/- | 1. |
| 3855. | +/- | 73. |
| 1.16 | +/- | 0. |
| 1.03 | +/- | 0. |
| 1.73 | +/- | 0. |
| 1.73 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -0.18 | +/- | 0. |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.20 | +/- | 0. |
| 3.29 | +/- | 0. |
| 0.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.07 |
| -9999.00 |
357-03-O
| 4244. | +/- | 4. |
| 4367. | +/- | 103. |
| 1.89 | +/- | 0. |
| 1.77 | +/- | 0. |
| 1.63 | +/- | 0. |
| 1.63 | +/- | -NaN |
| -0.80 | +/- | 0. |
| -0.80 | +/- | 0. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.23 | +/- | 0. |
| 4.71 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.97 |
| -9999.00 |

357-03-O
| 4055. | +/- | 3. |
| 4121. | +/- | 84. |
| 2.11 | +/- | 0. |
| 1.80 | +/- | 0. |
| 1.29 | +/- | 0. |
| 1.29 | +/- | -NaN |
| 0.51 | +/- | 0. |
| 0.51 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.19 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.98 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
357-03-O
| 4045. | +/- | 2. |
| 4162. | +/- | 86. |
| 1.50 | +/- | 0. |
| 1.40 | +/- | 0. |
| 1.75 | +/- | 0. |
| 1.75 | +/- | -NaN |
| -0.66 | +/- | 0. |
| -0.66 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.24 | +/- | 0. |
| 4.35 | +/- | 0. |
| 0.64 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.65 |
| -9999.00 |

357-03-O
| 3717. | +/- | 2. |
| 3800. | +/- | 76. |
| 1.51 | +/- | 0. |
| 1.30 | +/- | 0. |
| 1.41 | +/- | 0. |
| 1.41 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | 0. |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.19 | +/- | 0. |
| 2.75 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.22 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| 0.08 |
| -9999.00 |
STAR_WARN,COLORTE_WARN,SN_WARN
357-03-O
| 4246. | +/- | 6. |
| 4385. | +/- | 111. |
| 1.74 | +/- | 0. |
| 1.68 | +/- | 0. |
| 1.21 | +/- | 0. |
| 1.21 | +/- | -NaN |
| -1.18 | +/- | 0. |
| -1.17 | +/- | 0. |
| -0.14 | +/- | 0. |
| -0.14 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.25 | +/- | 0. |
| 5.88 | +/- | 0. |
| 0.77 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.14 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| -0.94 |
| -9999.00 |

357-03-O
| 3642. | +/- | 1. |
| 3731. | +/- | 68. |
| 1.10 | +/- | 0. |
| 0.95 | +/- | 0. |
| 1.73 | +/- | 0. |
| 1.73 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.21 | +/- | 0. |
| 3.00 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_WARN,COLORTE_WARN
357-03-O
| 8934. | +/- | 52. |
| 8800. | +/- | 352. |
| 4.27 | +/- | 0. |
| 4.50 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -1.57 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 85.70 | +/- | 0. |
| 1.93 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.58 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -1.36 |
| -9999.00 |
357-03-O
| 3387. | +/- | 1. |
| 3464. | +/- | 55. |
| 0.72 | +/- | 0. |
| 0.53 | +/- | 0. |
| 1.48 | +/- | 0. |
| 1.48 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.27 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.37 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.53 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.57 |
| -9999.00 |
357-03-O
| 3644. | +/- | 1. |
| 3722. | +/- | 69. |
| 1.14 | +/- | 0. |
| 0.95 | +/- | 0. |
| 1.52 | +/- | 0. |
| 1.52 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.24 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.56 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.54 |
| -9999.00 |
357-03-O
| 3754. | +/- | 2. |
| 3824. | +/- | 76. |
| 1.69 | +/- | 0. |
| 1.42 | +/- | 0. |
| 1.36 | +/- | 0. |
| 1.36 | +/- | -NaN |
| 0.43 | +/- | 0. |
| 0.43 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.04 | +/- | 0. |
| 2.30 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.45 |
| -9999.00 |
357-03-O
| 3512. | +/- | 1. |
| 3578. | +/- | 55. |
| 1.20 | +/- | 0. |
| 0.98 | +/- | 0. |
| 1.48 | +/- | 0. |
| 1.48 | +/- | -NaN |
| 0.52 | +/- | 0. |
| 0.52 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.18 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.56 |
| -9999.00 |
357-03-O
| 4409. | +/- | 3. |
| 4485. | +/- | 86. |
| 2.59 | +/- | 0. |
| 2.33 | +/- | 0. |
| 1.36 | +/- | 0. |
| 1.36 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.41 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.48 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.44 |
| -9999.00 |

357-03-O
| 4548. | +/- | 3. |
| 4625. | +/- | 79. |
| 2.64 | +/- | 0. |
| 2.35 | +/- | 0. |
| 1.44 | +/- | 0. |
| 1.44 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.73 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.18 |
| -9999.00 |

357-03-O
| 3988. | +/- | 3. |
| 4079. | +/- | 89. |
| 1.83 | +/- | 0. |
| 1.62 | +/- | 0. |
| 1.47 | +/- | 0. |
| 1.47 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -0.07 | +/- | 0. |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.19 | +/- | 0. |
| 3.08 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.16 |
| -9999.00 |

357-03-O
| 4746. | +/- | 12. |
| 4824. | +/- | 115. |
| 2.68 | +/- | 0. |
| 2.37 | +/- | 0. |
| 1.39 | +/- | 0. |
| 1.39 | +/- | -NaN |
| -0.43 | +/- | 0. |
| -0.42 | +/- | 0. |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.21 | +/- | 0. |
| 3.79 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -1.25 |
| -9999.00 |

PERSIST_JUMP_NEG
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 3378. | +/- | 18. |
| -10000. | +/- | -NaN |
| 0.63 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.20 | +/- | 0. |
| 1.20 | +/- | -NaN |
| 0.92 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.72 | +/- | 0. |
| 0.24 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.49 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 2.00 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.90 |
| -9999.00 |
357-03-O
| 4100. | +/- | 7. |
| 4215. | +/- | 98. |
| 1.95 | +/- | 0. |
| 1.80 | +/- | 0. |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| -0.61 | +/- | 0. |
| -0.61 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -0.23 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -0.19 | +/- | 0. |
| 4.23 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.33 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
357-03-O
| 3933. | +/- | 2. |
| 4031. | +/- | 85. |
| 1.58 | +/- | 0. |
| 1.41 | +/- | 0. |
| 1.86 | +/- | 0. |
| 1.86 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -0.23 | +/- | 0. |
| -0.13 | +/- | 0. |
| -0.13 | +/- | -NaN |
| 0.59 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.16 | +/- | 0. |
| 3.39 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.14 |
| -9999.00 |
357-03-O
| 4499. | +/- | 3. |
| 4575. | +/- | 80. |
| 2.87 | +/- | 0. |
| 2.63 | +/- | 0. |
| 1.20 | +/- | 0. |
| 1.20 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.48 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.22 |
| -9999.00 |

357-03-O
| 3631. | +/- | 1. |
| 3740. | +/- | 78. |
| 0.86 | +/- | 0. |
| 0.77 | +/- | 0. |
| 1.93 | +/- | 0. |
| 1.93 | +/- | -NaN |
| -0.48 | +/- | 0. |
| -0.48 | +/- | 0. |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.28 | +/- | 0. |
| 3.91 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.68 |
| -9999.00 |
357-03-O
| 3679. | +/- | 1. |
| 3761. | +/- | 73. |
| 1.48 | +/- | 0. |
| 1.27 | +/- | 0. |
| 1.56 | +/- | 0. |
| 1.56 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | 0. |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.20 | +/- | 0. |
| 2.69 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.22 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.16 |
| -9999.00 |
STAR_WARN,SN_WARN
357-03-O
| 3790. | +/- | 2. |
| 3881. | +/- | 86. |
| 1.34 | +/- | 0. |
| 1.18 | +/- | 0. |
| 1.54 | +/- | 0. |
| 1.54 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.16 | +/- | 0. |
| 3.05 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -0.11 |
| -9999.00 |
STAR_WARN,SN_WARN
357-03-O
| 4529. | +/- | 10. |
| 4607. | +/- | 107. |
| 2.67 | +/- | 0. |
| 2.37 | +/- | 0. |
| 0.67 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.16 | +/- | 0. |
| 0.16 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.69 | +/- | 0. |
| 0.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.16 |
| -9999.00 |

STAR_BAD
STAR_WARN,COLORTE_WARN
357-03-O
| 8593. | +/- | 40. |
| -10000. | +/- | -NaN |
| 4.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -1.34 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.28 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.47 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -1.65 |
| -9999.00 |
STAR_WARN,SN_WARN
357-03-O
| 4294. | +/- | 21. |
| 4419. | +/- | 118. |
| 1.68 | +/- | 0. |
| 1.58 | +/- | 0. |
| 0.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| -0.86 | +/- | 0. |
| -0.86 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.00 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -0.22 | +/- | 0. |
| 4.88 | +/- | 0. |
| 0.69 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -2.48 |
| -9999.00 |
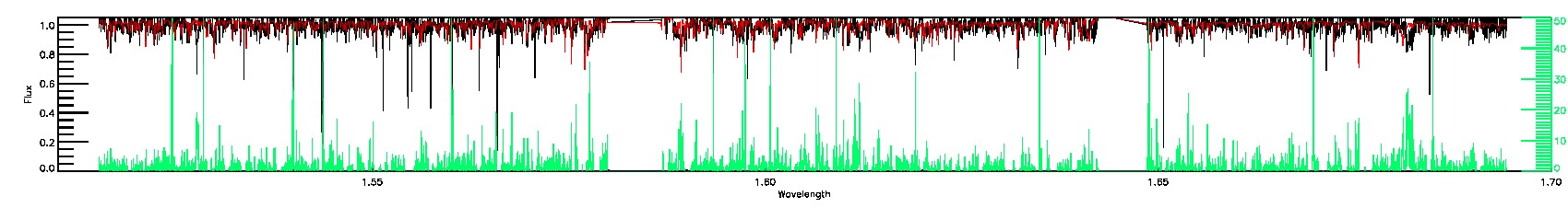
357-03-O
| 3642. | +/- | 1. |
| 3741. | +/- | 63. |
| 0.86 | +/- | 0. |
| 0.74 | +/- | 0. |
| 1.95 | +/- | 0. |
| 1.95 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.24 | +/- | 0. |
| 0.17 | +/- | 0. |
| 0.17 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.24 | +/- | 0. |
| 3.41 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.34 |
| -9999.00 |
357-03-O
| 3610. | +/- | 1. |
| 3743. | +/- | 68. |
| 0.27 | +/- | 0. |
| 0.25 | +/- | 0. |
| 2.62 | +/- | 0. |
| 2.62 | +/- | -NaN |
| -1.02 | +/- | 0. |
| -1.02 | +/- | 0. |
| -0.05 | +/- | 0. |
| -0.05 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.22 | +/- | 0. |
| 5.38 | +/- | 0. |
| 0.73 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -1.72 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
357-03-O
| 5681. | +/- | 80. |
| -10000. | +/- | -NaN |
| 2.21 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.76 | +/- | 1. |
| 4.76 | +/- | -NaN |
| -1.73 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.19 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.11 | +/- | 0. |
| 0.91 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -1.41 |
| -9999.00 |

STAR_WARN,SN_WARN
357-03-O
| 3696. | +/- | 3. |
| 3813. | +/- | 94. |
| 0.88 | +/- | 0. |
| 0.82 | +/- | 0. |
| 2.16 | +/- | 0. |
| 2.16 | +/- | -NaN |
| -0.68 | +/- | 0. |
| -0.68 | +/- | 0. |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.27 | +/- | 0. |
| 4.39 | +/- | 0. |
| 0.64 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -0.48 |
| -9999.00 |
357-03-O
| 4057. | +/- | 3. |
| 4147. | +/- | 91. |
| 1.87 | +/- | 0. |
| 1.64 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.43 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.17 | +/- | 0. |
| 0.14 | +/- | 0. |
| 3.04 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 1.00 |
| -9999.00 |

357-03-O
| 3792. | +/- | 1. |
| 3910. | +/- | 77. |
| 1.08 | +/- | 0. |
| 1.03 | +/- | 0. |
| 1.96 | +/- | 0. |
| 1.96 | +/- | -NaN |
| -0.70 | +/- | 0. |
| -0.70 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.27 | +/- | 0. |
| 4.45 | +/- | 0. |
| 0.65 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.86 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
357-03-O
| 3585. | +/- | 1. |
| 3654. | +/- | 69. |
| 1.36 | +/- | 0. |
| 1.13 | +/- | 0. |
| 1.44 | +/- | 0. |
| 1.44 | +/- | -NaN |
| 0.45 | +/- | 0. |
| 0.45 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.27 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.48 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 3344. | +/- | 16. |
| -10000. | +/- | -NaN |
| 0.32 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.40 | +/- | 0. |
| 2.40 | +/- | -NaN |
| 0.83 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.82 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.61 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 2.00 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 0.24 |
| -9999.00 |
357-03-O
| 4555. | +/- | 7. |
| 4635. | +/- | 102. |
| 2.94 | +/- | 0. |
| 2.74 | +/- | 0. |
| 0.79 | +/- | 0. |
| 0.79 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.85 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.07 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
357-03-O
| 4268. | +/- | 4. |
| 4358. | +/- | 94. |
| 2.27 | +/- | 0. |
| 2.04 | +/- | 0. |
| 1.29 | +/- | 0. |
| 1.29 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -0.02 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.03 | +/- | 0. |
| 3.00 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.14 |
| -9999.00 |

BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
357-03-O
| 6139. | +/- | 32. |
| -10000. | +/- | -NaN |
| 2.52 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.93 | +/- | 0. |
| 2.93 | +/- | -NaN |
| -1.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 0.34 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 15.03 | +/- | 0. |
| 1.18 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.27 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6332. | +/- | 115. |
| -10000. | +/- | -NaN |
| 2.65 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.61 | +/- | 1. |
| 1.61 | +/- | -NaN |
| -1.89 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| 0.99 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 92.26 | +/- | 0. |
| 1.97 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.21 |
| -9999.00 |
STAR_BAD
STAR_WARN,SN_WARN
357-03-O
| 6066. | +/- | 42. |
| -10000. | +/- | -NaN |
| 2.55 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.09 | +/- | 1. |
| 1.09 | +/- | -NaN |
| -1.96 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 17.77 | +/- | 0. |
| 1.25 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| -0.72 |
| -9999.00 |
STAR_WARN,SN_WARN
357-03-O
| 3848. | +/- | 3. |
| 3925. | +/- | 84. |
| 1.62 | +/- | 0. |
| 1.37 | +/- | 0. |
| 1.46 | +/- | 0. |
| 1.46 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.02 | +/- | 0. |
| 2.51 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.62 |
| -9999.00 |
357-03-O
| 4270. | +/- | 3. |
| 4350. | +/- | 90. |
| 2.45 | +/- | 0. |
| 2.19 | +/- | 0. |
| 1.26 | +/- | 0. |
| 1.26 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.61 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| 0.43 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
357-03-O
| 3324. | +/- | 1. |
| 3423. | +/- | 57. |
| 0.34 | +/- | 0. |
| 0.22 | +/- | 0. |
| 1.93 | +/- | 0. |
| 1.93 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.24 | +/- | 0. |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.22 | +/- | 0. |
| 3.41 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.17 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
357-03-O
| 8477. | +/- | 38. |
| 8362. | +/- | 344. |
| 4.28 | +/- | 0. |
| 4.69 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.03 | +/- | 0. |
| -2.02 | +/- | 1. |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.01 | +/- | 1. |
| 38.62 | +/- | 0. |
| 1.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.42 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -1.69 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -2.31 |
| -9999.00 |
357-03-O
| 3918. | +/- | 1. |
| 4044. | +/- | 86. |
| 1.15 | +/- | 0. |
| 1.12 | +/- | 0. |
| 2.03 | +/- | 0. |
| 2.03 | +/- | -NaN |
| -0.87 | +/- | 0. |
| -0.87 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.25 | +/- | 0. |
| 4.91 | +/- | 0. |
| 0.69 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -1.56 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 4872. | +/- | 79. |
| -10000. | +/- | -NaN |
| 2.26 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| -0.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.80 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.28 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.51 |
| -9999.00 |

STAR_WARN,SN_WARN
357-03-O
| 3953. | +/- | 3. |
| 4020. | +/- | 84. |
| 1.94 | +/- | 0. |
| 1.64 | +/- | 0. |
| 1.33 | +/- | 0. |
| 1.33 | +/- | -NaN |
| 0.49 | +/- | 0. |
| 0.49 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.29 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.22 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.77 |
| -9999.00 |
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN
357-03-O
| 6152. | +/- | 27. |
| -10000. | +/- | -NaN |
| 4.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.16 | +/- | 0. |
| 1.16 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.16 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.41 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -2.41 |
| -9999.00 |
357-03-O
| 3824. | +/- | 2. |
| 3919. | +/- | 83. |
| 1.39 | +/- | 0. |
| 1.24 | +/- | 0. |
| 1.65 | +/- | 0. |
| 1.65 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -0.16 | +/- | 0. |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.20 | +/- | 0. |
| 3.24 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.16 |
| -9999.00 |
STAR_WARN,COLORTE_WARN,SN_WARN
357-03-O
| 3348. | +/- | 3. |
| 3426. | +/- | 75. |
| 0.57 | +/- | 0. |
| 0.39 | +/- | 0. |
| 1.66 | +/- | 0. |
| 1.66 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.58 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.97 |
| -9999.00 |
STAR_WARN,SN_WARN
357-03-O
| 3956. | +/- | 4. |
| 4060. | +/- | 97. |
| 1.64 | +/- | 0. |
| 1.47 | +/- | 0. |
| 1.67 | +/- | 0. |
| 1.67 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -0.35 | +/- | 0. |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.22 | +/- | 0. |
| 3.62 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.62 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_WARN,COLORTE_WARN,SN_WARN
357-03-O
| 4017. | +/- | 4. |
| 4086. | +/- | 87. |
| 2.13 | +/- | 0. |
| 1.84 | +/- | 0. |
| 1.19 | +/- | 0. |
| 1.19 | +/- | -NaN |
| 0.45 | +/- | 0. |
| 0.45 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.27 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.47 |
| -9999.00 |

357-03-O
| 3554. | +/- | 1. |
| 3653. | +/- | 61. |
| 0.81 | +/- | 0. |
| 0.69 | +/- | 0. |
| 1.96 | +/- | 0. |
| 1.96 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -0.24 | +/- | 0. |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.24 | +/- | 0. |
| 3.41 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.62 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
357-03-O
| 3971. | +/- | 2. |
| 4050. | +/- | 83. |
| 1.72 | +/- | 0. |
| 1.47 | +/- | 0. |
| 1.50 | +/- | 0. |
| 1.50 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.07 | +/- | 0. |
| 2.62 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.38 |
| -9999.00 |
357-03-O
| 3447. | +/- | 1. |
| 3517. | +/- | 63. |
| 1.06 | +/- | 0. |
| 0.85 | +/- | 0. |
| 1.47 | +/- | 0. |
| 1.47 | +/- | -NaN |
| 0.44 | +/- | 0. |
| 0.44 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.01 | +/- | 0. |
| 2.29 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.98 |
| -9999.00 |
STAR_BAD
357-03-O
| 6963. | +/- | 16. |
| -10000. | +/- | -NaN |
| 3.85 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.34 | +/- | 0. |
| 1.34 | +/- | -NaN |
| -1.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.98 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -1.98 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.80 |
| -9999.00 |
357-03-O
| 3914. | +/- | 2. |
| 4000. | +/- | 83. |
| 1.62 | +/- | 0. |
| 1.40 | +/- | 0. |
| 1.57 | +/- | 0. |
| 1.57 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.16 | +/- | 0. |
| 2.85 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.16 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
357-03-O
| 3842. | +/- | 2. |
| 3912. | +/- | 80. |
| 1.80 | +/- | 0. |
| 1.52 | +/- | 0. |
| 1.40 | +/- | 0. |
| 1.40 | +/- | -NaN |
| 0.42 | +/- | 0. |
| 0.42 | +/- | 0. |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.31 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.45 |
| -9999.00 |
357-03-O
| 3657. | +/- | 1. |
| 3726. | +/- | 65. |
| 1.41 | +/- | 0. |
| 1.17 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.43 | +/- | -NaN |
| 0.47 | +/- | 0. |
| 0.47 | +/- | 0. |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.01 | +/- | 0. |
| 2.25 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 1.00 |
| -9999.00 |
357-03-O
| 3985. | +/- | 2. |
| 4105. | +/- | 94. |
| 1.23 | +/- | 0. |
| 1.18 | +/- | 0. |
| 2.12 | +/- | 0. |
| 2.12 | +/- | -NaN |
| -0.73 | +/- | 0. |
| -0.73 | +/- | 0. |
| -0.03 | +/- | 0. |
| -0.03 | +/- | -NaN |
| 0.18 | +/- | 0. |
| 0.18 | +/- | -NaN |
| 0.24 | +/- | 0. |
| 0.21 | +/- | 0. |
| 4.54 | +/- | 0. |
| 0.66 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -1.54 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
357-03-O
| 3874. | +/- | 1. |
| 3983. | +/- | 80. |
| 1.33 | +/- | 0. |
| 1.23 | +/- | 0. |
| 1.78 | +/- | 0. |
| 1.78 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -0.50 | +/- | 0. |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.13 | +/- | -NaN |
| 0.27 | +/- | 0. |
| 0.24 | +/- | 0. |
| 3.96 | +/- | 0. |
| 0.60 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.86 |
| -9999.00 |
357-03-O
| 3371. | +/- | 1. |
| 3462. | +/- | 57. |
| 0.32 | +/- | 0. |
| 0.18 | +/- | 0. |
| 1.69 | +/- | 0. |
| 1.69 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -0.04 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.13 | +/- | 0. |
| 0.10 | +/- | 0. |
| 3.03 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.10 |
| -9999.00 |
357-03-O
| 3732. | +/- | 2. |
| 3798. | +/- | 73. |
| 1.72 | +/- | 0. |
| 1.43 | +/- | 0. |
| 1.35 | +/- | 0. |
| 1.35 | +/- | -NaN |
| 0.52 | +/- | 0. |
| 0.52 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.19 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.99 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
357-03-O
| 3242. | +/- | 1. |
| 3314. | +/- | 52. |
| 0.65 | +/- | 0. |
| 0.45 | +/- | 0. |
| 1.55 | +/- | 0. |
| 1.55 | +/- | -NaN |
| 0.37 | +/- | 0. |
| 0.37 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.49 | +/- | 0. |
| 0.49 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.07 | +/- | 0. |
| 2.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.93 |
| -9999.00 |
357-03-O
| 3634. | +/- | 1. |
| 3729. | +/- | 70. |
| 1.19 | +/- | 0. |
| 1.05 | +/- | 0. |
| 1.53 | +/- | 0. |
| 1.53 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -0.14 | +/- | 0. |
| 0.21 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.20 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.21 | +/- | 0. |
| 3.21 | +/- | 0. |
| 0.51 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.05 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
357-03-O
| 4413. | +/- | 20. |
| -10000. | +/- | -NaN |
| 2.42 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.94 | +/- | 0. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.69 |
| -9999.00 |
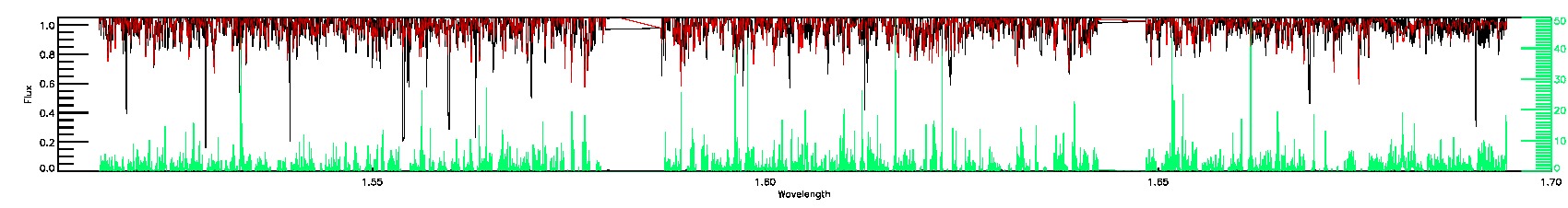
STAR_WARN,COLORTE_WARN,SN_WARN
357-03-O
| 4671. | +/- | 11. |
| 4730. | +/- | 108. |
| 2.87 | +/- | 0. |
| 2.51 | +/- | 0. |
| 1.13 | +/- | 0. |
| 1.13 | +/- | -NaN |
| 0.21 | +/- | 0. |
| 0.21 | +/- | 0. |
| 0.02 | +/- | 0. |
| 0.02 | +/- | -NaN |
| 0.29 | +/- | 0. |
| 0.29 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.61 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.60 |
| -9999.00 |

357-03-O
| 3921. | +/- | 1. |
| 4043. | +/- | 82. |
| 1.13 | +/- | 0. |
| 1.09 | +/- | 0. |
| 2.05 | +/- | 0. |
| 2.05 | +/- | -NaN |
| -0.81 | +/- | 0. |
| -0.81 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.23 | +/- | 0. |
| 4.74 | +/- | 0. |
| 0.68 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.80 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,SN_WARN
357-03-O
| 5928. | +/- | 39. |
| -10000. | +/- | -NaN |
| 2.93 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.35 | +/- | 1. |
| 1.35 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.24 | +/- | 1. |
| 0.35 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.17 |
| -9999.00 |
357-03-O
| 3769. | +/- | 2. |
| 3862. | +/- | 83. |
| 1.43 | +/- | 0. |
| 1.26 | +/- | 0. |
| 1.12 | +/- | 0. |
| 1.12 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -0.11 | +/- | 0. |
| 0.22 | +/- | 0. |
| 0.22 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.19 | +/- | 0. |
| 3.15 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.20 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.53 |
| -9999.00 |
STAR_WARN,SN_WARN
357-03-O
| 4277. | +/- | 15. |
| 4380. | +/- | 111. |
| 1.84 | +/- | 0. |
| 1.65 | +/- | 0. |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -0.35 | +/- | 0. |
| -0.17 | +/- | 0. |
| -0.17 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.11 | +/- | 0. |
| 3.62 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.18 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.96 |
| -9999.00 |
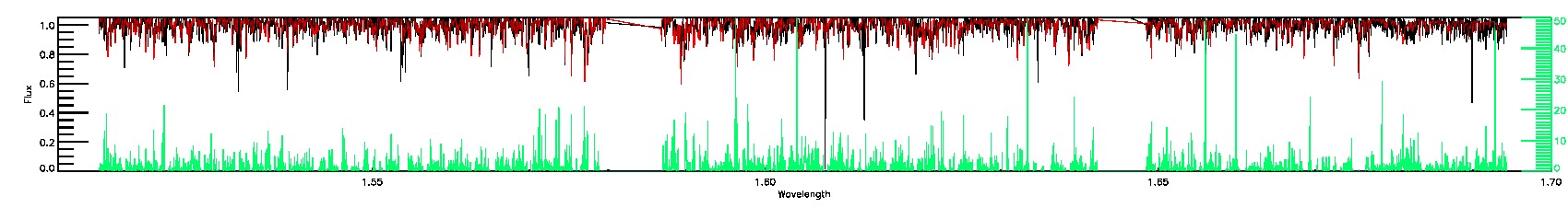
357-03-O
| 3341. | +/- | 1. |
| 3420. | +/- | 55. |
| 0.58 | +/- | 0. |
| 0.39 | +/- | 0. |
| 1.62 | +/- | 0. |
| 1.62 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.22 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -0.00 | +/- | 0. |
| 2.60 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.26 |
| -9999.00 |
357-03-O
| 3876. | +/- | 1. |
| 3980. | +/- | 80. |
| 1.47 | +/- | 0. |
| 1.33 | +/- | 0. |
| 1.74 | +/- | 0. |
| 1.74 | +/- | -NaN |
| -0.37 | +/- | 0. |
| -0.37 | +/- | 0. |
| 0.16 | +/- | 0. |
| 0.16 | +/- | -NaN |
| 0.15 | +/- | 0. |
| 0.15 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.23 | +/- | 0. |
| 3.67 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -1.04 |
| -9999.00 |
STAR_WARN,SN_WARN
357-03-O
| 4404. | +/- | 10. |
| 4533. | +/- | 117. |
| 1.70 | +/- | 0. |
| 1.61 | +/- | 0. |
| 2.24 | +/- | 0. |
| 2.24 | +/- | -NaN |
| -0.95 | +/- | 0. |
| -0.95 | +/- | 0. |
| -0.07 | +/- | 0. |
| -0.07 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.27 | +/- | 0. |
| 5.15 | +/- | 0. |
| 0.71 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -1.22 |
| -9999.00 |
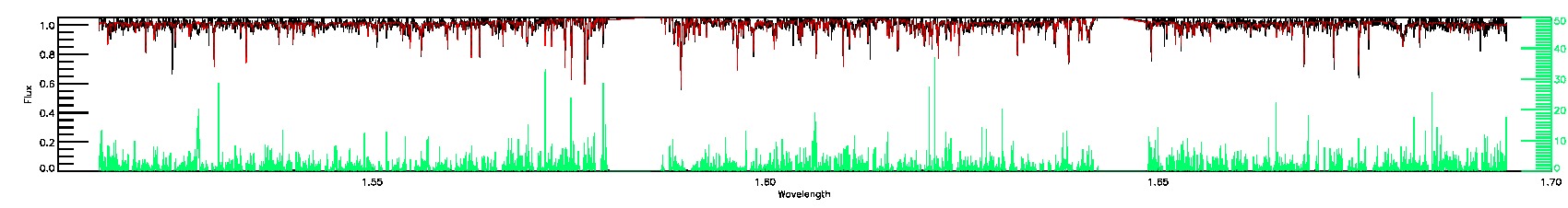
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6678. | +/- | 51. |
| -10000. | +/- | -NaN |
| 2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.28 | +/- | 0. |
| 1.28 | +/- | -NaN |
| -0.90 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.46 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 15.51 | +/- | 0. |
| 1.19 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| 0.99 |
| -9999.00 |
VERY_BRIGHT_NEIGHBOR
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6136. | +/- | 115. |
| -10000. | +/- | -NaN |
| 3.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.84 | +/- | 0. |
| 0.84 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 17.84 | +/- | 0. |
| 1.25 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 1.00 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6646. | +/- | 127. |
| -10000. | +/- | -NaN |
| 2.86 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.94 | +/- | 16. |
| 0.94 | +/- | -NaN |
| -2.09 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 9. |
| -9999.99 | +/- | -NaN |
| -0.43 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 8.13 | +/- | 2. |
| 0.91 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -1.98 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -2.26 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
357-03-O
| 5927. | +/- | 300. |
| -10000. | +/- | -NaN |
| 4.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.02 | +/- | 0. |
| 1.02 | +/- | -NaN |
| -0.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.32 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.46 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 2.00 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
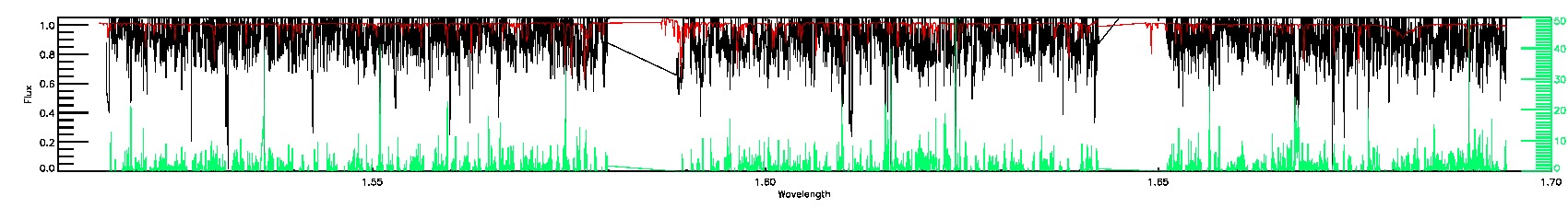
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
357-03-O
| 4648. | +/- | 59. |
| -10000. | +/- | -NaN |
| 1.86 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| -0.72 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.51 | +/- | 0. |
| 0.65 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.38 |
| -9999.00 |
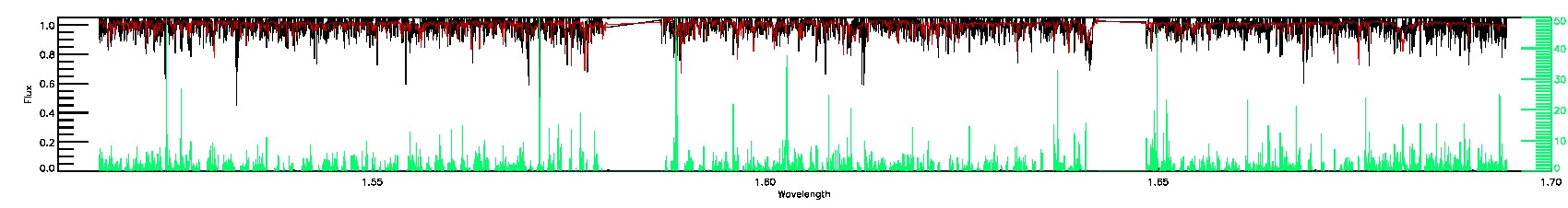
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 7891. | +/- | 219. |
| -10000. | +/- | -NaN |
| 2.55 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.97 | +/- | 1. |
| 1.97 | +/- | -NaN |
| -0.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.11 | +/- | 1. |
| 0.71 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| 0.80 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
357-03-O
| 4572. | +/- | 29. |
| -10000. | +/- | -NaN |
| 2.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.48 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| -1.53 |
| -9999.00 |

STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6156. | +/- | 138. |
| -10000. | +/- | -NaN |
| 3.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.76 | +/- | 0. |
| 4.76 | +/- | -NaN |
| -0.24 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.36 | +/- | 0. |
| 0.92 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 1.24 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -0.84 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6063. | +/- | 67. |
| -10000. | +/- | -NaN |
| 2.62 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.98 | +/- | 1. |
| 0.98 | +/- | -NaN |
| -1.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 20.37 | +/- | 0. |
| 1.31 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.08 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 5517. | +/- | 115. |
| -10000. | +/- | -NaN |
| 1.99 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.84 | +/- | 1. |
| 0.84 | +/- | -NaN |
| -1.41 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.41 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.76 | +/- | 0. |
| 0.83 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.24 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.51 |
| -9999.00 |

BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6854. | +/- | 109. |
| -10000. | +/- | -NaN |
| 2.74 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.74 | +/- | 1. |
| 0.74 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 18.91 | +/- | 0. |
| 1.28 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.30 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 7990. | +/- | 408. |
| -10000. | +/- | -NaN |
| 5.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.85 | +/- | 13. |
| 1.85 | +/- | -NaN |
| -1.76 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 12. |
| -9999.99 | +/- | -NaN |
| 0.17 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 95.32 | +/- | 1. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -2.33 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6250. | +/- | 97. |
| -10000. | +/- | -NaN |
| 2.63 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.80 | +/- | 0. |
| 2.80 | +/- | -NaN |
| -0.96 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.67 | +/- | 0. |
| 1.03 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| 0.97 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 4674. | +/- | 69. |
| -10000. | +/- | -NaN |
| 1.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -1.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.73 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.07 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.45 | +/- | 0. |
| 0.81 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.72 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.49 |
| -9999.00 |

BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6752. | +/- | 97. |
| -10000. | +/- | -NaN |
| 2.66 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.16 | +/- | 0. |
| 4.16 | +/- | -NaN |
| -0.92 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 13.59 | +/- | 0. |
| 1.13 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.23 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
357-03-O
| 3809. | +/- | 12. |
| -10000. | +/- | -NaN |
| 1.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -0.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.67 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.78 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
357-03-O
| 5912. | +/- | 91. |
| -10000. | +/- | -NaN |
| 2.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.23 | +/- | 2. |
| 2.23 | +/- | -NaN |
| -2.39 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.51 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 9. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.00 | +/- | 0. |
| 1.08 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.87 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| 0.99 |
| -9999.00 |
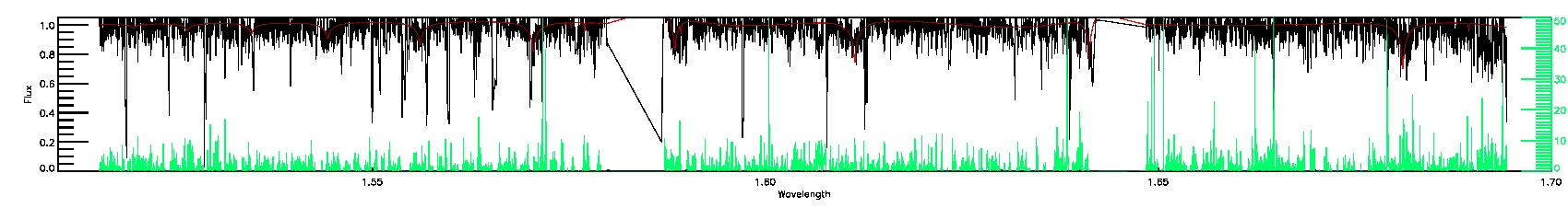
BRIGHT_NEIGHBOR
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6087. | +/- | 53. |
| -10000. | +/- | -NaN |
| 2.51 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.55 | +/- | 0. |
| 4.55 | +/- | -NaN |
| -1.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 18. |
| -9999.99 | +/- | -NaN |
| 0.53 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 11.28 | +/- | 0. |
| 1.05 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -2.46 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 5648. | +/- | 66. |
| -10000. | +/- | -NaN |
| 2.59 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.46 | +/- | 0. |
| 2.46 | +/- | -NaN |
| -1.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 16.42 | +/- | 0. |
| 1.22 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.46 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.64 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
357-03-O
| 5642. | +/- | 82. |
| -10000. | +/- | -NaN |
| 1.97 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.85 | +/- | 1. |
| 3.85 | +/- | -NaN |
| -1.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.25 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| 0.35 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.81 | +/- | 0. |
| 0.83 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.27 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.71 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
357-03-O
| 5787. | +/- | 188. |
| -10000. | +/- | -NaN |
| 3.58 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.94 | +/- | 1. |
| 0.94 | +/- | -NaN |
| -1.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.39 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.24 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
357-03-O
| 5520. | +/- | 73. |
| -10000. | +/- | -NaN |
| 1.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.10 | +/- | 1. |
| 2.10 | +/- | -NaN |
| -1.73 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.36 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.14 | +/- | 0. |
| 0.91 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.61 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.14 |
| -9999.00 |
| -1.57 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6766. | +/- | 80. |
| -10000. | +/- | -NaN |
| 2.55 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.39 | +/- | 1. |
| 1.39 | +/- | -NaN |
| -0.58 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| -0.42 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 73.94 | +/- | 0. |
| 1.87 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.67 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
357-03-O
| 4968. | +/- | 73. |
| -10000. | +/- | -NaN |
| 1.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.56 | +/- | 0. |
| 1.56 | +/- | -NaN |
| -1.90 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.97 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.32 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.67 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.97 | +/- | 0. |
| 0.95 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.95 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.90 |
| -9999.00 |
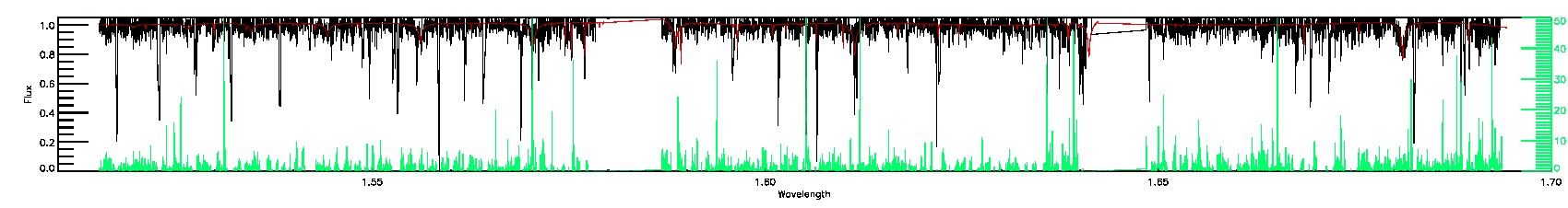
BRIGHT_NEIGHBOR
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
357-03-O
| 3953. | +/- | 35. |
| -10000. | +/- | -NaN |
| 0.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| -1.87 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.85 | +/- | 0. |
| 0.95 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| 0.95 |
| -9999.00 |
STAR_BAD
STAR_WARN,SN_WARN
357-03-O
| 6314. | +/- | 16. |
| -10000. | +/- | -NaN |
| 2.51 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.31 | +/- | 43. |
| 0.31 | +/- | -NaN |
| -2.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| -0.22 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 67.72 | +/- | 0. |
| 1.83 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.13 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 5514. | +/- | 23. |
| -10000. | +/- | -NaN |
| 2.72 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.66 | +/- | 0. |
| 0.66 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 31.20 | +/- | 0. |
| 1.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 1.50 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.20 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6241. | +/- | 43. |
| -10000. | +/- | -NaN |
| 2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.27 | +/- | 0. |
| 4.27 | +/- | -NaN |
| -1.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 6. |
| -9999.99 | +/- | -NaN |
| 0.60 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.87 | +/- | 0. |
| 0.27 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.36 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| -1.36 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
357-03-O
| 5612. | +/- | 69. |
| -10000. | +/- | -NaN |
| 1.63 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.78 | +/- | 1. |
| 4.78 | +/- | -NaN |
| -2.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.25 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.27 | +/- | 9. |
| -9999.99 | +/- | -NaN |
| 0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.43 | +/- | 0. |
| 1.02 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.40 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.49 |
| -9999.00 |
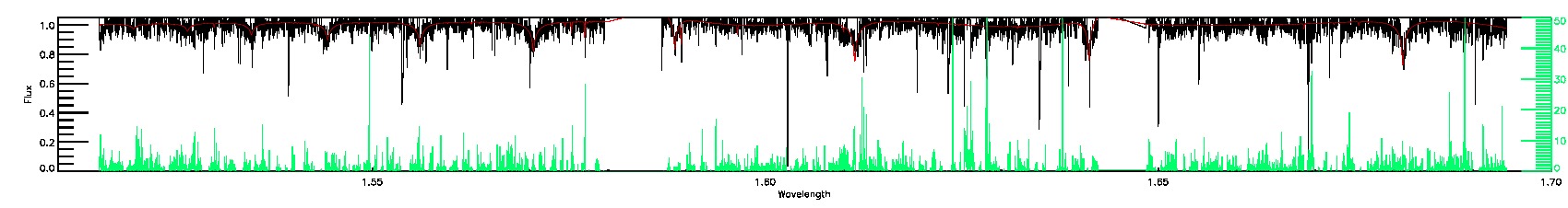
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6861. | +/- | 72. |
| -10000. | +/- | -NaN |
| 2.57 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.44 | +/- | 0. |
| 0.44 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 54.64 | +/- | 0. |
| 1.74 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.27 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6014. | +/- | 109. |
| -10000. | +/- | -NaN |
| 2.71 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.34 | +/- | 0. |
| 4.34 | +/- | -NaN |
| -1.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| 0.54 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 14.39 | +/- | 0. |
| 1.16 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.04 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
357-03-O
| 5954. | +/- | 95. |
| -10000. | +/- | -NaN |
| 2.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.93 | +/- | 1. |
| 1.93 | +/- | -NaN |
| -1.77 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 6. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.32 | +/- | 0. |
| 0.92 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -0.85 |
| -9999.00 |
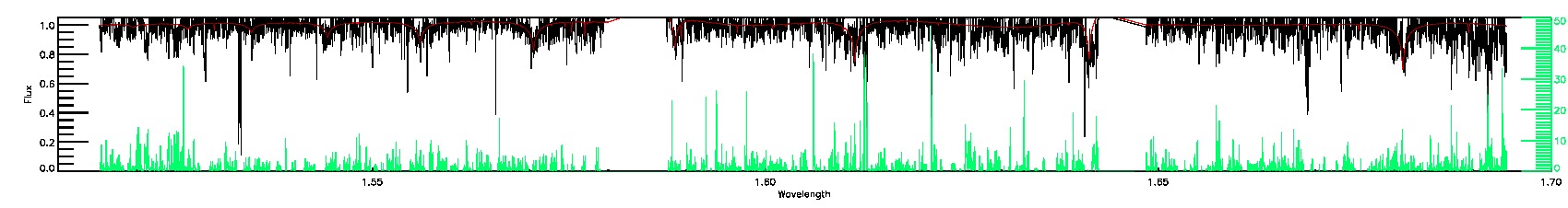
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 7415. | +/- | 146. |
| -10000. | +/- | -NaN |
| 3.32 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.67 | +/- | 0. |
| 2.67 | +/- | -NaN |
| -0.65 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.31 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.67 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
357-03-O
| 5825. | +/- | 66. |
| -10000. | +/- | -NaN |
| 2.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.79 | +/- | 1. |
| 4.79 | +/- | -NaN |
| -1.82 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.24 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.11 | +/- | 7. |
| -9999.99 | +/- | -NaN |
| 0.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.58 | +/- | 0. |
| 0.93 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -1.98 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -2.28 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
357-03-O
| 5631. | +/- | 104. |
| -10000. | +/- | -NaN |
| 1.92 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.45 | +/- | 1. |
| 0.45 | +/- | -NaN |
| -1.91 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.00 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.93 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 9.06 | +/- | 0. |
| 0.96 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.49 |
| -9999.00 |

BRIGHT_NEIGHBOR
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
357-03-O
| 4449. | +/- | 53. |
| -10000. | +/- | -NaN |
| 2.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| -0.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.66 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.27 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
357-03-O
| 5608. | +/- | 125. |
| -10000. | +/- | -NaN |
| 3.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| -0.56 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.11 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| 0.39 |
| -9999.00 |

VERY_BRIGHT_NEIGHBOR
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
357-03-O
| 6003. | +/- | 34. |
| -10000. | +/- | -NaN |
| 2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.54 | +/- | 0. |
| 3.54 | +/- | -NaN |
| -1.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| 0.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 21.71 | +/- | 0. |
| 1.34 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.17 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.01 |
| -9999.00 |
VERY_BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 7938. | +/- | 106. |
| -10000. | +/- | -NaN |
| 3.63 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.46 | +/- | 1. |
| 1.46 | +/- | -NaN |
| -1.31 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.31 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.30 | +/- | 1. |
| 0.72 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.32 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 4301. | +/- | 66. |
| -10000. | +/- | -NaN |
| 2.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| -1.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.23 | +/- | 0. |
| 0.79 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| 0.98 |
| -9999.00 |

STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6056. | +/- | 85. |
| -10000. | +/- | -NaN |
| 3.22 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.67 | +/- | 0. |
| 1.67 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.27 | +/- | 0. |
| 0.80 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.52 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
357-03-O
| 5601. | +/- | 148. |
| -10000. | +/- | -NaN |
| 2.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.32 | +/- | 1. |
| 3.32 | +/- | -NaN |
| -2.46 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.80 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| -0.13 | +/- | 19. |
| -9999.99 | +/- | -NaN |
| 0.58 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 12.46 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.85 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.47 |
| -9999.00 |
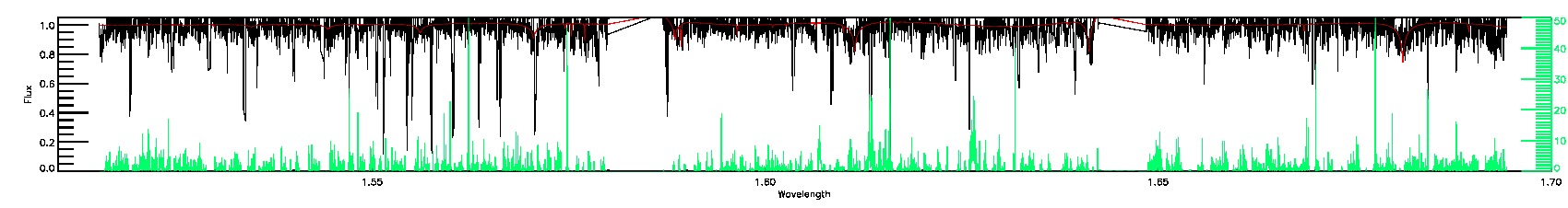
PERSIST_JUMP_NEG
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6247. | +/- | 102. |
| -10000. | +/- | -NaN |
| 2.73 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.12 | +/- | 1. |
| 1.12 | +/- | -NaN |
| -1.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| -0.70 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.03 | +/- | 1. |
| 0.70 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.35 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 5884. | +/- | 111. |
| -10000. | +/- | -NaN |
| 2.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.74 | +/- | 2. |
| 0.74 | +/- | -NaN |
| -1.88 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| -0.35 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.90 | +/- | 0. |
| 0.95 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| -2.43 |
| -9999.00 |
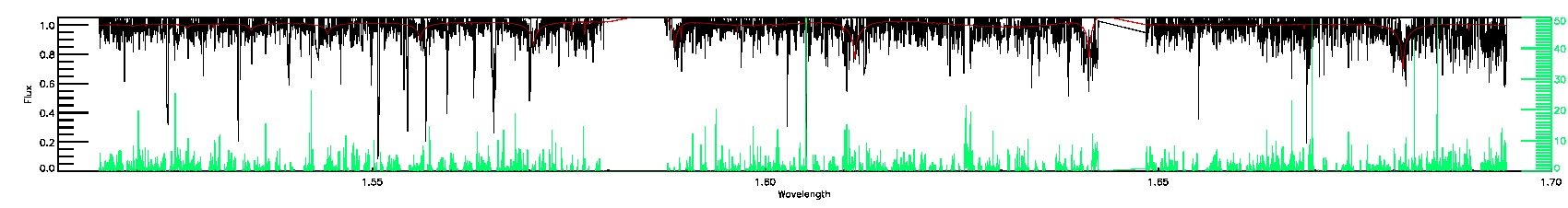
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 3720. | +/- | 23. |
| -10000. | +/- | -NaN |
| 0.69 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| -1.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.72 | +/- | 0. |
| 0.83 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -1.83 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 7041. | +/- | 158. |
| -10000. | +/- | -NaN |
| 3.57 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.62 | +/- | 1. |
| 0.62 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.77 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 78.63 | +/- | 0. |
| 1.90 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 1.00 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
357-03-O
| 5480. | +/- | 95. |
| -10000. | +/- | -NaN |
| 1.67 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.60 | +/- | 0. |
| 4.60 | +/- | -NaN |
| -1.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.29 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.54 | +/- | 0. |
| 0.82 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.60 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -2.50 |
| -9999.00 |

BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6164. | +/- | 84. |
| -10000. | +/- | -NaN |
| 2.60 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.64 | +/- | 0. |
| 1.64 | +/- | -NaN |
| 0.61 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.62 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 83.68 | +/- | 0. |
| 1.92 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.42 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 1.00 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6287. | +/- | 91. |
| -10000. | +/- | -NaN |
| 2.77 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.58 | +/- | 0. |
| 4.58 | +/- | -NaN |
| -1.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 14.08 | +/- | 0. |
| 1.15 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.67 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6838. | +/- | 90. |
| -10000. | +/- | -NaN |
| 2.92 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.82 | +/- | 0. |
| 1.82 | +/- | -NaN |
| -1.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 13. |
| -9999.99 | +/- | -NaN |
| 0.95 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 16.70 | +/- | 0. |
| 1.22 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 1.00 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 5700. | +/- | 85. |
| -10000. | +/- | -NaN |
| 2.56 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.10 | +/- | 0. |
| 4.10 | +/- | -NaN |
| -1.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.45 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 21.52 | +/- | 0. |
| 1.33 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.37 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -2.29 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
357-03-O
| 6230. | +/- | 29. |
| -10000. | +/- | -NaN |
| 2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.69 | +/- | 2. |
| 4.69 | +/- | -NaN |
| -1.78 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 18. |
| -9999.99 | +/- | -NaN |
| 0.37 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 14.49 | +/- | 1. |
| 1.16 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.64 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
357-03-O
| 6154. | +/- | 39. |
| -10000. | +/- | -NaN |
| 2.51 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.52 | +/- | 1. |
| 1.52 | +/- | -NaN |
| -1.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.27 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| 0.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 76.33 | +/- | 0. |
| 1.88 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.90 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
357-03-O
| 6313. | +/- | 94. |
| -10000. | +/- | -NaN |
| 3.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.00 | +/- | 0. |
| 1.00 | +/- | -NaN |
| -1.42 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.21 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.36 | +/- | 11. |
| -9999.99 | +/- | -NaN |
| 0.95 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 11.62 | +/- | 0. |
| 1.07 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.16 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -1.92 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6917. | +/- | 106. |
| -10000. | +/- | -NaN |
| 3.41 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.26 | +/- | 1. |
| 1.26 | +/- | -NaN |
| -1.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 | +/- | 6. |
| -9999.99 | +/- | -NaN |
| 0.26 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.41 | +/- | 0. |
| 0.81 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.79 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 5194. | +/- | 104. |
| -10000. | +/- | -NaN |
| 2.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.48 | +/- | 1. |
| 0.48 | +/- | -NaN |
| -0.58 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.14 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 1.41 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| 0.99 |
| -9999.00 |
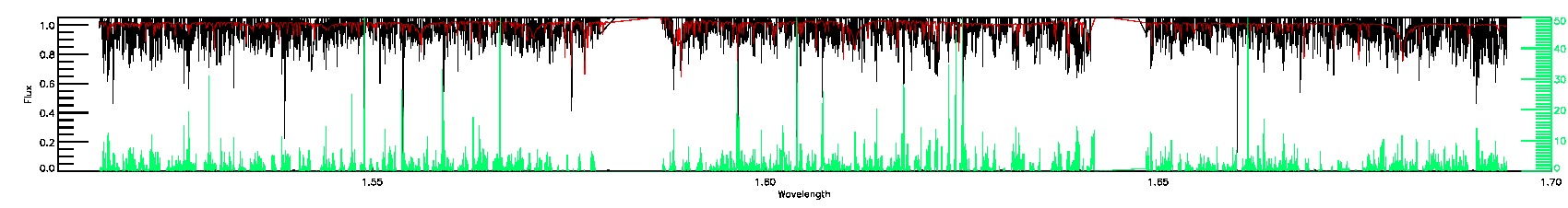
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 5975. | +/- | 88. |
| -10000. | +/- | -NaN |
| 2.62 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.64 | +/- | 0. |
| 4.64 | +/- | -NaN |
| -1.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.49 | +/- | 0. |
| 0.93 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.45 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
357-03-O
| 7724. | +/- | 108. |
| -10000. | +/- | -NaN |
| 3.64 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.09 | +/- | 1. |
| 1.09 | +/- | -NaN |
| -1.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.97 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 19.74 | +/- | 0. |
| 1.30 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.99 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6177. | +/- | 97. |
| -10000. | +/- | -NaN |
| 2.69 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.12 | +/- | 0. |
| 4.12 | +/- | -NaN |
| -1.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| 0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.48 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.49 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6565. | +/- | 310. |
| -10000. | +/- | -NaN |
| 4.34 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.69 | +/- | 1. |
| 1.69 | +/- | -NaN |
| -0.64 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 88.86 | +/- | 0. |
| 1.95 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 5860. | +/- | 103. |
| -10000. | +/- | -NaN |
| 2.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.61 | +/- | 1. |
| 4.61 | +/- | -NaN |
| -1.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.34 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 7. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.57 | +/- | 0. |
| 0.82 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -1.69 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.39 |
| -9999.00 |
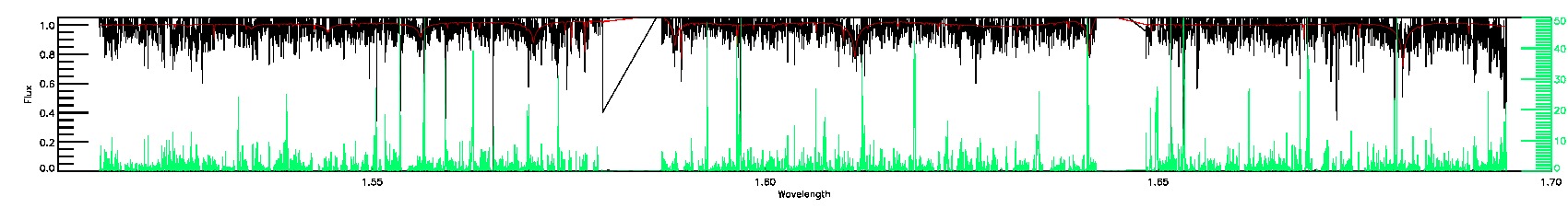
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6008. | +/- | 128. |
| -10000. | +/- | -NaN |
| 2.75 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.52 | +/- | 0. |
| 4.52 | +/- | -NaN |
| -1.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 9. |
| -9999.99 | +/- | -NaN |
| 0.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 14.98 | +/- | 0. |
| 1.18 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.08 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 5950. | +/- | 87. |
| -10000. | +/- | -NaN |
| 2.58 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.46 | +/- | 0. |
| 3.46 | +/- | -NaN |
| -1.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 6. |
| -9999.99 | +/- | -NaN |
| 0.42 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.41 | +/- | 0. |
| 0.87 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.75 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
357-03-O
| 5688. | +/- | 135. |
| -10000. | +/- | -NaN |
| 2.30 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.92 | +/- | 1. |
| 0.92 | +/- | -NaN |
| -1.39 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.31 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.68 | +/- | 0. |
| 0.82 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -2.47 |
| -9999.00 |

BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
357-03-O
| 5508. | +/- | 67. |
| -10000. | +/- | -NaN |
| 1.74 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.57 | +/- | 2. |
| 1.57 | +/- | -NaN |
| -2.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.11 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.61 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 12.69 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.35 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.49 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
357-03-O
| 7156. | +/- | 100. |
| -10000. | +/- | -NaN |
| 3.31 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.77 | +/- | 0. |
| 4.77 | +/- | -NaN |
| -1.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.42 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.35 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 20.49 | +/- | 0. |
| 1.31 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.37 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 1.35 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.95 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 7562. | +/- | 162. |
| -10000. | +/- | -NaN |
| 3.65 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.49 | +/- | 0. |
| 2.49 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.21 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| -0.61 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.87 | +/- | 1. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.60 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
357-03-O
| 7045. | +/- | 84. |
| -10000. | +/- | -NaN |
| 3.42 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.72 | +/- | 1. |
| 0.72 | +/- | -NaN |
| -0.86 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| 0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.06 | +/- | 0. |
| 0.31 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| -0.99 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
357-03-O
| 6871. | +/- | 92. |
| -10000. | +/- | -NaN |
| 2.95 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.48 | +/- | 0. |
| 4.48 | +/- | -NaN |
| -0.79 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 17.24 | +/- | 0. |
| 1.24 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.27 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.89 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 7337. | +/- | 120. |
| -10000. | +/- | -NaN |
| 3.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.19 | +/- | 1. |
| 1.19 | +/- | -NaN |
| -1.34 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 11. |
| -9999.99 | +/- | -NaN |
| 0.32 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 1.70 | +/- | 1. |
| 0.23 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.48 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.44 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6217. | +/- | 47. |
| -10000. | +/- | -NaN |
| 2.52 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.64 | +/- | 0. |
| 4.64 | +/- | -NaN |
| -1.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 7. |
| -9999.99 | +/- | -NaN |
| 0.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.16 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 0.80 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6504. | +/- | 41. |
| -10000. | +/- | -NaN |
| 2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.90 | +/- | 1. |
| 0.90 | +/- | -NaN |
| -1.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.25 | +/- | 13. |
| -9999.99 | +/- | -NaN |
| 0.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 28.22 | +/- | 0. |
| 1.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.48 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 1.00 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 4794. | +/- | 88. |
| -10000. | +/- | -NaN |
| 1.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.36 | +/- | 0. |
| 1.36 | +/- | -NaN |
| -1.85 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.22 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.44 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.69 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.71 | +/- | 0. |
| 0.94 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -1.27 |
| -9999.00 |
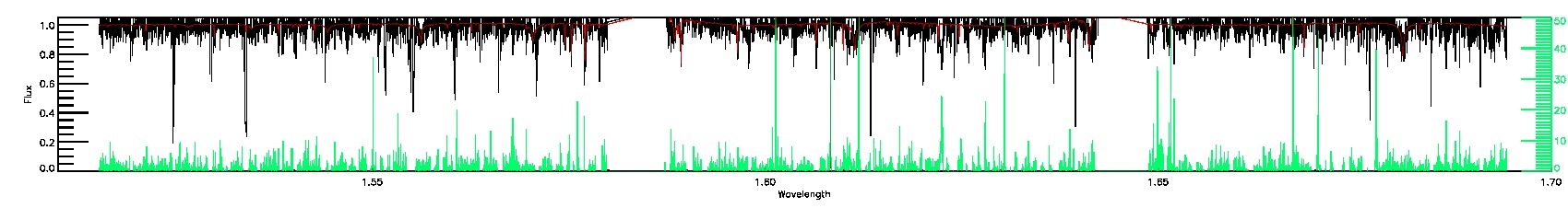
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
357-03-O
| 5887. | +/- | 128. |
| -10000. | +/- | -NaN |
| 2.66 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.11 | +/- | 0. |
| 1.11 | +/- | -NaN |
| -1.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.39 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.19 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.56 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.51 | +/- | 0. |
| 0.74 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.90 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |

BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
357-03-O
| 5950. | +/- | 88. |
| -10000. | +/- | -NaN |
| 2.34 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.72 | +/- | 0. |
| 4.72 | +/- | -NaN |
| -1.26 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.42 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.17 | +/- | 0. |
| 0.79 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.28 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.79 |
| -9999.00 |

BRIGHT_NEIGHBOR
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 4517. | +/- | 90. |
| -10000. | +/- | -NaN |
| 0.85 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.65 | +/- | 1. |
| 0.65 | +/- | -NaN |
| -2.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.92 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.69 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 11.01 | +/- | 0. |
| 1.04 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -1.69 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -2.28 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6999. | +/- | 108. |
| -10000. | +/- | -NaN |
| 2.59 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.33 | +/- | 1. |
| 1.33 | +/- | -NaN |
| -1.34 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 0.67 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 9.94 | +/- | 0. |
| 1.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| -1.19 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
357-03-O
| 5078. | +/- | 105. |
| -10000. | +/- | -NaN |
| 1.46 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.76 | +/- | 0. |
| 0.76 | +/- | -NaN |
| -1.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.42 | +/- | 0. |
| 0.73 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.10 |
| -9999.00 |

STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6702. | +/- | 45. |
| -10000. | +/- | -NaN |
| 2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.90 | +/- | 1. |
| 0.90 | +/- | -NaN |
| -1.17 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.14 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 15.81 | +/- | 1. |
| 1.20 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.33 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6823. | +/- | 125. |
| -10000. | +/- | -NaN |
| 3.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.29 | +/- | 0. |
| 2.29 | +/- | -NaN |
| -1.95 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 20. |
| -9999.99 | +/- | -NaN |
| 0.97 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 15.65 | +/- | 0. |
| 1.19 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.34 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.95 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 5951. | +/- | 93. |
| -10000. | +/- | -NaN |
| 2.34 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.58 | +/- | 1. |
| 4.58 | +/- | -NaN |
| -1.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.23 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 6. |
| -9999.99 | +/- | -NaN |
| 0.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.64 | +/- | 0. |
| 0.82 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.30 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.32 |
| -9999.00 |

VERY_BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
357-03-O
| 4841. | +/- | 89. |
| -10000. | +/- | -NaN |
| 1.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.84 | +/- | 1. |
| 0.84 | +/- | -NaN |
| -2.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.43 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.77 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.18 | +/- | 0. |
| 1.01 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.61 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.43 |
| -9999.00 |
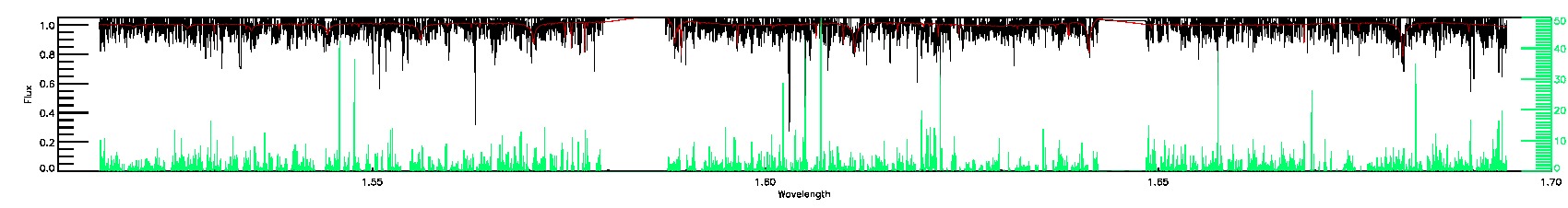
SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6823. | +/- | 95. |
| -10000. | +/- | -NaN |
| 3.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.75 | +/- | 0. |
| 3.75 | +/- | -NaN |
| -1.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.96 | +/- | 1. |
| 0.29 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.80 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 5686. | +/- | 53. |
| -10000. | +/- | -NaN |
| 2.61 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.15 | +/- | 0. |
| 1.15 | +/- | -NaN |
| -0.83 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 24.93 | +/- | 0. |
| 1.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6858. | +/- | 55. |
| -10000. | +/- | -NaN |
| 2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.05 | +/- | 0. |
| 2.05 | +/- | -NaN |
| -0.77 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 13.00 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.40 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6471. | +/- | 67. |
| -10000. | +/- | -NaN |
| 2.62 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.51 | +/- | 0. |
| 4.51 | +/- | -NaN |
| -1.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 7. |
| -9999.99 | +/- | -NaN |
| 0.39 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.37 | +/- | 0. |
| 1.02 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -2.09 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
357-03-O
| 6726. | +/- | 88. |
| -10000. | +/- | -NaN |
| 2.99 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.11 | +/- | 3. |
| 2.11 | +/- | -NaN |
| -2.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 8. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.89 | +/- | 1. |
| 0.28 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -2.38 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 5270. | +/- | 169. |
| -10000. | +/- | -NaN |
| 3.31 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.27 | +/- | 0. |
| 1.27 | +/- | -NaN |
| -0.57 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.13 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.98 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -2.44 |
| -9999.00 |
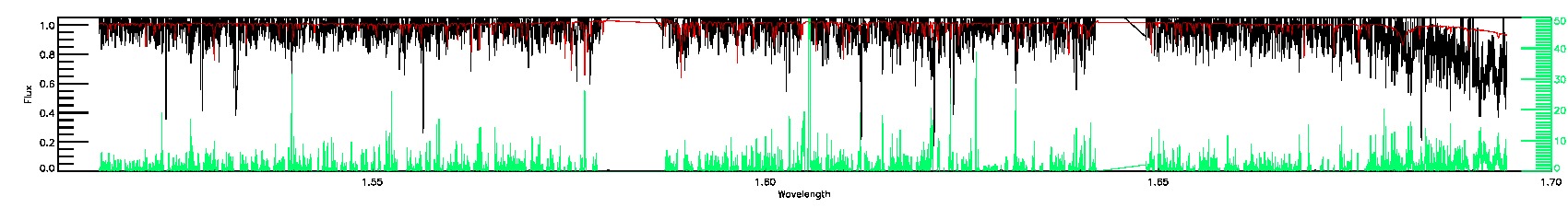
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 5392. | +/- | 122. |
| -10000. | +/- | -NaN |
| 1.85 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.55 | +/- | 1. |
| 4.55 | +/- | -NaN |
| -1.98 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.49 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| -0.15 | +/- | 7. |
| -9999.99 | +/- | -NaN |
| 0.72 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 9.40 | +/- | 0. |
| 0.97 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -1.98 |
| -9999.00 |
| -1.98 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.01 |
| -9999.00 |

BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
357-03-O
| 6076. | +/- | 92. |
| -10000. | +/- | -NaN |
| 2.59 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.38 | +/- | 0. |
| 1.38 | +/- | -NaN |
| -1.60 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.82 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 17.35 | +/- | 0. |
| 1.24 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -2.35 |
| -9999.00 |
