SUSPECT_BROAD_LINES
STAR_WARN,COLORTE_WARN
359-02-O
| 8912. | +/- | 61. |
| 8807. | +/- | 404. |
| 4.25 | +/- | 0. |
| 4.76 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.26 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.14 | +/- | 1. |
| 0.79 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.28 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -1.98 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -1.95 |
| -9999.00 |
STAR_WARN,SN_WARN
359-02-O
| 4341. | +/- | 16. |
| 4442. | +/- | 111. |
| 2.17 | +/- | 0. |
| 1.97 | +/- | 0. |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -0.28 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.14 | +/- | 0. |
| 0.14 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -0.11 | +/- | 0. |
| 3.48 | +/- | 0. |
| 0.54 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -0.59 |
| -9999.00 |

SUSPECT_BROAD_LINES
359-02-O
| 7301. | +/- | 24. |
| 7096. | +/- | 218. |
| 4.25 | +/- | 0. |
| 3.83 | +/- | 0. |
| 2.99 | +/- | 0. |
| 2.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| 0.09 | +/- | 0. |
| -0.14 | +/- | 0. |
| -0.14 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -0.31 | +/- | -NaN |
| 0.03 | +/- | 0. |
| 0.02 | +/- | 0. |
| 27.19 | +/- | 0. |
| 1.43 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.50 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 1.00 |
| -9999.00 |
STAR_WARN,SN_WARN
359-02-O
| 4317. | +/- | 12. |
| 4421. | +/- | 112. |
| 2.28 | +/- | 0. |
| 2.09 | +/- | 0. |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -0.36 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.16 | +/- | 0. |
| 3.66 | +/- | 0. |
| 0.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -0.25 |
| -9999.00 |

STAR_WARN,SN_WARN
359-02-O
| 4324. | +/- | 14. |
| 4442. | +/- | 116. |
| 1.74 | +/- | 0. |
| 1.61 | +/- | 0. |
| 1.07 | +/- | 0. |
| 1.07 | +/- | -NaN |
| -0.69 | +/- | 0. |
| -0.69 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.11 | +/- | 0. |
| 0.11 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.21 | +/- | 0. |
| 4.44 | +/- | 0. |
| 0.65 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -1.65 |
| -9999.00 |
STAR_WARN,SN_WARN
359-02-O
| 4322. | +/- | 14. |
| 4421. | +/- | 110. |
| 2.11 | +/- | 0. |
| 1.91 | +/- | 0. |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -0.25 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.12 | +/- | 0. |
| 0.12 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -0.05 | +/- | 0. |
| 3.42 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -0.11 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
359-02-O
| 8698. | +/- | 40. |
| 8590. | +/- | 353. |
| 3.87 | +/- | 0. |
| 4.34 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 28.55 | +/- | 0. |
| 1.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.33 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -2.13 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
359-02-O
| 7851. | +/- | 14. |
| 7656. | +/- | 223. |
| 4.32 | +/- | 0. |
| 3.96 | +/- | 0. |
| 1.27 | +/- | 0. |
| 1.27 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| 0.05 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.04 | +/- | -NaN |
| 0.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 87.02 | +/- | 0. |
| 1.94 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -2.31 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 7558. | +/- | 87. |
| -10000. | +/- | -NaN |
| 3.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.90 | +/- | 0. |
| 3.90 | +/- | -NaN |
| -1.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.77 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 22.45 | +/- | 0. |
| 1.35 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 1.47 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.02 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
359-02-O
| 3900. | +/- | 2. |
| 3991. | +/- | 86. |
| 1.63 | +/- | 0. |
| 1.42 | +/- | 0. |
| 1.39 | +/- | 0. |
| 1.39 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -0.05 | +/- | 0. |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.22 | +/- | 0. |
| 0.19 | +/- | 0. |
| 3.04 | +/- | 0. |
| 0.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 4376. | +/- | 20. |
| -10000. | +/- | -NaN |
| 2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.39 | +/- | 0. |
| 0.53 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.92 |
| -9999.00 |

359-02-O
| 4007. | +/- | 2. |
| 4079. | +/- | 82. |
| 2.08 | +/- | 0. |
| 1.79 | +/- | 0. |
| 0.84 | +/- | 0. |
| 0.84 | +/- | -NaN |
| 0.38 | +/- | 0. |
| 0.38 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.03 | +/- | 0. |
| 2.36 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.95 |
| -9999.00 |

STAR_WARN,COLORTE_WARN
359-02-O
| 8647. | +/- | 50. |
| 8537. | +/- | 378. |
| 4.27 | +/- | 0. |
| 4.72 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 27.55 | +/- | 0. |
| 1.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.05 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -1.97 |
| -9999.00 |
STAR_WARN,SN_WARN
359-02-O
| 3881. | +/- | 3. |
| 3967. | +/- | 89. |
| 1.34 | +/- | 0. |
| 1.17 | +/- | 0. |
| 1.36 | +/- | 0. |
| 1.36 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.03 | +/- | 0. |
| 0.03 | +/- | -NaN |
| 0.23 | +/- | 0. |
| 0.23 | +/- | -NaN |
| 0.06 | +/- | 0. |
| 0.03 | +/- | 0. |
| 2.84 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -0.17 |
| -9999.00 |
STAR_WARN,COLORTE_WARN,SN_WARN
359-02-O
| 4415. | +/- | 10. |
| 4482. | +/- | 103. |
| 2.76 | +/- | 0. |
| 2.43 | +/- | 0. |
| 0.83 | +/- | 0. |
| 0.83 | +/- | -NaN |
| 0.49 | +/- | 0. |
| 0.49 | +/- | 0. |
| 0.10 | +/- | 0. |
| 0.10 | +/- | -NaN |
| 0.28 | +/- | 0. |
| 0.28 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.22 | +/- | 0. |
| 0.35 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| 0.97 |
| -9999.00 |

359-02-O
| 3837. | +/- | 2. |
| 3906. | +/- | 79. |
| 1.76 | +/- | 0. |
| 1.47 | +/- | 0. |
| 1.22 | +/- | 0. |
| 1.22 | +/- | -NaN |
| 0.44 | +/- | 0. |
| 0.44 | +/- | 0. |
| 0.06 | +/- | 0. |
| 0.06 | +/- | -NaN |
| 0.26 | +/- | 0. |
| 0.26 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -0.02 | +/- | 0. |
| 2.28 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.74 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 4733. | +/- | 47. |
| -10000. | +/- | -NaN |
| 2.52 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.53 | +/- | 0. |
| 0.53 | +/- | -NaN |
| -0.64 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.31 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| -1.90 |
| -9999.00 |

STAR_WARN,SN_WARN
359-02-O
| 4714. | +/- | 38. |
| 4795. | +/- | 126. |
| 2.80 | +/- | 0. |
| 2.64 | +/- | 0. |
| 0.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| -0.41 | +/- | 0. |
| -0.41 | +/- | 0. |
| -0.02 | +/- | 0. |
| -0.02 | +/- | -NaN |
| 0.19 | +/- | 0. |
| 0.19 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.16 | +/- | 0. |
| 3.76 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| 1.00 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_WARN,COLORTE_WARN
359-02-O
| 9121. | +/- | 86. |
| 8998. | +/- | 434. |
| 4.01 | +/- | 0. |
| 4.34 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -1.83 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 15.51 | +/- | 0. |
| 1.19 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.87 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -1.65 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_WARN,SN_WARN
359-02-O
| 4433. | +/- | 16. |
| 4518. | +/- | 110. |
| 2.55 | +/- | 0. |
| 2.30 | +/- | 0. |
| 0.50 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.07 | +/- | 0. |
| 0.07 | +/- | 0. |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| 0.08 | +/- | 0. |
| 0.08 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -0.04 | +/- | 0. |
| 2.83 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.43 |
| -9999.00 |

359-02-O
| 8316. | +/- | 52. |
| 8192. | +/- | 371. |
| 4.23 | +/- | 0. |
| 4.56 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -1.83 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 29.70 | +/- | 0. |
| 1.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.42 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.98 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| -1.81 |
| -9999.00 |
STAR_BAD
STAR_WARN,COLORTE_WARN
359-02-O
| 6932. | +/- | 31. |
| -10000. | +/- | -NaN |
| 4.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.45 | +/- | 0. |
| 1.45 | +/- | -NaN |
| -2.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 94.99 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.16 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.40 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
359-02-O
| 15248. | +/- | 112. |
| -10000. | +/- | -NaN |
| 4.43 | +/- | 0. |
| 3.89 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| 0.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 81.49 | +/- | 0. |
| 1.91 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.45 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.14 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
359-02-O
| 8522. | +/- | 41. |
| 8414. | +/- | 335. |
| 3.80 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -2.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 56.55 | +/- | 0. |
| 1.75 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -2.34 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -2.14 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| -2.14 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -2.09 |
| -9999.00 |
STAR_BAD
STAR_WARN,COLORTE_WARN
359-02-O
| 8919. | +/- | 54. |
| -10000. | +/- | -NaN |
| 3.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -1.62 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.35 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.61 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -1.69 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -1.69 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -1.66 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
359-02-O
| 8239. | +/- | 24. |
| 8041. | +/- | 241. |
| 3.67 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 47.83 | +/- | 0. |
| 1.68 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.14 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -0.24 |
| -9999.00 |
STAR_BAD,COLORTE_BAD
STAR_WARN,COLORTE_WARN
359-02-O
| 5291. | +/- | 25. |
| -10000. | +/- | -NaN |
| 2.96 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.28 | +/- | 1. |
| 1.28 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.34 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.75 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |

STAR_WARN,SN_WARN
359-02-O
| 4934. | +/- | 54. |
| 5007. | +/- | 142. |
| 2.77 | +/- | 0. |
| 2.67 | +/- | 0. |
| 1.17 | +/- | 0. |
| 1.17 | +/- | -NaN |
| -0.83 | +/- | 0. |
| -0.83 | +/- | 0. |
| 0.09 | +/- | 0. |
| 0.09 | +/- | -NaN |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.30 | +/- | 0. |
| 4.81 | +/- | 0. |
| 0.68 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -2.14 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -1.18 |
| -9999.00 |
STAR_WARN,COLORTE_WARN
359-02-O
| 13772. | +/- | 99. |
| -10000. | +/- | -NaN |
| 4.36 | +/- | 0. |
| 3.80 | +/- | 0. |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| 0.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| 0.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 86.22 | +/- | 0. |
| 1.94 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.33 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| 0.15 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 4778. | +/- | 43. |
| -10000. | +/- | -NaN |
| 2.94 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.55 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.19 | +/- | 0. |
| 0.50 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.17 |
| -9999.00 |

BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 5986. | +/- | 66. |
| -10000. | +/- | -NaN |
| 2.69 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.77 | +/- | 0. |
| 4.77 | +/- | -NaN |
| -1.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.39 | +/- | 0. |
| 1.02 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.42 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -1.93 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 7351. | +/- | 80. |
| -10000. | +/- | -NaN |
| 3.67 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.52 | +/- | 0. |
| 3.52 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 15.09 | +/- | 0. |
| 1.18 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.30 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -1.99 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 3638. | +/- | 24. |
| -10000. | +/- | -NaN |
| 0.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.41 | +/- | 0. |
| 0.41 | +/- | -NaN |
| -1.90 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.98 | +/- | 0. |
| 0.95 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -1.69 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| -1.93 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -2.42 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,SN_WARN
359-02-O
| 7122. | +/- | 31. |
| -10000. | +/- | -NaN |
| 3.84 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.79 | +/- | 0. |
| 4.79 | +/- | -NaN |
| 0.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 13.40 | +/- | 0. |
| 1.13 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.50 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.14 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.20 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 3812. | +/- | 43. |
| -10000. | +/- | -NaN |
| 2.99 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| -1.96 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.21 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.41 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 9.31 | +/- | 0. |
| 0.97 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -1.98 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -2.47 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5603. | +/- | 109. |
| -10000. | +/- | -NaN |
| 2.61 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.52 | +/- | 0. |
| 1.52 | +/- | -NaN |
| -1.21 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.22 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.55 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 15.58 | +/- | 0. |
| 1.19 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.50 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.69 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6452. | +/- | 133. |
| -10000. | +/- | -NaN |
| 2.67 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.99 | +/- | 1. |
| 0.99 | +/- | -NaN |
| -1.91 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 11. |
| -9999.99 | +/- | -NaN |
| 0.56 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 88.84 | +/- | 0. |
| 1.95 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.75 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7966. | +/- | 252. |
| -10000. | +/- | -NaN |
| 3.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.65 | +/- | 1. |
| 0.65 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 11.89 | +/- | 0. |
| 1.08 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -2.43 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5730. | +/- | 47. |
| -10000. | +/- | -NaN |
| 3.83 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.66 | +/- | 1. |
| 0.66 | +/- | -NaN |
| -0.30 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.46 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.53 | +/- | 0. |
| 0.74 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.42 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| -2.43 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7779. | +/- | 160. |
| -10000. | +/- | -NaN |
| 3.92 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.77 | +/- | 1. |
| 0.77 | +/- | -NaN |
| 0.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.65 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 88.37 | +/- | 0. |
| 1.95 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6274. | +/- | 116. |
| -10000. | +/- | -NaN |
| 3.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.70 | +/- | 0. |
| 0.70 | +/- | -NaN |
| 0.88 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.61 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.48 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 1.00 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7963. | +/- | 296. |
| -10000. | +/- | -NaN |
| 5.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.56 | +/- | 7. |
| 1.56 | +/- | -NaN |
| -1.66 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| -0.27 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 4.55 | +/- | 1. |
| 0.66 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| 0.34 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 5891. | +/- | 134. |
| -10000. | +/- | -NaN |
| 2.32 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.54 | +/- | 0. |
| 2.54 | +/- | -NaN |
| -1.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.35 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.25 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.92 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.00 | +/- | 0. |
| 0.84 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.80 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.48 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 10280. | +/- | 415. |
| -10000. | +/- | -NaN |
| 3.42 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -1.00 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 11.81 | +/- | 3. |
| 1.07 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.16 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -1.31 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 5786. | +/- | 112. |
| -10000. | +/- | -NaN |
| 2.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.56 | +/- | 1. |
| 0.56 | +/- | -NaN |
| -1.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.34 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.20 | +/- | 8. |
| -9999.99 | +/- | -NaN |
| 0.65 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.75 | +/- | 0. |
| 0.76 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.37 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6414. | +/- | 90. |
| -10000. | +/- | -NaN |
| 2.65 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.40 | +/- | 1. |
| 0.40 | +/- | -NaN |
| -0.76 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.42 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| -0.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 28.11 | +/- | 0. |
| 1.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.45 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -2.16 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5824. | +/- | 146. |
| -10000. | +/- | -NaN |
| 2.45 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.59 | +/- | 1. |
| 1.59 | +/- | -NaN |
| -1.51 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 11. |
| -9999.99 | +/- | -NaN |
| 0.72 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.16 | +/- | 0. |
| 0.85 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.90 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.04 |
| -9999.00 |

BRIGHT_NEIGHBOR
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5838. | +/- | 130. |
| -10000. | +/- | -NaN |
| 2.90 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.45 | +/- | 1. |
| 0.45 | +/- | -NaN |
| -0.96 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 14.84 | +/- | 0. |
| 1.17 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.50 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -2.20 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7128. | +/- | 125. |
| -10000. | +/- | -NaN |
| 3.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.63 | +/- | 0. |
| 0.63 | +/- | -NaN |
| 0.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 18.42 | +/- | 0. |
| 1.27 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.48 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.09 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 9986. | +/- | 1698. |
| -10000. | +/- | -NaN |
| 4.31 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -0.67 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 16.32 | +/- | 1. |
| 1.21 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.35 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.69 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6438. | +/- | 43. |
| -10000. | +/- | -NaN |
| 2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.67 | +/- | 1. |
| 0.67 | +/- | -NaN |
| -1.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 6. |
| -9999.99 | +/- | -NaN |
| 0.34 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 30.31 | +/- | 0. |
| 1.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| 0.99 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6680. | +/- | 154. |
| -10000. | +/- | -NaN |
| 3.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| 0.34 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 93.82 | +/- | 0. |
| 1.97 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.50 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -2.35 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 5520. | +/- | 128. |
| -10000. | +/- | -NaN |
| 2.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.30 | +/- | 1. |
| 1.30 | +/- | -NaN |
| -1.73 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.37 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.14 | +/- | 0. |
| 0.91 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.20 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.24 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7940. | +/- | 193. |
| -10000. | +/- | -NaN |
| 3.85 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.33 | +/- | 0. |
| 1.33 | +/- | -NaN |
| 0.61 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.46 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 33.08 | +/- | 0. |
| 1.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6160. | +/- | 79. |
| -10000. | +/- | -NaN |
| 2.52 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.06 | +/- | 0. |
| 2.06 | +/- | -NaN |
| -0.56 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.18 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.30 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.67 | +/- | 0. |
| 0.94 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -0.75 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6089. | +/- | 108. |
| -10000. | +/- | -NaN |
| 2.72 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.81 | +/- | 0. |
| 1.81 | +/- | -NaN |
| -0.67 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.60 | +/- | 0. |
| 0.93 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 1.50 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 1.00 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 4526. | +/- | 82. |
| -10000. | +/- | -NaN |
| 2.21 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -1.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.41 | +/- | 0. |
| 0.73 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| -2.39 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 3848. | +/- | 27. |
| -10000. | +/- | -NaN |
| 0.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -1.73 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.11 | +/- | 0. |
| 0.91 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -2.49 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 3788. | +/- | 28. |
| -10000. | +/- | -NaN |
| 0.83 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| -1.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.57 | +/- | 0. |
| 0.82 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -1.76 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7118. | +/- | 120. |
| -10000. | +/- | -NaN |
| 3.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.65 | +/- | 0. |
| 2.65 | +/- | -NaN |
| -0.45 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.77 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 16.44 | +/- | 0. |
| 1.22 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| 0.49 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 4625. | +/- | 403. |
| -10000. | +/- | -NaN |
| 2.63 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 1.54 | +/- | 2. |
| 1.54 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.34 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 12.76 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.22 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 1.16 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -9999.00 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7332. | +/- | 8. |
| -10000. | +/- | -NaN |
| 2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.25 | +/- | 0. |
| 1.25 | +/- | -NaN |
| 0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.60 | +/- | 0. |
| 0.75 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.50 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| 1.00 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6776. | +/- | 125. |
| -10000. | +/- | -NaN |
| 3.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.75 | +/- | 1. |
| 0.75 | +/- | -NaN |
| -1.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.94 | +/- | 1. |
| 0.69 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -1.82 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6038. | +/- | 83. |
| -10000. | +/- | -NaN |
| 2.74 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.17 | +/- | 0. |
| 1.17 | +/- | -NaN |
| -0.71 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 16.01 | +/- | 0. |
| 1.20 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 1.00 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5879. | +/- | 118. |
| -10000. | +/- | -NaN |
| 2.72 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.29 | +/- | 0. |
| 1.29 | +/- | -NaN |
| -1.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 | +/- | 7. |
| -9999.99 | +/- | -NaN |
| 0.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 23.87 | +/- | 0. |
| 1.38 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.37 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5738. | +/- | 69. |
| -10000. | +/- | -NaN |
| 2.75 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.23 | +/- | 0. |
| 1.23 | +/- | -NaN |
| -0.74 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.10 | +/- | 0. |
| 1.08 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 1.50 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 1.00 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6480. | +/- | 111. |
| -10000. | +/- | -NaN |
| 2.65 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.84 | +/- | 0. |
| 1.84 | +/- | -NaN |
| -1.35 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 36. |
| -9999.99 | +/- | -NaN |
| 0.95 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 29.10 | +/- | 0. |
| 1.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.47 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6177. | +/- | 107. |
| -10000. | +/- | -NaN |
| 2.62 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.07 | +/- | 1. |
| 1.07 | +/- | -NaN |
| -1.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.36 | +/- | 0. |
| 0.92 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.35 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7100. | +/- | 181. |
| -10000. | +/- | -NaN |
| 3.89 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.47 | +/- | 1. |
| 1.47 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.97 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 43.51 | +/- | 0. |
| 1.64 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.40 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7354. | +/- | 165. |
| -10000. | +/- | -NaN |
| 3.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.76 | +/- | 0. |
| 4.76 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.42 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 33.39 | +/- | 0. |
| 1.52 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.22 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -2.34 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 3166. | +/- | 1. |
| -10000. | +/- | -NaN |
| 0.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| 0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.77 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 1.22 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.11 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 3004. | +/- | 2. |
| -10000. | +/- | -NaN |
| 2.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.81 | +/- | 0. |
| 0.81 | +/- | -NaN |
| -0.80 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.72 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.72 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| 1.99 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5501. | +/- | 1. |
| -10000. | +/- | -NaN |
| 3.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.11 | +/- | 0. |
| 1.11 | +/- | -NaN |
| -1.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.46 | +/- | 0. |
| 0.87 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -1.23 |
| -9999.00 |
PERSIST_JUMP_NEG
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 3647. | +/- | 21. |
| -10000. | +/- | -NaN |
| -0.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.51 | +/- | 0. |
| 0.51 | +/- | -NaN |
| -0.46 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.52 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.64 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.52 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.87 | +/- | 0. |
| 0.59 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.31 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| 1.52 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 4201. | +/- | 12. |
| -10000. | +/- | -NaN |
| 2.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| 0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.74 | +/- | 0. |
| 0.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.62 |
| -9999.00 |

STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 4693. | +/- | 110. |
| -10000. | +/- | -NaN |
| 2.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| -0.68 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.41 | +/- | 0. |
| 0.64 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -2.11 |
| -9999.00 |
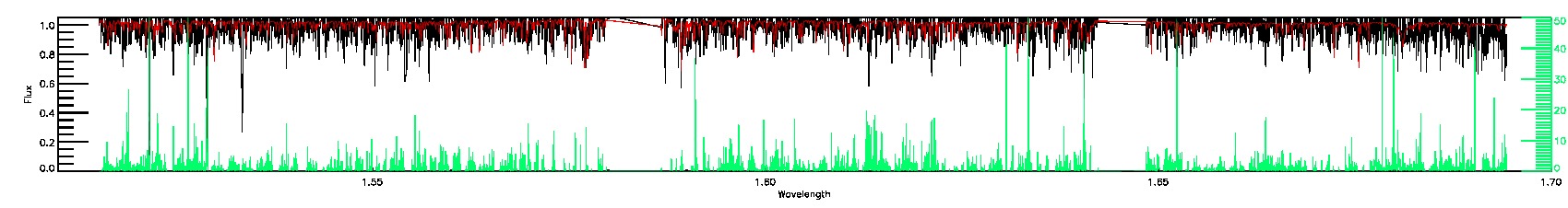
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6375. | +/- | 132. |
| -10000. | +/- | -NaN |
| 3.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.47 | +/- | 0. |
| 4.47 | +/- | -NaN |
| -1.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 9.36 | +/- | 0. |
| 0.97 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.46 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5566. | +/- | 69. |
| -10000. | +/- | -NaN |
| 5.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.26 | +/- | 1. |
| 1.26 | +/- | -NaN |
| -0.42 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.68 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 24.83 | +/- | 0. |
| 1.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.50 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -1.69 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| 0.04 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5547. | +/- | 97. |
| -10000. | +/- | -NaN |
| 1.59 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -1.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.28 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.13 | +/- | 7. |
| -9999.99 | +/- | -NaN |
| 0.21 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.87 | +/- | 0. |
| 0.84 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.45 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| 1.99 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 1.00 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5976. | +/- | 116. |
| -10000. | +/- | -NaN |
| 2.65 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.87 | +/- | 1. |
| 0.87 | +/- | -NaN |
| -1.30 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.25 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 16.54 | +/- | 0. |
| 1.22 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.11 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 7446. | +/- | 97. |
| -10000. | +/- | -NaN |
| 3.46 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.78 | +/- | 1. |
| 1.78 | +/- | -NaN |
| -1.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 0.60 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.59 | +/- | 0. |
| 0.20 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -1.67 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7875. | +/- | 176. |
| -10000. | +/- | -NaN |
| 3.74 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.03 | +/- | 1. |
| 1.03 | +/- | -NaN |
| 0.74 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.86 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 94.32 | +/- | 0. |
| 1.97 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.39 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6621. | +/- | 85. |
| -10000. | +/- | -NaN |
| 2.68 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.88 | +/- | 0. |
| 2.88 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.14 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 25.83 | +/- | 0. |
| 1.41 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.50 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -2.14 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.69 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6363. | +/- | 97. |
| -10000. | +/- | -NaN |
| 2.58 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.48 | +/- | 0. |
| 2.48 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 88.47 | +/- | 0. |
| 1.95 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 5004. | +/- | 94. |
| -10000. | +/- | -NaN |
| 2.80 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.49 | +/- | 0. |
| 0.49 | +/- | -NaN |
| -0.87 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.93 | +/- | 0. |
| 0.69 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.42 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| 1.26 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.43 |
| -9999.00 |

VERY_BRIGHT_NEIGHBOR
STAR_BAD
STAR_WARN,SN_WARN
359-02-O
| 6318. | +/- | 50. |
| -10000. | +/- | -NaN |
| 2.61 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.76 | +/- | 0. |
| 4.76 | +/- | -NaN |
| -1.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 0.63 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.41 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.23 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| 0.99 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5708. | +/- | 134. |
| -10000. | +/- | -NaN |
| 2.69 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.83 | +/- | 1. |
| 0.83 | +/- | -NaN |
| -1.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 20.20 | +/- | 0. |
| 1.31 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.99 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 4244. | +/- | 22. |
| -10000. | +/- | -NaN |
| 2.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| -0.70 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.45 | +/- | 0. |
| 0.65 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.68 |
| -9999.00 |
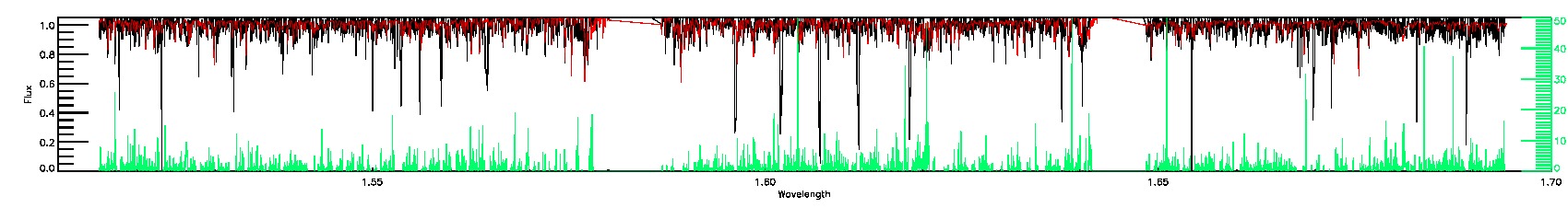
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6430. | +/- | 155. |
| -10000. | +/- | -NaN |
| 3.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.53 | +/- | 1. |
| 1.53 | +/- | -NaN |
| -0.96 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 11.44 | +/- | 0. |
| 1.06 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.43 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| 0.83 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7040. | +/- | 123. |
| -10000. | +/- | -NaN |
| 3.62 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.44 | +/- | 0. |
| 0.44 | +/- | -NaN |
| -0.88 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 0.91 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 21.80 | +/- | 0. |
| 1.34 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.48 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 3606. | +/- | 34. |
| -10000. | +/- | -NaN |
| 0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.30 | +/- | 0. |
| 0.30 | +/- | -NaN |
| -1.87 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.85 | +/- | 0. |
| 0.95 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| -1.52 |
| -9999.00 |
VERY_BRIGHT_NEIGHBOR
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 7561. | +/- | 115. |
| -10000. | +/- | -NaN |
| 3.82 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.55 | +/- | 0. |
| 1.55 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.32 | +/- | 0. |
| 0.92 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.47 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| 0.55 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6062. | +/- | 128. |
| -10000. | +/- | -NaN |
| 2.69 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.38 | +/- | 1. |
| 1.38 | +/- | -NaN |
| -1.41 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.08 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 78.89 | +/- | 0. |
| 1.90 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.07 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 4439. | +/- | 20. |
| -10000. | +/- | -NaN |
| 2.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.40 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.35 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.41 | +/- | 0. |
| 0.38 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.64 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 3588. | +/- | 25. |
| -10000. | +/- | -NaN |
| 0.58 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| -1.86 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.46 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.79 | +/- | 0. |
| 0.94 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -2.49 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 4250. | +/- | 87. |
| -10000. | +/- | -NaN |
| 1.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.35 | +/- | 0. |
| 0.35 | +/- | -NaN |
| -1.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.32 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.58 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.15 | +/- | 0. |
| 0.79 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.31 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -1.28 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6078. | +/- | 124. |
| -10000. | +/- | -NaN |
| 2.75 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.21 | +/- | 0. |
| 1.21 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 23.32 | +/- | 0. |
| 1.37 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.85 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6787. | +/- | 165. |
| -10000. | +/- | -NaN |
| 3.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.27 | +/- | 1. |
| 1.27 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.57 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.76 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.46 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -1.61 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.45 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 3847. | +/- | 33. |
| -10000. | +/- | -NaN |
| 1.46 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| -1.52 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.34 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.19 | +/- | 0. |
| 0.86 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| 0.06 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 3491. | +/- | 25. |
| -10000. | +/- | -NaN |
| 0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.76 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.34 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6173. | +/- | 136. |
| -10000. | +/- | -NaN |
| 3.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.82 | +/- | 0. |
| 0.82 | +/- | -NaN |
| 0.59 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 86.56 | +/- | 0. |
| 1.94 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 1.33 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -1.77 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 4908. | +/- | 90. |
| -10000. | +/- | -NaN |
| 1.64 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.38 | +/- | 0. |
| 0.38 | +/- | -NaN |
| -0.98 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.42 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.23 | +/- | 0. |
| 0.72 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.37 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -1.18 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 5604. | +/- | 165. |
| -10000. | +/- | -NaN |
| 2.70 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.58 | +/- | 1. |
| 0.58 | +/- | -NaN |
| -1.62 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.87 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| 0.35 | +/- | 8. |
| -9999.99 | +/- | -NaN |
| 0.79 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.62 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.60 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -2.41 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 5473. | +/- | 101. |
| -10000. | +/- | -NaN |
| 2.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.67 | +/- | 0. |
| 0.67 | +/- | -NaN |
| -1.57 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.94 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.33 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.88 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.40 | +/- | 0. |
| 0.87 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.96 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -1.15 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7733. | +/- | 162. |
| -10000. | +/- | -NaN |
| 3.82 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.29 | +/- | 1. |
| 1.29 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.24 | +/- | 1. |
| 0.63 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.45 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.48 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7435. | +/- | 153. |
| -10000. | +/- | -NaN |
| 3.88 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.44 | +/- | 0. |
| 1.44 | +/- | -NaN |
| -1.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 6. |
| -9999.99 | +/- | -NaN |
| 0.97 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 15.55 | +/- | 0. |
| 1.19 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.99 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6159. | +/- | 74. |
| -10000. | +/- | -NaN |
| 2.63 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.46 | +/- | 0. |
| 4.46 | +/- | -NaN |
| -1.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 9. |
| -9999.99 | +/- | -NaN |
| 0.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.18 | +/- | 0. |
| 1.09 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -1.32 |
| -9999.00 |
PERSIST_JUMP_NEG
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 3314. | +/- | 60. |
| -10000. | +/- | -NaN |
| 2.80 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.97 | +/- | 1. |
| 0.97 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.19 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 0.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.76 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.80 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 2.00 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -1.55 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 5495. | +/- | 103. |
| -10000. | +/- | -NaN |
| 1.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.14 | +/- | 1. |
| 1.14 | +/- | -NaN |
| -2.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.41 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| -0.26 | +/- | 14. |
| -9999.99 | +/- | -NaN |
| 0.70 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 12.65 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.40 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| -1.10 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 4680. | +/- | 82. |
| -10000. | +/- | -NaN |
| 2.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -0.93 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.10 | +/- | 0. |
| 0.71 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -1.93 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.26 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 5353. | +/- | 130. |
| -10000. | +/- | -NaN |
| 1.80 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.52 | +/- | 1. |
| 0.52 | +/- | -NaN |
| -2.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.46 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| -0.23 | +/- | 9. |
| -9999.99 | +/- | -NaN |
| 0.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.45 | +/- | 0. |
| 1.02 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.44 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -2.49 |
| -9999.00 |
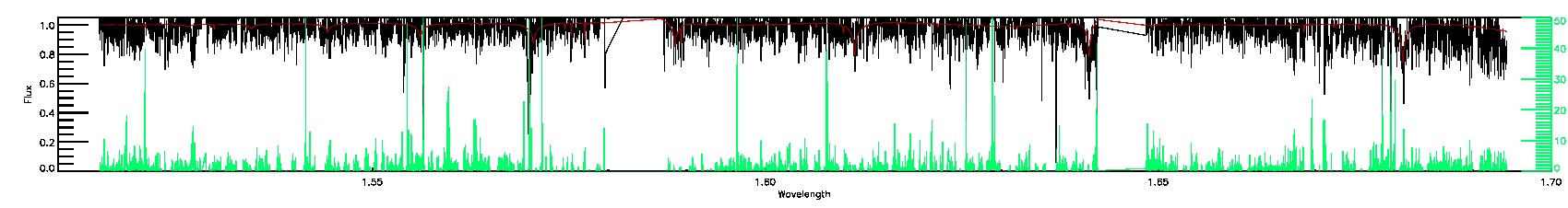
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 4534. | +/- | 82. |
| -10000. | +/- | -NaN |
| 0.76 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.80 | +/- | 1. |
| 0.80 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 0.70 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 12.76 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.69 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.44 |
| -9999.00 |

359-02-O
| 3710. | +/- | 2. |
| 3787. | +/- | 80. |
| 1.57 | +/- | 0. |
| 1.33 | +/- | 0. |
| 1.31 | +/- | 0. |
| 1.31 | +/- | -NaN |
| 0.25 | +/- | 0. |
| 0.26 | +/- | 0. |
| 0.01 | +/- | 0. |
| 0.01 | +/- | -NaN |
| 0.37 | +/- | 0. |
| 0.37 | +/- | -NaN |
| 0.04 | +/- | 0. |
| 0.00 | +/- | 0. |
| 2.55 | +/- | 0. |
| 0.41 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.57 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 4516. | +/- | 34. |
| -10000. | +/- | -NaN |
| 2.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.39 | +/- | 0. |
| 0.39 | +/- | -NaN |
| -0.88 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.94 | +/- | 0. |
| 0.69 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.86 |
| -9999.00 |
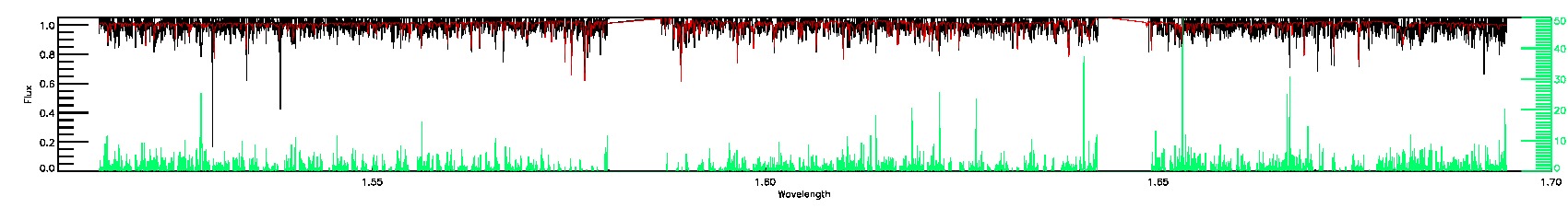
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6900. | +/- | 104. |
| -10000. | +/- | -NaN |
| 3.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.32 | +/- | 2. |
| 1.32 | +/- | -NaN |
| -1.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 37. |
| -9999.99 | +/- | -NaN |
| 0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.66 | +/- | 1. |
| 0.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -1.57 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6720. | +/- | 45. |
| -10000. | +/- | -NaN |
| 2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.63 | +/- | 1. |
| 1.63 | +/- | -NaN |
| -1.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.59 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 60.49 | +/- | 0. |
| 1.78 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.47 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 1.00 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6587. | +/- | 101. |
| -10000. | +/- | -NaN |
| 2.68 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.54 | +/- | 0. |
| 4.54 | +/- | -NaN |
| -1.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.75 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 11.53 | +/- | 0. |
| 1.06 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.40 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -1.47 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 1.00 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 5960. | +/- | 86. |
| -10000. | +/- | -NaN |
| 1.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.08 | +/- | 1. |
| 2.08 | +/- | -NaN |
| -1.58 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.30 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.31 | +/- | 8. |
| -9999.99 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.46 | +/- | 0. |
| 0.87 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -1.26 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| 0.23 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 4821. | +/- | 405. |
| -10000. | +/- | -NaN |
| 3.54 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 1.30 | +/- | 10. |
| 1.30 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 12.76 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6583. | +/- | 112. |
| -10000. | +/- | -NaN |
| 2.60 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.80 | +/- | 1. |
| 0.80 | +/- | -NaN |
| -0.30 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 88.02 | +/- | 0. |
| 1.94 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.39 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 6561. | +/- | 34. |
| -10000. | +/- | -NaN |
| 2.51 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.76 | +/- | 3. |
| 0.76 | +/- | -NaN |
| -2.35 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 18. |
| -9999.99 | +/- | -NaN |
| 0.45 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 24.24 | +/- | 0. |
| 1.38 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -2.24 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 6707. | +/- | 95. |
| -10000. | +/- | -NaN |
| 3.64 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.33 | +/- | 1. |
| 0.33 | +/- | -NaN |
| 0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 9.47 | +/- | 0. |
| 0.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 5110. | +/- | 76. |
| -10000. | +/- | -NaN |
| 1.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.58 | +/- | 1. |
| 0.58 | +/- | -NaN |
| -1.56 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.21 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.39 | +/- | 0. |
| 0.87 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.95 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD
STAR_WARN,SN_WARN
359-02-O
| 8655. | +/- | 258. |
| -10000. | +/- | -NaN |
| 4.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| -1.45 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 91.69 | +/- | 0. |
| 1.96 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.75 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -1.69 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.78 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 4445. | +/- | 64. |
| -10000. | +/- | -NaN |
| 2.64 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.43 | +/- | 0. |
| 0.43 | +/- | -NaN |
| -0.95 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.17 | +/- | 0. |
| 0.71 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.27 |
| -9999.00 |

PERSIST_JUMP_NEG,SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7240. | +/- | 201. |
| -10000. | +/- | -NaN |
| 3.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.71 | +/- | 2. |
| 0.71 | +/- | -NaN |
| -0.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 88.37 | +/- | 0. |
| 1.95 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
BRIGHT_NEIGHBOR,PERSIST_JUMP_NEG
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5626. | +/- | 118. |
| -10000. | +/- | -NaN |
| 2.63 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.37 | +/- | 2. |
| 1.37 | +/- | -NaN |
| -1.45 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| -0.30 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.72 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -1.69 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| 0.78 |
| -9999.00 |
PERSIST_JUMP_NEG,SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 4942. | +/- | 1174. |
| -10000. | +/- | -NaN |
| 4.18 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| -2.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| -0.08 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 91.73 | +/- | 1. |
| 1.96 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.47 |
| -9999.00 |

STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5957. | +/- | 187. |
| -10000. | +/- | -NaN |
| 3.45 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.24 | +/- | 1. |
| 1.24 | +/- | -NaN |
| -1.71 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.60 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.33 | +/- | 15. |
| -9999.99 | +/- | -NaN |
| 0.59 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.04 | +/- | 0. |
| 0.91 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.60 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.50 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 5507. | +/- | 184. |
| -10000. | +/- | -NaN |
| 3.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.34 | +/- | 2. |
| 2.34 | +/- | -NaN |
| -2.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.26 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| -0.11 | +/- | 10. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 9.80 | +/- | 0. |
| 0.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.14 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -9999.00 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 5784. | +/- | 120. |
| -10000. | +/- | -NaN |
| 2.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.09 | +/- | 2. |
| 1.09 | +/- | -NaN |
| -1.54 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.27 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| -0.40 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.28 | +/- | 0. |
| 0.86 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.39 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -2.38 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6087. | +/- | 125. |
| -10000. | +/- | -NaN |
| 2.58 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.90 | +/- | 8. |
| 0.90 | +/- | -NaN |
| -2.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 10.70 | +/- | 1. |
| 1.03 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.50 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5559. | +/- | 755. |
| -10000. | +/- | -NaN |
| 4.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.04 | +/- | 20. |
| 1.04 | +/- | -NaN |
| -2.48 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 9. |
| -9999.99 | +/- | -NaN |
| -0.32 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 3.12 | +/- | 4. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.52 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 4837. | +/- | 96. |
| -10000. | +/- | -NaN |
| 1.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.59 | +/- | 1. |
| 0.59 | +/- | -NaN |
| -2.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.69 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.34 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.98 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.08 | +/- | 0. |
| 1.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.91 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.48 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 6147. | +/- | 95. |
| -10000. | +/- | -NaN |
| 2.89 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.78 | +/- | 0. |
| 4.78 | +/- | -NaN |
| -0.71 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 13.19 | +/- | 0. |
| 1.12 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 4198. | +/- | 44. |
| -10000. | +/- | -NaN |
| 1.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| -1.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.61 | +/- | 0. |
| 0.75 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.45 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5671. | +/- | 101. |
| -10000. | +/- | -NaN |
| 2.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.43 | +/- | 1. |
| 2.43 | +/- | -NaN |
| -1.70 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.39 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.41 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.97 | +/- | 0. |
| 0.90 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.63 |
| -9999.00 |

STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5115. | +/- | 1340. |
| -10000. | +/- | -NaN |
| 5.05 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.82 | +/- | 11. |
| 0.82 | +/- | -NaN |
| -1.81 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 95.81 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -2.14 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -2.34 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 5475. | +/- | 92. |
| -10000. | +/- | -NaN |
| 2.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.80 | +/- | 1. |
| 0.80 | +/- | -NaN |
| -2.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.40 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 7. |
| -9999.99 | +/- | -NaN |
| 0.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 9.77 | +/- | 0. |
| 0.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.28 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.81 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -2.20 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 5228. | +/- | 102. |
| -10000. | +/- | -NaN |
| 1.57 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.93 | +/- | 1. |
| 1.93 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.21 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| -0.08 | +/- | 11. |
| -9999.99 | +/- | -NaN |
| 0.52 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.76 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -2.00 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -2.50 |
| -9999.00 |

STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6730. | +/- | 166. |
| -10000. | +/- | -NaN |
| 3.53 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.12 | +/- | 0. |
| 2.12 | +/- | -NaN |
| -0.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.35 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 66.68 | +/- | 0. |
| 1.82 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.03 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5568. | +/- | 110. |
| -10000. | +/- | -NaN |
| 2.88 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.97 | +/- | 2. |
| 0.97 | +/- | -NaN |
| -1.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 93.24 | +/- | 0. |
| 1.97 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.74 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.48 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6405. | +/- | 85. |
| -10000. | +/- | -NaN |
| 2.55 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.84 | +/- | 1. |
| 1.84 | +/- | -NaN |
| -1.84 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.18 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.34 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.90 | +/- | 1. |
| 0.28 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.92 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6382. | +/- | 151. |
| -10000. | +/- | -NaN |
| 3.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.32 | +/- | 1. |
| 0.32 | +/- | -NaN |
| -0.98 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.96 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 85.04 | +/- | 0. |
| 1.93 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.45 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 4988. | +/- | 99. |
| -10000. | +/- | -NaN |
| 1.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.10 | +/- | 2. |
| 2.10 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 18. |
| -9999.99 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.76 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -2.50 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 4523. | +/- | 113. |
| -10000. | +/- | -NaN |
| 0.92 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.83 | +/- | 1. |
| 2.83 | +/- | -NaN |
| -2.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.39 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.72 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -1.23 |
| -9999.00 |

BRIGHT_NEIGHBOR
STAR_WARN,SN_WARN
359-02-O
| 7943. | +/- | 59. |
| 7747. | +/- | 315. |
| 4.30 | +/- | 0. |
| 3.94 | +/- | 0. |
| 4.29 | +/- | 0. |
| 4.29 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -0.13 | +/- | -NaN |
| 0.80 | +/- | 0. |
| 0.80 | +/- | -NaN |
| 0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 19.91 | +/- | 0. |
| 1.30 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 4658. | +/- | 99. |
| -10000. | +/- | -NaN |
| 1.21 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.57 | +/- | 0. |
| 0.57 | +/- | -NaN |
| -2.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.71 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.53 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.94 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.92 | +/- | 0. |
| 1.04 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.94 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.25 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.34 |
| -9999.00 |

BRIGHT_NEIGHBOR
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5594. | +/- | 68. |
| -10000. | +/- | -NaN |
| 1.66 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.13 | +/- | 0. |
| 1.13 | +/- | -NaN |
| -2.24 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.95 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| 0.70 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.97 | +/- | 0. |
| 1.04 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.95 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.91 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 3004. | +/- | 0. |
| -10000. | +/- | -NaN |
| 3.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.06 | +/- | 0. |
| 3.06 | +/- | -NaN |
| 0.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.26 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.50 | +/- | 0. |
| 0.40 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.45 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 1.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5640. | +/- | 113. |
| -10000. | +/- | -NaN |
| 2.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.58 | +/- | 1. |
| 3.58 | +/- | -NaN |
| -1.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.61 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.26 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.70 | +/- | 0. |
| 0.83 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.49 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -2.47 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6629. | +/- | 130. |
| -10000. | +/- | -NaN |
| 3.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.71 | +/- | 0. |
| 4.71 | +/- | -NaN |
| -1.30 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 0.62 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.22 | +/- | 0. |
| 1.09 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.21 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.85 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5139. | +/- | 68. |
| -10000. | +/- | -NaN |
| 1.24 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.09 | +/- | 0. |
| 1.09 | +/- | -NaN |
| -2.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.20 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 0.85 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.66 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -1.04 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.49 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 5625. | +/- | 88. |
| -10000. | +/- | -NaN |
| 1.71 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| -1.76 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.97 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.15 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.26 | +/- | 0. |
| 0.92 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.96 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -2.15 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6678. | +/- | 102. |
| -10000. | +/- | -NaN |
| 2.86 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.87 | +/- | 1. |
| 0.87 | +/- | -NaN |
| -1.42 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.81 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 27.77 | +/- | 0. |
| 1.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5531. | +/- | 107. |
| -10000. | +/- | -NaN |
| 2.88 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.53 | +/- | 6. |
| 0.53 | +/- | -NaN |
| -2.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.26 | +/- | 56. |
| -9999.99 | +/- | -NaN |
| -0.14 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 5.46 | +/- | 1. |
| 0.74 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.43 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 3005. | +/- | 1. |
| -10000. | +/- | -NaN |
| 1.21 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.96 | +/- | 0. |
| 0.96 | +/- | -NaN |
| -2.45 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.57 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.40 | +/- | 0. |
| 1.09 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.00 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 2.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -2.41 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 4998. | +/- | 883. |
| -10000. | +/- | -NaN |
| 4.28 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.80 | +/- | 15. |
| 0.80 | +/- | -NaN |
| -2.46 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.25 | +/- | 7. |
| -9999.99 | +/- | -NaN |
| -0.14 | +/- | 12. |
| -9999.99 | +/- | -NaN |
| -0.36 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 12.46 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.25 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| -0.88 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -9999.00 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 5419. | +/- | 123. |
| -10000. | +/- | -NaN |
| 1.98 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.51 | +/- | 1. |
| 0.51 | +/- | -NaN |
| -1.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.49 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.49 | +/- | 8. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.62 | +/- | 0. |
| 0.82 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.65 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.95 |
| -9999.00 |

STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5535. | +/- | 155. |
| -10000. | +/- | -NaN |
| 3.67 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.64 | +/- | 3. |
| 0.64 | +/- | -NaN |
| -2.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 11. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.96 | +/- | 1. |
| 0.95 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.49 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7014. | +/- | 114. |
| -10000. | +/- | -NaN |
| 3.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.91 | +/- | 0. |
| 0.91 | +/- | -NaN |
| 0.41 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 36.60 | +/- | 0. |
| 1.56 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -0.10 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5822. | +/- | 115. |
| -10000. | +/- | -NaN |
| 2.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.96 | +/- | 1. |
| 0.96 | +/- | -NaN |
| -2.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.65 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 11. |
| -9999.99 | +/- | -NaN |
| 0.77 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.61 | +/- | 0. |
| 1.03 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.87 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -1.87 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -2.21 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 5217. | +/- | 140. |
| -10000. | +/- | -NaN |
| 2.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.43 | +/- | 0. |
| 2.43 | +/- | -NaN |
| -2.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.99 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.26 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 0.73 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.11 | +/- | 0. |
| 1.00 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -9999.00 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6219. | +/- | 38. |
| -10000. | +/- | -NaN |
| 2.51 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.79 | +/- | 1. |
| 1.79 | +/- | -NaN |
| -0.93 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 78.51 | +/- | 0. |
| 1.89 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.68 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 5849. | +/- | 95. |
| -10000. | +/- | -NaN |
| 2.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.40 | +/- | 2. |
| 1.40 | +/- | -NaN |
| -2.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 7. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 11.22 | +/- | 0. |
| 1.05 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -2.14 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -2.28 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.28 |
| -9999.00 |

STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 3925. | +/- | 31. |
| -10000. | +/- | -NaN |
| 1.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -1.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.37 | +/- | 0. |
| 0.73 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -1.77 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| -1.18 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6027. | +/- | 148. |
| -10000. | +/- | -NaN |
| 3.34 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.68 | +/- | 0. |
| 1.68 | +/- | -NaN |
| 0.32 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 11.41 | +/- | 0. |
| 1.06 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 1.00 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 3346. | +/- | 32. |
| -10000. | +/- | -NaN |
| 2.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.66 | +/- | 0. |
| 1.66 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.76 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.76 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -2.46 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 5742. | +/- | 73. |
| -10000. | +/- | -NaN |
| 2.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.78 | +/- | 0. |
| 4.78 | +/- | -NaN |
| -1.60 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.48 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.54 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.48 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.37 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6438. | +/- | 138. |
| -10000. | +/- | -NaN |
| 3.21 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.98 | +/- | 1. |
| 0.98 | +/- | -NaN |
| -1.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.18 | +/- | 12. |
| -9999.99 | +/- | -NaN |
| 0.94 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 17.61 | +/- | 0. |
| 1.25 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.21 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 4263. | +/- | 36. |
| -10000. | +/- | -NaN |
| 1.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -1.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.65 | +/- | 0. |
| 0.75 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -1.67 |
| -9999.00 |

STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5534. | +/- | 172. |
| -10000. | +/- | -NaN |
| 3.70 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.72 | +/- | 8. |
| 1.72 | +/- | -NaN |
| -2.42 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 22. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 5.08 | +/- | 2. |
| 0.71 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.52 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6218. | +/- | 126. |
| -10000. | +/- | -NaN |
| 2.96 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.19 | +/- | 0. |
| 4.19 | +/- | -NaN |
| -1.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.51 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 11.01 | +/- | 0. |
| 1.04 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 5333. | +/- | 125. |
| -10000. | +/- | -NaN |
| 1.85 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.82 | +/- | 2. |
| 1.82 | +/- | -NaN |
| -2.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 13. |
| -9999.99 | +/- | -NaN |
| 0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.66 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.47 |
| -9999.00 |

BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 5683. | +/- | 92. |
| -10000. | +/- | -NaN |
| 2.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.78 | +/- | 0. |
| 4.78 | +/- | -NaN |
| -1.32 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.41 | +/- | 0. |
| 0.81 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.32 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5723. | +/- | 148. |
| -10000. | +/- | -NaN |
| 2.86 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.76 | +/- | 1. |
| 1.76 | +/- | -NaN |
| -2.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 20. |
| -9999.99 | +/- | -NaN |
| 0.85 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.87 | +/- | 1. |
| 0.90 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.17 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 8940. | +/- | 746. |
| -10000. | +/- | -NaN |
| 3.72 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.00 | +/- | 0. |
| 10.00 | +/- | -NaN |
| 0.46 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 93.71 | +/- | 0. |
| 1.97 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.95 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -0.59 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -1.32 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| 1.00 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7986. | +/- | 207. |
| -10000. | +/- | -NaN |
| 3.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.29 | +/- | 0. |
| 4.29 | +/- | -NaN |
| 0.91 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.69 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 81.53 | +/- | 0. |
| 1.91 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 0.73 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5733. | +/- | 102. |
| -10000. | +/- | -NaN |
| 2.89 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.21 | +/- | 0. |
| 1.21 | +/- | -NaN |
| -0.59 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.24 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 14.32 | +/- | 0. |
| 1.16 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.49 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 1.50 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.03 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 4332. | +/- | 47. |
| -10000. | +/- | -NaN |
| 2.34 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.33 | +/- | 0. |
| 0.33 | +/- | -NaN |
| -0.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.80 | +/- | 0. |
| 0.58 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.45 |
| -9999.00 |
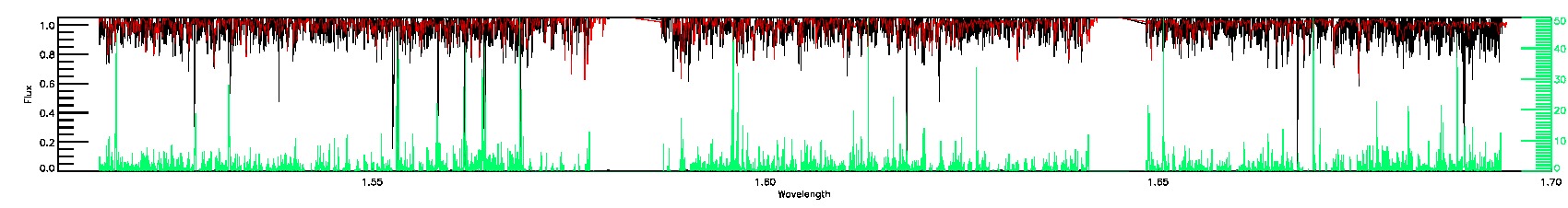
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6860. | +/- | 130. |
| -10000. | +/- | -NaN |
| 3.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.46 | +/- | 1. |
| 2.46 | +/- | -NaN |
| -1.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 0.90 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 22.20 | +/- | 0. |
| 1.35 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -2.22 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.79 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6957. | +/- | 111. |
| -10000. | +/- | -NaN |
| 3.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.70 | +/- | 0. |
| 1.70 | +/- | -NaN |
| -0.95 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| 0.60 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 28.32 | +/- | 0. |
| 1.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.18 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -0.53 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.28 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7863. | +/- | 179. |
| -10000. | +/- | -NaN |
| 3.45 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.04 | +/- | 0. |
| 1.04 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 19.22 | +/- | 0. |
| 1.28 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| -2.47 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6003. | +/- | 79. |
| -10000. | +/- | -NaN |
| 2.57 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.81 | +/- | 0. |
| 2.81 | +/- | -NaN |
| -1.82 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| 0.56 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.72 | +/- | 0. |
| 0.83 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.17 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| -1.57 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| -0.86 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -1.56 |
| -9999.00 |
| -2.18 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.05 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.11 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 3849. | +/- | 27. |
| -10000. | +/- | -NaN |
| 1.21 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| -1.35 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.51 | +/- | 0. |
| 0.81 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.81 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -1.41 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -0.99 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 5612. | +/- | 128. |
| -10000. | +/- | -NaN |
| 2.14 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.68 | +/- | 1. |
| 1.68 | +/- | -NaN |
| -1.77 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.91 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 8.33 | +/- | 0. |
| 0.92 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.85 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.04 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -1.93 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.69 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -1.79 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -2.48 |
| -9999.00 |

VERY_BRIGHT_NEIGHBOR
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6312. | +/- | 89. |
| -10000. | +/- | -NaN |
| 2.63 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.36 | +/- | 0. |
| 0.36 | +/- | -NaN |
| -1.87 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.13 | +/- | 7. |
| -9999.99 | +/- | -NaN |
| 0.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 30.32 | +/- | 0. |
| 1.48 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| -2.27 |
| -9999.00 |
| -2.06 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 5196. | +/- | 78. |
| -10000. | +/- | -NaN |
| 1.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.29 | +/- | 1. |
| 2.29 | +/- | -NaN |
| -2.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.33 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.45 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.54 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 10.22 | +/- | 0. |
| 1.01 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.89 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| -1.74 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.13 |
| -9999.00 |

STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6055. | +/- | 152. |
| -10000. | +/- | -NaN |
| 3.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.99 | +/- | 0. |
| 0.99 | +/- | -NaN |
| 0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.45 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 1.07 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.30 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -2.07 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.89 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7962. | +/- | 152. |
| -10000. | +/- | -NaN |
| 4.21 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.01 | +/- | 2. |
| 1.01 | +/- | -NaN |
| -1.54 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.26 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.30 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 10.34 | +/- | 1. |
| 1.01 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -1.93 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -0.91 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6926. | +/- | 64. |
| -10000. | +/- | -NaN |
| 2.53 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.00 | +/- | 2. |
| 1.00 | +/- | -NaN |
| -2.01 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.14 | +/- | 12. |
| -9999.99 | +/- | -NaN |
| 0.45 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 61.84 | +/- | 0. |
| 1.79 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -2.32 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -1.69 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -1.99 |
| -9999.00 |
| -1.39 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -2.01 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.16 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5853. | +/- | 90. |
| -10000. | +/- | -NaN |
| 2.58 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.20 | +/- | 1. |
| 1.20 | +/- | -NaN |
| -1.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 6. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 11.03 | +/- | 0. |
| 1.04 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| -2.12 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.58 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 3778. | +/- | 41. |
| -10000. | +/- | -NaN |
| 1.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.46 | +/- | 0. |
| 0.46 | +/- | -NaN |
| -1.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.97 | +/- | 0. |
| 0.84 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.11 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -1.40 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 7310. | +/- | 62. |
| -10000. | +/- | -NaN |
| 3.22 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.23 | +/- | 1. |
| 1.23 | +/- | -NaN |
| -1.82 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| -0.41 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 2.48 | +/- | 0. |
| 0.39 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.31 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -1.34 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -2.10 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -1.50 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 4386. | +/- | 21. |
| -10000. | +/- | -NaN |
| 2.22 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.90 | +/- | 0. |
| 0.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.35 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -0.25 |
| -9999.00 |
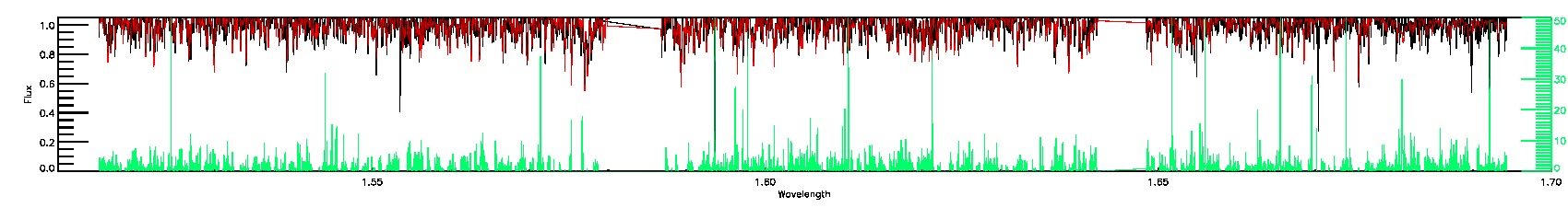
VERY_BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6347. | +/- | 299. |
| -10000. | +/- | -NaN |
| 3.85 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.03 | +/- | 1. |
| 2.03 | +/- | -NaN |
| -0.75 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.30 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.02 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.41 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 4565. | +/- | 92. |
| -10000. | +/- | -NaN |
| 0.56 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.28 | +/- | 1. |
| 1.28 | +/- | -NaN |
| -2.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.42 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.51 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 12.66 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -1.49 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.63 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.16 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.36 |
| -9999.00 |
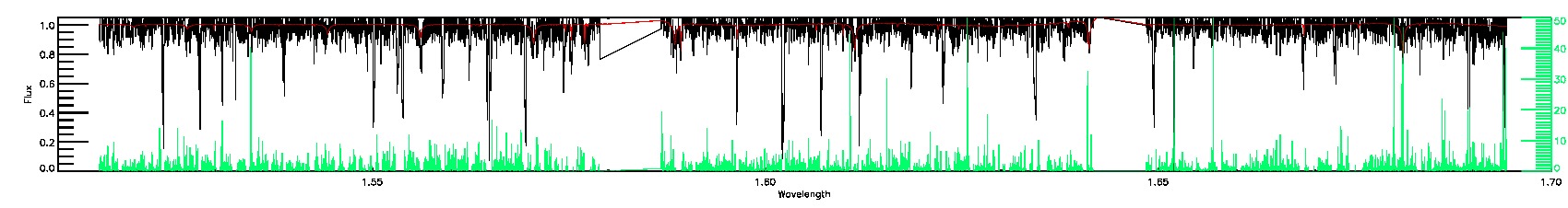
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6407. | +/- | 160. |
| -10000. | +/- | -NaN |
| 3.58 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.10 | +/- | 1. |
| 1.10 | +/- | -NaN |
| -1.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| 0.55 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 23.01 | +/- | 0. |
| 1.36 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -1.78 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.52 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.99 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 5259. | +/- | 78. |
| -10000. | +/- | -NaN |
| 1.62 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.13 | +/- | 1. |
| 1.13 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.33 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.37 | +/- | 13. |
| -9999.99 | +/- | -NaN |
| 0.52 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.75 | +/- | 0. |
| 1.11 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.83 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.37 |
| -9999.00 |

STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5761. | +/- | 135. |
| -10000. | +/- | -NaN |
| 2.71 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.83 | +/- | 0. |
| 0.83 | +/- | -NaN |
| -0.60 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.21 | +/- | 0. |
| 0.62 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 1.07 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| -1.67 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| 1.00 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 5243. | +/- | 93. |
| -10000. | +/- | -NaN |
| 1.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.54 | +/- | 3. |
| 1.54 | +/- | -NaN |
| -2.46 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.03 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| -0.35 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 12.46 | +/- | 0. |
| 1.10 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.14 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.26 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.46 |
| -9999.00 |

VERY_BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7205. | +/- | 118. |
| -10000. | +/- | -NaN |
| 3.66 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.51 | +/- | 0. |
| 4.51 | +/- | -NaN |
| -0.89 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 0.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.06 | +/- | 1. |
| 0.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -2.09 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -2.30 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.38 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6702. | +/- | 130. |
| -10000. | +/- | -NaN |
| 3.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.87 | +/- | 1. |
| 1.87 | +/- | -NaN |
| -1.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 | +/- | 8. |
| -9999.99 | +/- | -NaN |
| 0.86 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 77.52 | +/- | 0. |
| 1.89 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| 1.00 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 6212. | +/- | 28. |
| -10000. | +/- | -NaN |
| 2.51 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.25 | +/- | 0. |
| 4.25 | +/- | -NaN |
| -1.60 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.32 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.10 | +/- | 0. |
| 1.08 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.15 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -1.62 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -1.60 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 1.00 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7225. | +/- | 122. |
| -10000. | +/- | -NaN |
| 3.52 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.15 | +/- | 0. |
| 2.15 | +/- | -NaN |
| -0.35 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.24 | +/- | 6. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 17.16 | +/- | 0. |
| 1.23 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.43 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.36 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| 1.00 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6657. | +/- | 99. |
| -10000. | +/- | -NaN |
| 2.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.93 | +/- | 0. |
| 0.93 | +/- | -NaN |
| 0.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 41.15 | +/- | 0. |
| 1.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.17 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.76 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| -0.07 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.51 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.33 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 3943. | +/- | 26. |
| -10000. | +/- | -NaN |
| 0.97 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.31 | +/- | 0. |
| 0.31 | +/- | -NaN |
| -1.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 5.65 | +/- | 0. |
| 0.75 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -0.99 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -1.43 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -1.02 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -0.25 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 4692. | +/- | 54. |
| -10000. | +/- | -NaN |
| 1.56 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.79 | +/- | 1. |
| 0.79 | +/- | -NaN |
| -1.35 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.60 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.11 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.53 | +/- | 0. |
| 0.81 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 2.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.51 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.99 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 5274. | +/- | 114. |
| -10000. | +/- | -NaN |
| 2.13 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.78 | +/- | 1. |
| 1.78 | +/- | -NaN |
| -2.42 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.18 | +/- | 12. |
| -9999.99 | +/- | -NaN |
| -0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.15 | +/- | 0. |
| 1.08 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.12 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -1.31 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.40 |
| -9999.00 |

VERY_BRIGHT_NEIGHBOR
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5643. | +/- | 98. |
| -10000. | +/- | -NaN |
| 2.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.61 | +/- | 0. |
| 3.61 | +/- | -NaN |
| -2.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.83 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 9.50 | +/- | 0. |
| 0.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.26 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -1.69 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -2.00 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 3571. | +/- | 21. |
| -10000. | +/- | -NaN |
| 0.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| -1.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.11 | +/- | 0. |
| 0.85 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -1.36 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -1.44 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| -1.82 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| -1.51 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| -1.11 |
| -9999.00 |
| -1.42 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 3000. | +/- | 0. |
| -10000. | +/- | -NaN |
| 2.55 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.68 | +/- | 0. |
| 0.68 | +/- | -NaN |
| -1.30 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.21 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.66 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.33 | +/- | 0. |
| 0.80 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.55 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 1.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -1.64 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5999. | +/- | 116. |
| -10000. | +/- | -NaN |
| 2.73 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.58 | +/- | 1. |
| 1.58 | +/- | -NaN |
| -1.24 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 30.89 | +/- | 0. |
| 1.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.06 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -1.54 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.23 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7409. | +/- | 144. |
| -10000. | +/- | -NaN |
| 3.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 3.78 | +/- | 0. |
| 3.78 | +/- | -NaN |
| -0.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 0.72 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.98 | +/- | 1. |
| 0.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.28 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| -0.42 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| 0.80 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -1.59 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6695. | +/- | 135. |
| -10000. | +/- | -NaN |
| 2.91 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.23 | +/- | 0. |
| 2.23 | +/- | -NaN |
| 0.52 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.48 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.22 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 73.67 | +/- | 0. |
| 1.87 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.47 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.27 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| 0.23 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 6105. | +/- | 88. |
| -10000. | +/- | -NaN |
| 2.67 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.94 | +/- | 0. |
| 1.94 | +/- | -NaN |
| -1.25 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.41 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.45 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.10 | +/- | 0. |
| 0.61 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.18 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -0.46 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -1.59 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.21 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5018. | +/- | 101. |
| -10000. | +/- | -NaN |
| 1.26 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.11 | +/- | 1. |
| 1.11 | +/- | -NaN |
| -2.42 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.14 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.11 | +/- | 9. |
| -9999.99 | +/- | -NaN |
| 0.87 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 12.17 | +/- | 0. |
| 1.09 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.25 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| -1.92 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| -2.19 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -2.35 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6255. | +/- | 78. |
| -10000. | +/- | -NaN |
| 2.60 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.80 | +/- | 0. |
| 4.80 | +/- | -NaN |
| -1.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 0.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 18.34 | +/- | 0. |
| 1.26 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.19 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.04 |
| -9999.00 |
BRIGHT_NEIGHBOR,SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6509. | +/- | 96. |
| -10000. | +/- | -NaN |
| 2.71 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.81 | +/- | 1. |
| 0.81 | +/- | -NaN |
| -0.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| -0.29 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 89.47 | +/- | 0. |
| 1.95 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -2.06 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -1.95 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -0.49 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.45 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -1.06 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.37 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6481. | +/- | 125. |
| -10000. | +/- | -NaN |
| 2.76 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.00 | +/- | 6. |
| 1.00 | +/- | -NaN |
| -1.89 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 22. |
| -9999.99 | +/- | -NaN |
| 0.29 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 95.85 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -1.73 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -1.48 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.83 |
| -9999.00 |
| -1.90 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.22 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 4657. | +/- | 34. |
| -10000. | +/- | -NaN |
| 2.80 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.34 | +/- | 0. |
| 0.34 | +/- | -NaN |
| 0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.39 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.64 | +/- | 0. |
| 0.42 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| -1.50 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| -1.46 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| 0.77 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5638. | +/- | 41. |
| -10000. | +/- | -NaN |
| 3.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.88 | +/- | 0. |
| 0.88 | +/- | -NaN |
| -0.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.39 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 17.51 | +/- | 0. |
| 1.24 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| 0.33 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.40 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.29 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.89 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6783. | +/- | 118. |
| -10000. | +/- | -NaN |
| 3.62 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.97 | +/- | 0. |
| 1.97 | +/- | -NaN |
| -1.71 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 10. |
| -9999.99 | +/- | -NaN |
| 0.96 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 9.35 | +/- | 0. |
| 0.97 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.13 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -1.71 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -1.86 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.68 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.29 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -1.55 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7132. | +/- | 133. |
| -10000. | +/- | -NaN |
| 2.99 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.63 | +/- | 0. |
| 2.63 | +/- | -NaN |
| -0.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 1.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 43.27 | +/- | 0. |
| 1.64 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.20 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.61 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.97 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| -0.01 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.39 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6996. | +/- | 122. |
| -10000. | +/- | -NaN |
| 3.22 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.51 | +/- | 0. |
| 1.51 | +/- | -NaN |
| -0.31 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.01 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.08 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.41 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 15.14 | +/- | 0. |
| 1.18 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.49 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| 0.44 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.73 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.57 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -2.50 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6097. | +/- | 265. |
| -10000. | +/- | -NaN |
| 3.41 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.99 | +/- | 6. |
| 0.99 | +/- | -NaN |
| -2.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 14. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 20.90 | +/- | 1. |
| 1.32 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.15 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.00 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -0.00 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.15 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.36 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5656. | +/- | 164. |
| -10000. | +/- | -NaN |
| 2.95 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.28 | +/- | 6. |
| 1.28 | +/- | -NaN |
| -2.49 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 24. |
| -9999.99 | +/- | -NaN |
| -0.10 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 1.63 | +/- | 0. |
| 0.21 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.01 |
| -9999.00 |
| -0.44 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| -2.49 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 5906. | +/- | 121. |
| -10000. | +/- | -NaN |
| 2.71 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.60 | +/- | 2. |
| 0.60 | +/- | -NaN |
| -2.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.96 | +/- | 7. |
| -9999.99 | +/- | -NaN |
| -0.36 | +/- | 9. |
| -9999.99 | +/- | -NaN |
| 0.93 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 10.58 | +/- | 0. |
| 1.02 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.96 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| -2.20 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| -0.31 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| 0.86 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| -1.94 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.97 |
| -9999.00 |
| -1.89 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| -2.48 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,ROTATION_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 5501. | +/- | 118. |
| -10000. | +/- | -NaN |
| 2.21 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.55 | +/- | 0. |
| 4.55 | +/- | -NaN |
| -1.69 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.65 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.19 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.94 | +/- | 0. |
| 0.90 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.91 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| -2.08 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -1.40 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -1.85 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -1.96 |
| -9999.00 |
| -1.38 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.14 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6510. | +/- | 85. |
| -10000. | +/- | -NaN |
| 2.75 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.45 | +/- | 1. |
| 1.45 | +/- | -NaN |
| -0.43 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 95.04 | +/- | 0. |
| 1.98 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.07 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.12 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -2.17 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| 0.60 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -0.17 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -1.21 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -1.49 |
| -9999.00 |
| 1.00 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6256. | +/- | 103. |
| -10000. | +/- | -NaN |
| 2.58 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.99 | +/- | 0. |
| 0.99 | +/- | -NaN |
| -0.96 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.18 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 0.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 66.74 | +/- | 0. |
| 1.82 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.48 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.38 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| 0.32 |
| -9999.00 |
| 0.63 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.44 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -0.93 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| 0.97 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6302. | +/- | 50. |
| -10000. | +/- | -NaN |
| 3.23 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.99 | +/- | 0. |
| 0.99 | +/- | -NaN |
| -0.28 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.22 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 14.62 | +/- | 0. |
| 1.16 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.21 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.52 |
| -9999.00 |
| 0.16 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.58 |
| -9999.00 |
| -0.34 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.74 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.13 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.08 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5638. | +/- | 109. |
| -10000. | +/- | -NaN |
| 2.59 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.99 | +/- | 1. |
| 0.99 | +/- | -NaN |
| -1.40 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.67 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 20.50 | +/- | 0. |
| 1.31 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.30 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| -0.48 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| -0.67 |
| -9999.00 |
| -1.05 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| 0.70 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| -1.91 |
| -9999.00 |
| -1.55 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -2.11 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| -0.66 |
| -9999.00 |
| -2.16 |
| -9999.00 |
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5817. | +/- | 96. |
| -10000. | +/- | -NaN |
| 2.18 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.81 | +/- | 1. |
| 0.81 | +/- | -NaN |
| -1.61 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.10 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.02 | +/- | 6. |
| -9999.99 | +/- | -NaN |
| 0.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.57 | +/- | 0. |
| 0.88 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -2.29 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -2.33 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.31 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.03 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.88 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -2.14 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.40 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,SN_WARN
359-02-O
| 3703. | +/- | 7. |
| -10000. | +/- | -NaN |
| 0.82 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.12 | +/- | 0. |
| 1.12 | +/- | -NaN |
| -0.89 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.16 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.97 | +/- | 0. |
| 0.70 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.14 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.19 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| -0.90 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| 0.23 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -0.98 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -0.54 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -0.43 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -1.67 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6569. | +/- | 162. |
| -10000. | +/- | -NaN |
| 3.94 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.82 | +/- | 0. |
| 0.82 | +/- | -NaN |
| -0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.20 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 27.61 | +/- | 0. |
| 1.44 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.05 |
| -9999.00 |
| 0.04 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| -0.02 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.34 |
| -9999.00 |
| -0.65 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -1.70 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -2.46 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.24 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.78 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 1.00 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5724. | +/- | 122. |
| -10000. | +/- | -NaN |
| 2.33 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.97 | +/- | 0. |
| 0.97 | +/- | -NaN |
| -1.26 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.36 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| -0.44 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.50 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.18 | +/- | 0. |
| 0.79 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.58 |
| -9999.00 |
| 0.58 |
| -9999.00 |
| -0.41 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| 0.78 |
| -9999.00 |
| -0.19 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| -0.21 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.85 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| -1.58 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.44 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 4107. | +/- | 35. |
| -10000. | +/- | -NaN |
| 0.36 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.32 | +/- | 0. |
| 0.32 | +/- | -NaN |
| -1.35 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.03 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.06 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.11 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.51 | +/- | 0. |
| 0.81 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.15 |
| -9999.00 |
| -1.42 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -0.09 |
| -9999.00 |
| -1.12 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| -0.60 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| -0.57 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.18 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -1.72 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| -0.77 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| -0.64 |
| -9999.00 |
| -0.83 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| -1.15 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -1.35 |
| -9999.00 |

STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6247. | +/- | 87. |
| -10000. | +/- | -NaN |
| 2.58 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.46 | +/- | 0. |
| 4.46 | +/- | -NaN |
| -1.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.38 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.27 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 7.11 | +/- | 0. |
| 0.85 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.47 |
| -9999.00 |
| -0.08 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.66 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.11 |
| -9999.00 |
| -1.14 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -1.20 |
| -9999.00 |
| 0.59 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.52 |
| -9999.00 |
| -0.39 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -2.03 |
| -9999.00 |
| -2.13 |
| -9999.00 |
| -1.07 |
| -9999.00 |
| 0.24 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| -0.15 |
| -9999.00 |
| -1.17 |
| -9999.00 |
| -1.24 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.96 |
| -9999.00 |
BRIGHT_NEIGHBOR
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7108. | +/- | 146. |
| -10000. | +/- | -NaN |
| 3.80 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.66 | +/- | 0. |
| 4.66 | +/- | -NaN |
| -1.32 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 | +/- | 5. |
| -9999.99 | +/- | -NaN |
| 0.60 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 6.85 | +/- | 0. |
| 0.84 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.13 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.01 |
| -9999.00 |
| 0.55 |
| -9999.00 |
| -1.80 |
| -9999.00 |
| 0.65 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.46 |
| -9999.00 |
| -0.37 |
| -9999.00 |
| -2.41 |
| -9999.00 |
| -1.18 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| 0.05 |
| -9999.00 |
| -1.64 |
| -9999.00 |
| -1.63 |
| -9999.00 |
| -0.84 |
| -9999.00 |
| -1.16 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -1.79 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7793. | +/- | 191. |
| -10000. | +/- | -NaN |
| 3.24 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.89 | +/- | 0. |
| 0.89 | +/- | -NaN |
| 0.95 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.39 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 30.96 | +/- | 0. |
| 1.49 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.02 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.75 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.92 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| -0.32 |
| -9999.00 |
| 0.93 |
| -9999.00 |
| -1.75 |
| -9999.00 |
| 1.00 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 5793. | +/- | 94. |
| -10000. | +/- | -NaN |
| 2.08 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.53 | +/- | 1. |
| 0.53 | +/- | -NaN |
| -2.21 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.35 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.14 | +/- | 10. |
| -9999.99 | +/- | -NaN |
| 0.85 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 10.76 | +/- | 0. |
| 1.03 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.58 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| 0.28 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.81 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| -2.39 |
| -9999.00 |
| -2.31 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.38 |
| -9999.00 |
| -1.53 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -2.37 |
| -9999.00 |
| -2.35 |
| -9999.00 |
| -2.21 |
| -9999.00 |

SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6749. | +/- | 118. |
| -10000. | +/- | -NaN |
| 3.15 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.75 | +/- | 1. |
| 0.75 | +/- | -NaN |
| -0.51 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 0.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 70.88 | +/- | 0. |
| 1.85 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.08 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| 0.61 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| -1.30 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.40 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -1.28 |
| -9999.00 |
| -0.06 |
| -9999.00 |
| 0.87 |
| -9999.00 |
| -0.35 |
| -9999.00 |
| -2.47 |
| -9999.00 |
| -0.23 |
| -9999.00 |
| -1.10 |
| -9999.00 |
| -0.89 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.79 |
| -9999.00 |
| -1.03 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -0.20 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.49 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,ROTATION_WARN,SN_WARN
359-02-O
| 3279. | +/- | 15. |
| -10000. | +/- | -NaN |
| 2.83 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.78 | +/- | 0. |
| 0.78 | +/- | -NaN |
| -0.67 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.19 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.02 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.37 | +/- | 0. |
| 0.64 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.30 |
| -9999.00 |
| 0.25 |
| -9999.00 |
| 1.99 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| 0.12 |
| -9999.00 |
| -0.82 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.04 |
| -9999.00 |
| 0.08 |
| -9999.00 |
| -0.85 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.33 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.70 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| -0.71 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6805. | +/- | 115. |
| -10000. | +/- | -NaN |
| 3.37 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.55 | +/- | 0. |
| 2.55 | +/- | -NaN |
| -0.91 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 8. |
| -9999.99 | +/- | -NaN |
| 0.34 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 18.28 | +/- | 0. |
| 1.26 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.40 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| -1.37 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| -0.56 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -1.25 |
| -9999.00 |
| -1.65 |
| -9999.00 |
| -0.76 |
| -9999.00 |
| 0.64 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -2.25 |
| -9999.00 |
| -1.13 |
| -9999.00 |
| -0.92 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.42 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6792. | +/- | 94. |
| -10000. | +/- | -NaN |
| 2.86 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.41 | +/- | 0. |
| 2.41 | +/- | -NaN |
| 0.17 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.67 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 14.38 | +/- | 0. |
| 1.16 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| -0.18 |
| -9999.00 |
| 0.75 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| 0.48 |
| -9999.00 |
| 0.09 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| 0.83 |
| -9999.00 |
| -1.84 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -0.16 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| 0.47 |
| -9999.00 |
| 0.07 |
| -9999.00 |
| 0.17 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -2.43 |
| -9999.00 |
| 0.86 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6322. | +/- | 125. |
| -10000. | +/- | -NaN |
| 3.07 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.29 | +/- | 0. |
| 1.29 | +/- | -NaN |
| -0.80 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.44 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.26 | +/- | 4. |
| -9999.99 | +/- | -NaN |
| 0.47 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.70 | +/- | 0. |
| 0.67 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.44 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| -0.38 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.68 |
| -9999.00 |
| 0.68 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.37 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| 0.36 |
| -9999.00 |
| 0.39 |
| -9999.00 |
| 0.43 |
| -9999.00 |
| 0.96 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.71 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| -0.47 |
| -9999.00 |
| -0.80 |
| -9999.00 |
| -1.35 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -0.50 |
| -9999.00 |
| -0.96 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| -2.48 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7973. | +/- | 184. |
| -10000. | +/- | -NaN |
| 3.96 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.09 | +/- | 1. |
| 1.09 | +/- | -NaN |
| -1.26 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.46 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.01 | +/- | 6. |
| -9999.99 | +/- | -NaN |
| 0.96 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 29.47 | +/- | 0. |
| 1.47 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.46 |
| -9999.00 |
| -0.45 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| -2.24 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -2.23 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -2.02 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.98 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| 0.72 |
| -9999.00 |
| 0.88 |
| -9999.00 |
| -1.23 |
| -9999.00 |
| -1.22 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -1.33 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -2.34 |
| -9999.00 |
| -0.95 |
| -9999.00 |
| -9999.00 |
| -9999.00 |
| -1.66 |
| -9999.00 |
| -1.33 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 7491. | +/- | 130. |
| -10000. | +/- | -NaN |
| 3.12 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.22 | +/- | 1. |
| 1.22 | +/- | -NaN |
| -0.55 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.10 | +/- | 3. |
| -9999.99 | +/- | -NaN |
| 0.38 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 28.89 | +/- | 0. |
| 1.46 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.04 |
| -9999.00 |
| 0.49 |
| -9999.00 |
| 0.10 |
| -9999.00 |
| 0.54 |
| -9999.00 |
| -0.55 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.90 |
| -9999.00 |
| 0.20 |
| -9999.00 |
| -1.00 |
| -9999.00 |
| -0.10 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.71 |
| -9999.00 |
| 0.62 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| -0.72 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -0.11 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| -0.94 |
| -9999.00 |
| -0.87 |
| -9999.00 |
| -0.22 |
| -9999.00 |
| -0.14 |
| -9999.00 |
| -2.42 |
| -9999.00 |
| -2.30 |
| -9999.00 |
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6707. | +/- | 148. |
| -10000. | +/- | -NaN |
| 3.75 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 1.15 | +/- | 1. |
| 1.15 | +/- | -NaN |
| -0.78 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.05 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.33 | +/- | 1. |
| -9999.99 | +/- | -NaN |
| 0.56 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 2.85 | +/- | 0. |
| 0.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.09 |
| -9999.00 |
| -0.05 |
| -9999.00 |
| 0.42 |
| -9999.00 |
| 0.56 |
| -9999.00 |
| -2.50 |
| -9999.00 |
| 0.27 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.67 |
| -9999.00 |
| -1.01 |
| -9999.00 |
| 1.00 |
| -9999.00 |
| -0.69 |
| -9999.00 |
| 0.69 |
| -9999.00 |
| 0.30 |
| -9999.00 |
| 0.45 |
| -9999.00 |
| -2.45 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| -0.25 |
| -9999.00 |
| -0.28 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| 0.22 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| 0.41 |
| -9999.00 |
| -0.91 |
| -9999.00 |
| -0.73 |
| -9999.00 |
| -2.48 |
| -9999.00 |
| 0.24 |
| -9999.00 |
SUSPECT_BROAD_LINES
STAR_BAD,CHI2_BAD,SN_BAD
STAR_WARN,CHI2_WARN,SN_WARN
359-02-O
| 6481. | +/- | 124. |
| -10000. | +/- | -NaN |
| 2.97 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 4.60 | +/- | 0. |
| 4.60 | +/- | -NaN |
| -0.62 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| -0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.09 | +/- | 2. |
| -9999.99 | +/- | -NaN |
| 0.53 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 28.18 | +/- | 0. |
| 1.45 | +/- | -NaN |
| 0.00 | +/- | 0. |
| -9999.99 | +/- | -NaN |
| 0.00 |
| -9999.00 |
| 0.50 |
| -9999.00 |
| 0.06 |
| -9999.00 |
| 0.91 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.97 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| 0.89 |
| -9999.00 |
| 0.79 |
| -9999.00 |
| -0.29 |
| -9999.00 |
| -1.27 |
| -9999.00 |
| 0.99 |
| -9999.00 |
| 0.53 |
| -9999.00 |
| 0.82 |
| -9999.00 |
| 0.95 |
| -9999.00 |
| -1.09 |
| -9999.00 |
| -2.21 |
| -9999.00 |
| 0.03 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.84 |
| -9999.00 |
| -0.62 |
| -9999.00 |
| 0.94 |
| -9999.00 |
| 0.26 |
| -9999.00 |
| -1.08 |
| -9999.00 |
| -2.49 |
| -9999.00 |
| -2.42 |
| -9999.00 |
